Abstract
Arising from B. J. Venters & B. F. Pugh Nature 502, 53–58 (2013); doi:10.1038/nature12535
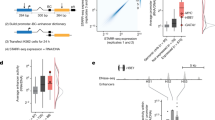
How cells locate the regions to initiate transcription is an open question, because core promoter elements (CPEs) are found in only a small fraction of core promoters1,2,3,4. A recent study5 measured 159,117 DNA binding regions of transcription factor IIB (TFIIB) by ChIP-exo (chromatin immunoprecipitation with lambda exonuclease digestion followed by high-throughput sequencing) in human cells, found four degenerate CPEs—upstream and downstream TFIIB recognition elements (BREu and BREd), TATA and initiator element (INR)—in nearly all of them, and concluded that these regions represent sites of transcription initiation marked by universal CPEs. We show that the claimed universality of CPEs is explained by the low specificities of the patterns used and that the same match frequencies are obtained with two negative controls (randomized sequences and scrambled patterns). Our analyses also cast doubt on the biological significance of most of the 150,753 non-messenger-RNA-associated ChIP-exo peaks, 72% of which lie within repetitive regions. There is a Retraction accompanying this Brief Communication Arising by Venters, B. J. & Pugh, B. F. Nature 511, http://dx.doi.org/10.1038/nature13588 (2014).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Ohler, U., Liao, G. C., Niemann, H. & Rubin, G. M. Computational analysis of core promoters in the Drosophila genome. Genome Biol. 3, RESEARCH0087 (2002)
Kadonaga, J. T. Perspectives on the RNA polymerase II core promoter. Wiley Interdiscip. Rev. Dev. Biol. 1, 40–51 (2012)
Lenhard, B., Sandelin, A. & Carnici, P. Metazoan promoters: emerging characteristics and insights into transcriptional regulation. Nature Rev. Genet. 13, 233–245 (2012)
Hartmann, H., Guthöhrlein, E. W., Siebert, M., Luehr, S. & Söding, J. P-value-based regulatory motif discovery using positional weight matrices. Genome Res. 23, 181–194 (2013)
Venters, B. J. & Pugh, B. F. Genomic organization of human transcription initiation complexes. Nature 502, 53–58 (2013)
Deininger, P. L. & Batzer, M. A. Mammalian retroelements. Genome Res. 12, 1455–1465 (2002)
Author information
Authors and Affiliations
Contributions
M.S. performed research and J.S. guided research; both M.S. and J.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
Declared none.
PowerPoint slides
Rights and permissions
About this article
Cite this article
Siebert, M., Söding, J. Universality of core promoter elements?. Nature 511, E11–E12 (2014). https://doi.org/10.1038/nature13587
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13587
This article is cited by
-
Chromatin-enriched lncRNAs can act as cell-type specific activators of proximal gene transcription
Nature Structural & Molecular Biology (2017)
-
Exome-based Variant Detection in Core Promoters
Scientific Reports (2016)
-
Bilaterian-like promoters in the highly compact Amphimedon queenslandica genome
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.