Abstract
Macromolecular complexes are essential to conserved biological processes, but their prevalence across animals is unclear. By combining extensive biochemical fractionation with quantitative mass spectrometry, here we directly examined the composition of soluble multiprotein complexes among diverse metazoan models. Using an integrative approach, we generated a draft conservation map consisting of more than one million putative high-confidence co-complex interactions for species with fully sequenced genomes that encompasses functional modules present broadly across all extant animals. Clustering reveals a spectrum of conservation, ranging from ancient eukaryotic assemblies that have probably served cellular housekeeping roles for at least one billion years, ancestral complexes that have accrued contemporary components, and rarer metazoan innovations linked to multicellularity. We validated these projections by independent co-fractionation experiments in evolutionarily distant species, affinity purification and functional analyses. The comprehensiveness, centrality and modularity of these reconstructed interactomes reflect their fundamental mechanistic importance and adaptive value to animal cell systems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hartwell, L. H., Hopfield, J. J., Leibler, S. & Murray, A. W. From molecular to modular cell biology. Nature 402, C47–C52 (1999)
Alberts, B. The cell as a collection of protein machines: Preparing the next generation of molecular biologists. Cell 92, 291–294 (1998)
Butland, G. et al. Interaction network containing conserved and essential protein complexes in Escherichia coli. Nature 433, 531–537 (2005)
Krogan, N. J. et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440, 637–643 (2006)
Guruharsha, K. G. et al. A protein complex network of Drosophila melanogaster. Cell 147, 690–703 (2011)
Havugimana, P. C. et al. A census of human soluble protein complexes. Cell 150, 1068–1081 (2012)
Stelzl, U. et al. A human protein-protein interaction network: a resource for annotating the proteome. Cell 122, 957–968 (2005)
Li, S. et al. A map of the interactome network of the metazoan C-elegans. Science 303, 540–543 (2004)
Hu, P. et al. Global functional atlas of Escherichia coli encompassing previously uncharacterized proteins. PLoS Biol. 7, e1000096 (2009)
Rolland, T. et al. A proteome-scale map of the human interactome network. Cell 159, 1212–1226 (2014)
Sharan, R. et al. Conserved patterns of protein interaction in multiple species. Proc. Natl Acad. Sci. USA 102, 1974–1979 (2005)
Gandhi, T. K. B. et al. Analysis of the human protein interactome and comparison with yeast, worm and fly interaction datasets. Nature Genet. 38, 285–293 (2006)
Tan, K., Shlomi, T., Feizi, H., Ideker, T. & Sharan, R. Transcriptional regulation of protein complexes within and across species. Proc. Natl Acad. Sci. USA 104, 1283–1288 (2007)
Singh, R., Xu, J. B. & Berger, B. Global alignment of multiple protein interaction networks with application to functional orthology detection. Proc. Natl Acad. Sci. USA 105, 12763–12768 (2008)
Yu, H. et al. Annotation transfer between genomes: protein–protein interologs and protein–DNA regulogs. Genome Res. 14, 1107–1118 (2004)
Ideker, T. & Krogan, N. J. Differential network biology. Mol. Syst. Biol. 8, 565 (2012)
Kiemer, L. & Cesareni, G. Comparative interactomics: comparing apples and pears? Trends Biotechnol. 25, 448–454 (2007)
von Mering, C. et al. Comparative assessment of large-scale data sets of protein–protein interactions. Nature 417, 399–403 (2002)
Malovannaya, A. et al. Analysis of the human endogenous coregulator complexome. Cell 145, 787–799 (2011)
Lee, I., Blom, U. M., Wang, P. I., Shim, J. E. & Marcotte, E. M. Prioritizing candidate disease genes by network-based boosting of genome-wide association data. Genome Res. 21, 1109–1121 (2011)
Uhlen, M. et al. Towards a knowledge-based Human Protein Atlas. Nature Biotechnol. 28, 1248–1250 (2010)
McKusick, V. A. Mendelian Inheritance in Man: A Catalog of Human Genes and Genetic Disorders. (Johns Hopkins Univ. Press, 1998)
Kim, M. S. et al. A draft map of the human proteome. Nature 509, 575–581 (2014)
Rubin, G. M. et al. Comparative genomics of the eukaryotes. Science 287, 2204–2215 (2000)
Bezginov, A., Clark, G. W., Charlebois, R. L., Dar, V. U. N. & Tillier, E. R. M. Coevolution reveals a network of human proteins originating with multicellularity. Mol. Biol. Evol. 30, 332–346 (2013)
Stumpf, M. P. H. et al. Estimating the size of the human interactome. Proc. Natl Acad. Sci. USA 105, 6959–6964 (2008)
Hart, G. T., Ramani, A. K. & Marcotte, E. M. How complete are current yeast and human protein-interaction networks? Genome Biol. 7, 120 (2006)
Eisenberg, E. & Levanon, E. Y. Preferential attachment in the protein network evolution. Phys. Rev. Lett. 91, 138701 (2003)
Knoll, A. H. The early evolution of eukaryotes: a geological perspective. Science 256, 622–627 (1992)
Burstein, E. et al. COMMD proteins, a novel family of structural and functional homologs of MURR1. J. Biol. Chem. 280, 22222–22232 (2005)
van de Sluis, B., Rothuizen, J., Pearson, P. L., van Oost, B. A. & Wijmenga, C. Identification of a new copper metabolism gene by positional cloning in a purebred dog population. Hum. Mol. Genet. 11, 165–173 (2002)
McDonald, F. J. COMMD1 and ion transport proteins: what is the COMMection? Focus on “COMMD1 interacts with the COOH terminus of NKCC1 in Calu-3 airway epithelial cells to modulate NKCC1 ubiquitination”. Am. J. Physiol. Cell Physiol. 305, C129–C130 (2013)
Kolanczyk, M. et al. Missense variant in CCDC22 causes X-linked recessive intellectual disability with features of Ritscher-Schinzel/3C syndrome. Eur. J. Hum. Genet. 109, 1–6 (2014)
Voineagu, I. et al. CCDC22: a novel candidate gene for syndromic X-linked intellectual disability. Mol. Psychiatry 17, 4–7 (2012)
Toledo, C. M. et al. BuGZ is required for Bub3 stability, Bub1 kinetochore function, and chromosome alignment. Dev. Cell 28, 282–294 (2014)
Kotake, Y. et al. Splicing factor SF3b as a target of the antitumor natural product pladienolide. Nature Chem. Biol. 3, 570–575 (2007)
Croft, D. et al. The Reactome pathway knowledgebase. Nucleic Acids Res. 42, D472–D477 (2014)
Ovádi, J. Cell Architecture and Metabolite Channeling. (RG Landes Company, 1995)
Ruepp, A. et al. CORUM: the comprehensive resource of mammalian protein complexes-2009. Nucleic Acids Res. 38, D497–D501 (2010)
Warde-Farley, D. et al. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 38, W214–W220 (2010)
Franceschini, A. et al. STRING v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 41, D808–D815 (2013)
Pu, S., Wong, J., Turner, B., Cho, E. & Wodak, S. J. Up-to-date catalogues of yeast protein complexes. Nucleic Acids Res. 37, 825–831 (2009)
Kirkwood, K. J., Ahmad, Y., Larance, M. & Lamond, A. I. Characterization of native protein complexes and protein isoform variation using size-fractionation-based quantitative proteomics. Mol. Cell. Proteomics 12, 3851–3873 (2013)
Turner, B. et al. iRefWeb: interactive analysis of consolidated protein interaction data and their supporting evidence. Database 2010, baq023 (2010)
Stark, C. et al. BioGRID: a general repository for interaction datasets. Nucleic Acids Res. 34, D535–D539 (2006)
Uhlen, M. et al. Tissue-based map of the human proteome. Science 347, 6220 (2015)
de Bie, P. et al. Characterization of COMMD protein–protein interactions in NF-κB signalling. Biochem. J. 398, 63–71 (2006)
Phillips-Krawczak, C. A. et al. COMMD1 is linked to the WASH complex and regulates endosomal trafficking of the copper transporter ATP7A. Mol. Biol. Cell 26, 91–103 (2015)
Yanai, I., Peshkin, L., Jorgensen, P. & Kirschner, M. W. Mapping gene expression in two Xenopus species: evolutionary constraints and developmental flexibility. Dev. Cell 20, 483–496 (2011)
Acknowledgements
We thank G. Bader, P. Kim, G. Moreno-Hagelsieb, S. Pu and S. Wodak for critical suggestions, illustrator A. Syrett for expert help drafting figures, T. Kwon (University of Texas) for X. laevis gene models, and K. Foltz (University of California, Santa Barbara), A. Brehm (Philipps-University Marburg), P. Paddison (Fred Hutchinson Cancer Research Center), J. Smith (Woods Hole Marine Biological Laboratory), P. Zandstra and J. Moffat (University of Toronto) for providing biological specimens and reagents. We thank members of the Emili and Marcotte laboratories for assistance and guidance, and SciNet (University of Toronto) and the Texas Advanced Computing Center (University of Texas) for high-performance computing resources. This work was supported by grants from the CIHR, NSERC, ORF and the CFI to A.E., from the CIHR and Heart and Stroke to J. P., from the NIH (F32GM112495) to K.D., and from the NIH, NSF, CPRIT, and Welch Foundation (F-1515) to E.M.M.
Author information
Authors and Affiliations
Contributions
A.E. and E.M.M. designed and co-supervised the project. C.W. performed proteomic experiments, aided by P.C.H. B.B. coordinated data analysis, aided by S.Ph., K.D. and S.S., and guided by E.M.M. E.R.T., G.Cl., A.B., J.P., X.X., K.C., G.Cr., C.W. and S.Ph. analysed network and conservation data. C.W., F.T., O.K., J.K., S.Pa., O.P., Z.N., D.R.B., X.G., R.H.M, M.S., J.G., M.B., W.B.D. and J.B.W. contributed validation experiments. S.Ph. designed the web portal. C.W., B.B., E.M.M. and A.E. drafted the manuscript. All authors discussed results and contributed edits.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Performance measures.
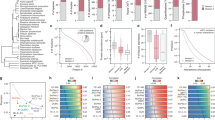
a, Performance benchmarks, measuring the precision and recall of our method and data in identifying known co-complex interactions from a withheld reference set of annotated human complexes (from CORUM39; as in Fig. 2b). Fivefold cross-validation against this withheld set shows strong performance gains, beyond a baseline achieved using only human and mouse co-fractionation data along with additional evidence from independent protein interaction screens5,19 and a functional gene network20 (far-left curve), made by integrating co-fractionation data from the additional non-human animal species (as indicated). ‘All data’ and ‘Fractionation data only’ curves include biochemical fractionation data from all five input species: human, mouse, urchin, fly and worm; the latter curve omits all external data. In all cases, at least two species were required to show supporting biochemical evidence. Recall refers to the fraction of 4,528 total positive interactions derived from the withheld human CORUM complexes. b, All 16,655 interactions were identified at least in two species, half (49%, 8,121) found in three or more species. c, Among these high-confidence co-complex interactions, 8,981 (54%) were not reported in iRefWeb44 (v13.0), BioGRID45 (v3.2.119) or CORUM reference (Supplementary Table 2) for any of the five input species or in yeast; half (46%, 4,128) of these novel co-complex interactions display evidence of co-fractionation in three or more species. d, Final precision/recall performance on withheld interaction test set. A support vector machine classifier was trained using interactions derived from our training set of CORUM complexes, then ∼1 million protein pairs found to co-elute in at least two of the five input species were scored by the classifier. Black curve shows precision and recall for ranked list of co-eluting pairs, with recall representing the fraction recovered of 4,528 total positive interactions derived from the withheld set of merged human CORUM complexes, and precision measured using co-eluting pairs where both members of the pair are contained in the set of proteins represented in the CORUM withheld set. The top 16,655 pairs, giving a cumulative precision of 67.5% and recall of 23.0% on this withheld test set, form the high-confidence set of co-complex protein–protein interactions (blue circle). The highest-scoring interactions were clustered using the two-stage approach described in the Supplementary Methods, yielding a final set of 7,669 interactions, which form the 981 identified complexes (red circle; precision = 90.0%, recall = 20.8%).
Extended Data Figure 2 Properties of protein elution profiles.
a, Distribution of global protein tissue expression pattern similarity, measured as the Pearson correlation coefficient of protein abundance across 30 human tissues23, showing markedly higher correlations for 16,468 protein–protein pairs of putative co-complex interaction partners compared to the same number of randomized pairs of proteins in the network which were not predicted to interact. b, Heat map illustrating the low to moderate cross-species Spearman’s rank correlation coefficients in the elution profiles observed between orthologous proteins during mixed-bed ion exchange chromatography under standardized conditions, highlighting the shift in absolute chromatographic retention times in different species. This variation indicates that the conservation of co-fractionation by putatively interacting proteins is not merely a trivial result stemming from fixed column-retention times. c, The degree of co-fractionation is measured as the correlation coefficient between elution profiles. Spatial proximity is calculated from the mean of residue pair distances between components of multisubunit complexes with known three-dimensional structures (see Supplementary Methods).
Extended Data Figure 3 Derivation of complexes.
a, The 2,153 proteins present in the 981 derived metazoan complexes participate in multiple assemblies (‘moonlighting’) to an extent comparable to the sharing of subunits reported for literature-derived complexes (CORUM). For comparison, we examined the 1,550 unique proteins from the full CORUM set of 1,216 human complexes passing our selection criteria for supporting evidence (‘Unmerged’) and the 1,461 unique proteins from the non-redundant set of 501 merged complexes used as the reference for splitting our training and testing sets, with some of the largest complexes removed to avoid bias in training (‘Merged’; see ‘Optimizing the two-stage clustering’ in Supplementary Methods for details). b, Schematic of 981 identified complexes containing 2,153 unique proteins. In this graphical representation, 7,669 co-complex interactions are shown as lines, and proteins as nodes. Red and green interactions were previously annotated in CORUM. Red interactions were used in training the classifier and/or clustering procedure, while green interactions were held out for validation purposes. Grey interactions were not previously annotated in CORUM.
Extended Data Figure 4 Properties of new and old proteins and complexes.
a, The 2,153 protein components in the conserved animal complexes tend to be more ancient than the 2,301 proteins reported in the CORUM reference complexes or in two recent large-scale protein interaction assays, based on either the 7,062 proteins found by affinity purification/mass spectrometry (AP/MS; E. L. Huttlin et al., BioGRID preprint 166968, http://thebiogrid.org/166968/publication/) or the 3,667 proteins analysed by yeast two-hybrid assays (Y2H)10. Ages are derived from OMA (Orthologous Matrix database) as in ref. 25. b, Annotation rates (mean count of annotation terms per protein) of old and new proteins in the derived complexes and pairwise PPIs, compared with proteins in the CORUM reference complex set. Old proteins (defined by OMA) from the complexes generally exhibited higher annotation rates than new proteins. c, Differential enrichment of old, mixed and metazoan-specific protein complexes for functional annotations (select GO-slim biological process terms shown, top) and protein domains (Pfam, bottom).
Extended Data Figure 5 Abundance and expression trends for proteins in complexes.
Proteins within the identified complexes tend to be ubiquitously expressed across human tissues. a, b, Pie charts show the proportions of proteins with varying tissue expression patterns, from a recently published human tissue proteome map46, comparing the full set of 20,258 human proteins (a) with the 2,131 proteins within the identified complexes (b). Consistent with these observations, 91% of the protein components in the complexes were expressed in >15 tissues in data from a reference human proteome23, compared to less than half (46%) of the 17,294 proteins in the overall reference set (Z-test P < 0.001). c, d, The distributions of average mRNA (c, data from EBI accession E-MTAB-1733) and protein (d, data from PaxDb integrated data set, 9606-H.sapiens_whole_organism-integrated_data set) abundances for all proteins identified and those within complexes. Evolutionarily old proteins (defined by OMA as described in ref. 25 and mentioned earlier) tend towards higher abundances, even for proteins in reference complexes.
Extended Data Figure 6 Additional validation data.
a, Confirmation of MIB2 interactions by co-immunoprecipitation. Extract (∼10 mg protein) from cultured human HCT116 cells expressing Flag-tagged MIB2 or control (WT) cells was incubated with 100 µl anti-Flag M2 resin for 4 h while gently rotating at 4 °C. After extensive washing with RIPA buffer, co-purifying proteins bound to the beads were eluted by the addition of 25 µl Laemmli loading buffer at 95 °C. Polypeptides were separated by SDS–PAGE and immunoblotted using Flag, VPS4A, VPS4B or IST1 antibodies as indicated (expanded gel images provided in Supplementary Information). b, Protein co-complex interactions reported in the CYC2008 yeast protein complex database42 are reconstructed accurately from the co-fractionation data, regardless of whether the full set of co-fractionation plus external data are used to derive protein interactions (‘All data’, see also Fig. 4b) or if the external yeast data was specifically excluded from the analyses (‘All data, excluding yeast’).
Extended Data Figure 7 Agreement of derived complexes’ molecular weights with measurement by HPLC and density centrifugation.
a, CORUM reference complexes’ inferred molecular weights (MW) are consistent with their components’ average cumulative size-exclusion chromatograms. The molecular weight of each complex was calculated as the sum of putative component molecular weights, assuming 1:1 stoichiometry. Data from ref. 43 were analysed as in Fig. 4c and show a similar trend as for the derived complexes. b, Derived complexes’ inferred molecular weights are broadly consistent with their components’ average cumulative ultracentrifugation profiles on a sucrose density gradient. Average profiles are plotted for X. laevis orthologues, based on a preparation of haemoglobin-depleted heart and liver proteins separated on a 7–47% sucrose density gradient, as described in the Supplementary Methods.
Extended Data Figure 8 Distribution of uncharacterized proteins and novel interactions across the 981 derived complexes.
Complexes were sorted by median age (defined by OMA). Among 2,153 unique proteins, 293 (red) lack Gene Ontology (GO) functional annotations, while 1,756 of 7,665 co-complex interactions are novel (light green) (not listed in iRefWeb curation database).
Extended Data Figure 9 Properties of the Commander complex.
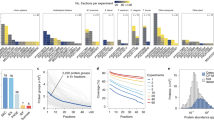
The automatically derived 8 subunit Commander complex (Fig. 3b) was subsequently extended to 13 subunits (COMMD1 to 10, CCDC22, CCDC93, and SH3GLB1) based on combined analysis of AP/MS (Fig. 4a), size-exclusion chromatograms43 (Fig. 4d), published pairwise interactions30,47,48, and analysis of elution profiles of the remaining COMM-domain-containing proteins, as shown here. Example protein elution profiles are plotted for Commander complex subunits observed from: HEK293 cell nuclear extract (a); sea urchin embryonic (5 days post-fertilization) extract (b); and fly SL2 cell nuclear extract (c); each fractionated by heparin affinity chromatography. d, Co-expression of Commander complex subunits during embryonic development of X. tropicalis (plotting mean ± s.d. of three clutches; data from ref. 49). e, Messenger RNA expression patterns of Commander complex subunits in stage 15 X. laevis embryos. Images show coordinated spatial expression in early vertebrate embryogenesis, as measured by in situ hybridization (three embryos examined). f, Knockdown of Commd2 induced marked head and eye defects in developing X. laevis. Top, Commd2 antisense knockdown significantly decreased eye size, shown for stage 38 tadpoles (from three clutches; control n = 47 animals, one eye each; ***P < 0.0001, two-sided Mann–Whitney test); phenotypes were consistent between translation blocking (MOatg; n = 60) morpholino reagents, splice site blocking (MOsp; n = 50) morpholinos, and knockdowns of interaction partner Commd3 (see Fig. 5a). Bottom, Commd2-knockdown induced altered Pax6 patterning in the embryonic eye (control n = 8 animals, two eyes each; MO n = 11). g, Commd2/3-knockdown animals show altered neural patterning. Changes in stage 15 X. laevis embryos, measured by in situ hybridization (assayed in duplicates; five embryos per treatment), seen upon knockdown but not on controls: the forebrain marker PAX6 was expanded, while the mid-brain marker EN2 was strongly reduced. Notably, while expression of KROX20/EGR1 in rhombomere R3 was shifted posteriorly, expression in R5 was strongly reduced or entirely absent. Panels in Fig. 5b are reproduced from this figure and are directly comparable. h, Confirmation of splice-blocking Commd2 morpholino activity. Images and schematic show the basis and results of RT–PCR and agarose gel electrophoresis obtained with the corresponding X. laevis knockdown tadpoles.
Extended Data Figure 10 Supporting data for BUB3 and CCDC97 experiments.
a, Sequence alignment showing conservation of ZNF207 GLEBS domain. b, Targeted CRISPR/Cas9-induced knockout of CCDC97 in two independent lines of human HEK293 cells, as verified by western blotting (expanded gel images provided in Supplementary Information). c, Loss of CCDC97 impairs cell growth. Lines show growth curves of control versus knockout cell lines in two biological replicate assays.
Supplementary information
Supplementary Information
This file contains Supplementary Methods and Data and additional references. (PDF 389 kb)
Supplementary Tables
This file contains Supplementary Tables 1-9 as follows: (1) Sample information (2) Human protein interactions + interologs detected in other 8 experimentally studied species + CORUM interaction reference standards (3) List co-complex interactions projected for 122 sequenced eukaryotic species (4) Final set of (981) conserved animal protein complexes (5) Protein age and conservation profile across 122 sequenced eukaryotic species (6) GOSlim domain, disease and phenotype enrichment results (7) Human disease annotations (8) Consecutive pathway and metabolic enzymes and (9) 36 common metabolites excluded from Recon2. (ZIP 16512 kb)
Supplementary Information
This file contains the Western blots gels for Extended Data Figures 6a and 10b. (PDF 1782 kb)
Rights and permissions
About this article
Cite this article
Wan, C., Borgeson, B., Phanse, S. et al. Panorama of ancient metazoan macromolecular complexes. Nature 525, 339–344 (2015). https://doi.org/10.1038/nature14877
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14877
This article is cited by
-
Analysis of the effects of importin α1 on the nuclear translocation of IL-1α in HeLa cells
Scientific Reports (2024)
-
Molecular complex detection in protein interaction networks through reinforcement learning
BMC Bioinformatics (2023)
-
Co-fractionation–mass spectrometry to characterize native mitochondrial protein assemblies in mammalian neurons and brain
Nature Protocols (2023)
-
Targeted cross-linker delivery for the in situ mapping of protein conformations and interactions in mitochondria
Nature Communications (2023)
-
Mapping protein states and interactions across the tree of life with co-fractionation mass spectrometry
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.