Abstract
DNA double-strand break (DSB) repair by homologous recombination has evolved to maintain genetic integrity in all organisms1. Although many reactions that occur during homologous recombination are known1,2,3, it is unclear where, when and how they occur in cells. Here, by using conventional and super-resolution microscopy, we describe the progression of DSB repair in live Escherichia coli. Specifically, we investigate whether homologous recombination can occur efficiently between distant sister loci that have segregated to opposite halves of an E. coli cell. We show that a site-specific DSB in one sister can be repaired efficiently using distant sister homology. After RecBCD processing of the DSB, RecA is recruited to the cut locus, where it nucleates into a bundle that contains many more RecA molecules than can associate with the two single-stranded DNA regions that form at the DSB. Mature bundles extend along the long axis of the cell, in the space between the bulk nucleoid and the inner membrane. Bundle formation is followed by pairing, in which the two ends of the cut locus relocate at the periphery of the nucleoid and together move rapidly towards the homology of the uncut sister. After sister locus pairing, RecA bundles disassemble and proteins that act late in homologous recombination are recruited to give viable recombinants 1–2-generation-time equivalents after formation of the initial DSB. Mutated RecA proteins that do not form bundles are defective in sister pairing and in DSB-induced repair. This work reveals an unanticipated role of RecA bundles in channelling the movement of the DNA DSB ends, thereby facilitating the long-range homology search that occurs before the strand invasion and transfer reactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Cromie, G. A., Connelly, J. C. & Leach, D. R. Recombination at double-strand breaks and DNA ends: conserved mechanisms from phage to humans. Mol. Cell 8, 1163–1174 (2001)
Cox, M. M. Regulation of bacterial RecA protein function. Crit. Rev. Biochem. Mol. Biol. 42, 41–63 (2007)
Dillingham, M. S. & Kowalczykowski, S. C. RecBCD enzyme and the repair of double-stranded DNA breaks. Microbiol. Mol. Biol. Rev. 72, 642–671 (2008)
Lesterlin, C., Gigant, E., Boccard, F. & Espeli, O. Sister chromatid interactions in bacteria revealed by a site-specific recombination assay. EMBO J. 31, 3468–3479 (2012)
Reyes-Lamothe, R., Possoz, C., Danilova, O. & Sherratt, D. J. Independent positioning and action of Escherichia coli replisomes in live cells. Cell 133, 90–102 (2008)
Monteilhet, C., Perrin, A., Thierry, A., Colleaux, L. & Dujon, B. Purification and characterization of the in vitro activity of I-Sce I, a novel and highly specific endonuclease encoded by a group I intron. Nucleic Acids Res. 18, 1407–1413 (1990)
Bzymek, M., Thayer, N. H., Oh, S. D., Kleckner, N. & Hunter, N. Double Holliday junctions are intermediates of DNA break repair. Nature 464, 937–941 (2010)
Renzette, N. et al. Localization of RecA in Escherichia coli K-12 using RecA–GFP. Mol. Microbiol. 57, 1074–1085 (2005)
Handa, N., Amitani, I., Gumlaw, N., Sandler, S. J. & Kowalczykowski, S. C. Single molecule analysis of a red fluorescent RecA protein reveals a defect in nucleoprotein filament nucleation that relates to its reduced biological functions. J. Biol. Chem. 284, 18664–18673 (2009)
Rehrauer, W. M. & Kowalczykowski, S. C. Alteration of the nucleoside triphosphate (NTP) catalytic domain within Escherichia coli recA protein attenuates NTP hydrolysis but not joint molecule formation. J. Biol. Chem. 268, 1292–1297 (1993)
Bakshi, S., Siryaporn, A., Goulian, M. & Weisshaar, J. C. Superresolution imaging of ribosomes and RNA polymerase in live Escherichia coli cells. Mol. Microbiol. 85, 21–38 (2012)
Garvey, N., St John, A. C. & Witkin, E. M. Evidence for RecA protein association with the cell membrane and for changes in the levels of major outer membrane proteins in SOS-induced Escherichia coli cells. J. Bacteriol. 163, 870–876 (1985)
Bell, J. C., Plank, J. L., Dombrowski, C. C. & Kowalczykowski, S. C. Direct imaging of RecA nucleation and growth on single molecules of SSB-coated ssDNA. Nature 491, 274–278 (2012)
Feinstein, E. et al. Single-molecule studies of the stringency factors and rates governing the polymerization of RecA on double-stranded DNA. Nucleic Acids Res. 39, 3781–3791 (2011)
Brenner, S. L., Zlotnick, A. & Griffith, J. D. RecA protein self-assembly. Multiple discrete aggregation states. J. Mol. Biol. 204, 959–972 (1988)
DiCapua, E., Schnarr, M., Ruigrok, R. W., Lindner, P. & Timmins, P. A. Complexes of RecA protein in solution. A study by small angle neutron scattering. J. Mol. Biol. 214, 557–570 (1990)
Chen, Z., Yang, H. & Pavletich, N. P. Mechanism of homologous recombination from the RecA–ssDNA/dsDNA structures. Nature 453, 489–494 (2008)
Story, R. M., Weber, I. T. & Steitz, T. A. The structure of the E. coli recA protein monomer and polymer. Nature 355, 318–325 (1992)
Yu, X., VanLoock, M. S., Yang, S., Reese, J. T. & Egelman, E. H. What is the structure of the RecA-DNA filament? Curr. Protein Pept. Sci. 5, 73–79 (2004)
Levin-Zaidman, S. et al. Ordered intracellular RecA–DNA assemblies: a potential site of in vivo RecA-mediated activities. Proc. Natl Acad. Sci. USA 97, 6791–6796 (2000)
Renzette, N. & Sandler, S. J. Requirements for ATP binding and hydrolysis in RecA function in Escherichia coli . Mol. Microbiol. 67, 1347–1359 (2008)
Kidane, D. & Graumann, P. L. Dynamic formation of RecA filaments at DNA double strand break repair centers in live cells. J. Cell Biol. 170, 357–366 (2005)
Gayathri, P. et al. A bipolar spindle of antiparallel ParM filaments drives bacterial plasmid segregation. Science 338, 1334–1337 (2012)
Bernhardt, T. G. & de Boer, P. A. Screening for synthetic lethal mutants in Escherichia coli and identification of EnvC (YibP) as a periplasmic septal ring factor with murein hydrolase activity. Mol. Microbiol. 52, 1255–1269 (2004)
Guzman, L. M., Belin, D., Carson, M. J. & Beckwith, J. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J. Bacteriol. 177, 4121–4130 (1995)
Nielsen, H. J., Ottesen, J. R., Youngren, B., Austin, S. J. & Hansen, F. G. The Escherichia coli chromosome is organized with the left and right chromosome arms in separate cell halves. Mol. Microbiol. 62, 331–338 (2006)
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000)
Gustafsson, M. G. et al. Three-dimensional resolution doubling in wide-field fluorescence microscopy by structured illumination. Biophys. J. 94, 4957–4970 (2008)
Schermelleh, L. et al. Subdiffraction multicolor imaging of the nuclear periphery with 3D structured illumination microscopy. Science 320, 1332–1336 (2008)
Sliusarenko, O., Heinritz, J., Emonet, T. & Jacobs-Wagner, C. High-throughput, subpixel precision analysis of bacterial morphogenesis and intracellular spatio-temporal dynamics. Mol. Microbiol. 80, 612–627 (2011)
Meile, J. C. et al. The terminal region of the E. coli chromosome localises at the periphery of the nucleoid. BMC Microbiol. 11, 28 (2011)
Badrinarayanan, A., Reyes-Lamothe, R., Uphoff, S., Leake, M. C. & Sherratt, D. J. In vivo architecture and action of bacterial structural maintenance of chromosome proteins. Science 338, 528–531 (2012)
Stohl, E. A. et al. Escherichia coli RecX inhibits RecA recombinase and coprotease activities in vitro and in vivo . J. Biol. Chem. 278, 2278–2285 (2003)
Sommer, S., Boudsocq, F., Devoret, R. & Bailone, A. Specific RecA amino acid changes affect RecA–UmuD′C interaction. Mol. Microbiol. 28, 281–291 (1998)
Sassanfar, M. & Roberts, J. Constitutive and UV-mediated activation of RecA protein: combined effects of recA441 and recF143 mutations and of addition of nucleosides and adenine. J. Bacteriol. 173, 5869–5875 (1991)
Karu, A. E. & Belk, E. D. Induction of E. coli recA protein via recBC and alternate pathways: quantitation by enzyme-linked immunosorbent assay (ELISA). Mol. Gen. Genet. 185, 275–282 (1982)
Wang, X., Lesterlin, C., Reyes-Lamothe, R., Ball, G. & Sherratt, D. J. Replication and segregation of an Escherichia coli chromosome with two replication origins. Proc. Natl Acad. Sci. USA 108, E243–E250 (2011)
Lesterlin, C., Pages, C., Dubarry, N., Dasgupta, S. & Cornet, F. Asymmetry of chromosome replichores renders the DNA translocase activity of FtsK essential for cell division and cell shape maintenance in Escherichia coli . PLoS Genet. 4, e1000288 (2008)
White, M. A., Eykelenboom, J. K., Lopez-Vernaza, M. A., Wilson, E. & Leach, D. R. Non-random segregation of sister chromosomes in Escherichia coli . Nature 455, 1248–1250 (2008)
Espeli, O., Mercier, R. & Boccard, F. DNA dynamics vary according to macrodomain topography in the E. coli chromosome. Mol. Microbiol. 68, 1418–1427 (2008)
McGinness, K. E., Baker, T. A. & Sauer, R. T. Engineering controllable protein degradation. Mol. Cell 22, 701–707 (2006)
Reyes-Lamothe, R., Sherratt, D. J. & Leake, M. C. Stoichiometry and architecture of active DNA replication machinery in Escherichia coli . Science 328, 498–501 (2010)
Helmstetter, C. E. et al. in Escherichia coli and Salmonella Vol. 2 (ed. Neidhardt, F. C. ) 1627–1639 (ASM, 1996)
Kuwada, N. J., Cheveralls, K. C., Traxler, B. & Wiggins, P. A. Mapping the driving forces of chromosome structure and segregation in Escherichia coli . Nucleic Acids Res. (2013)
Acknowledgements
This work was supported by an EMBO long-term fellowship to C.L. (LTF-535-2009); by a Wellcome Trust Programme Grant to D.J.S. (WT083469MA); and by a Wellcome Trust Strategic Award (091911) supporting advanced microscopy at Micron Oxford. We thank I. Dobbie for his support and advice on imaging. We thank D. Leach and S. Kowalczykowski for the gift of strains and plasmids, N. Dubarry for critical reading of the manuscript and M, Stouf for the gift of pMS11. We thank E. Birt and K. Prosser for preliminary experiments.
Author information
Authors and Affiliations
Contributions
D.J.S. and C.L. conceived the project, interpreted the data and wrote the paper. C.L. designed and undertook experiments. G.B. developed analytical tools for microscopy data and provided advice on the manuscript. L.S. provided expertise on 3D-SIM and advice on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 DSB induction.
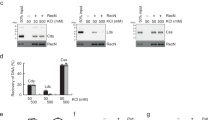
a, DSBs were introduced using the I-SceI endonuclease. The I-SceI cutting site (I-SceICS ) is an 18 bp site (5′-TAGGGATAACAGGGTAAT-3′) that was inserted 1,067 kb from oriC on the right replichore (codA-cynR intergenic region). I-SceICS is flanked by an upstream parS-P1 site (USe; 1.2 kb upstream of the cut) and a downstream parS-PMT1 site (DSe; 0.6 kb downstream of the cut) that binds different fluorescent ParB proteins. In vivo localization of USe and DSe was monitored in live cells by expressing the fluorescent fusions of the corresponding ParBP1- and ParBPMT1-binding proteins from pFHC2973 or pMS11. The I-SceI gene controlled by the arabinose-inducible promoter was expressed from the araB locus of the chromosome (782 kb from oriC), or alternatively expressed from a plasmid when a higher level of cutting was required. Two alleles of the I-SceI gene were used, namely the wt allele and an unstable allele carrying a degron tag (I-SceIMycDas ). The construction of the system is detailed in Methods. b, Cell viability of Rec+ and recA cells was assayed by plating efficiency after induction of I-SceIMycDas . Colony-forming units (c.f.u. ml−1) were counted 0, 30, 60, 120 and 180 min after addition of 0.2% arabinose in a steady-state population (OD600 nm = 0.2). Values at t = 0 were normalized to 1. Bottom, Southern blot shows the efficiency of DSB induction for the same time points, and the level of DSBs measured from three independent experiments. A cynT probe (500 bp) was used to visualize the substrate: a 12 kb SacII fragment containing the I-SceI cut site, or the product of restriction by I-SceI (downstream 3.1 kb fragment). Conditions were used in which 60 min after I-SceI was induced, most cells with segregated sisters received no more than a single DSB, consistent with the overall level of cutting observed biochemically (∼57%) and the dependence on RecA for survival. c, Flow cytometry showing the DNA content distribution of TB28 strains in the stationary phase, during exponential growth (M9 glucose 30 °C) and after rifampicin/cephalexin run-out. Subpopulations harbouring 1 or 2 chromosome equivalents after run-out indicate the number of replication origins per cell (1.91), which was used to calculate the C (DNA synthesis period) + D (time between completion of DNA replication and cell division) period (74 min) using Helmstetter’s model (origin per cell = 2(C+D)/t) (ref. 43). A summary of the cell cycle parameters is shown. All experiments were performed in identical growth conditions in which cells harbour only one replicating chromosome, with initiation of replication occurring ∼6 min after birth in the T = 80 min cell cycle. d, Snapshot analysis of USe localization before (left) and after (right) 60 min of DSB induction showed an identical pattern compared with DSe (Fig. 1A). From left to right: ratio of cells with 0, 1 or 2 foci; dot plots of USe foci positions as a function of cell length; histograms of USe position along the cell long axis and the ISD (in μm) are shown. Bottom, a plot of ISD against cell length shows that DSB induction triggers the global decrease of ISD values and generates a population of long cells with reduced ISD (framed in green), revealing the pairing of sister loci.
Extended Data Figure 2 Consequences of DSB induction.
A, Analysis of DSB-induced DSe sister pairing in Rec+ cells. a, Time-lapse particle tracking of DSe foci (5 min per frame) with indicative images (every 20 min) showing pairing between segregated sister loci (DSB was induced 30 min before the beginning of time-lapse acquisition). The red dashed line indicates the sister-pairing event and the grey area indicates the pairing period. b, DSe focus fluorescence intensity quantification performed on two consecutive frames—that is, before and after pairing—shows that the one focus observed after pairing retained 104 ± 15% of the added fluorescence of the previous sister focus (normalized at 100%), consistent with colocalization of the two localization arrays. c, Distances between DSe and USe focus centres remained unchanged upon DSB induction in Rec+ and recA cells, indicating that both DNA ends remain coincident throughout cutting, pairing and repair. d, Summary diagram of sister pairing that occurred 103 ± 46 min after DSB induction and lasted for 50 ± 34 min (grey area). Once initiated, pairing occurred rapidly (<5 min) and engaged both DSB ends. B, a, DSB induction generated a variety of cell types harbouring 0, 1 or 2 DSe foci with 0, 1 or 2 USe due to loss of the DSe and/or USe foci. One image for each of the nine cell types is shown with their quantification 0, 60 and 120 min after DSB induction in Rec+ and recA strains. These data are displayed using a square density representation. This was obtained with a custom MATLAB code that uses the data generated by the spotfinderM function of MicrobeTracker software. Only cells >3.3 μm are considered in order to focus the analysis on the fraction of cells in which the I-SceI cut site region (codA-cynR) has already segregated. b, Two-colour time-lapse analysis after DSB induction showed that DSe and USe foci belonging to the same DSB disappeared within 0–5 min from one another in 71% of cases (green and red lines), and recovered within 0–10 min from one another in 87% of cases (green and red arrows), consistent with bidirectional processing of the two ends of the DSB by RecBCD (see Supplementary Discussion). C, Time-lapse analysis of RecBCD-mediated complete degradation of the nucleoid after DSB induction in recA cells. One example is shown with nucleoid DNA stained with DAPI (green), cell membranes stained with FM4-64 (red). The decrease in fluorescence intensity of nucleoids undergoing degradation (red curve) was measured and reflects the rate of DNA degradation in vivo (Supplementary Video 1). Decrease in fluorescence of non-degraded nucleoids is accounted for by photobleaching of the DAPI (blue curve). The overall rate of DNA degradation was 23.2 kb min−1 (386 bp s−1); that is, 193 bp s−1 per RecBCD complex (n = 38 cells analysed).
Extended Data Figure 3 RecA bundle characterization.
A, Survival after ultraviolet (UV) irradiation shows complementation of recA-gfp allele sensitivity (SS0385 strain) by expression of a wild-type recA gene in the fhuB ectopic locus of the chromosome or from a plasmid (pSC101-recA). RecA–GFP strain (TB28, recA-gfp, fhuB::recA) showed a level of UV resistance similar to the wild-type (wt) strain. B, a, Snapshot analysis of RecA–GFP structures induced by DSBs. Each image was obtained by projection of a three-dimensional stack (15 z-sections of 125 nm z-distance) of 3D-SIM images for RecA–GFP (green) merged with wide-field images for DSe (red). Before DSB induction, RecA is diffuse in most cells (95%) and spontaneous spots are distant from DSe foci. Sixty minutes after DSB induction, RecA spots form in close proximity to a DSe focus and RecA bundles are observed in 29% of total cells, regardless of whether cells have one or two DSe foci. b, Bundle length (nm), and the thickness of the large central body and the thinner extensions is shown in the histogram on the left (nm). The width of the thinner extensions (≤100 nm) corresponds to the lateral resolution limit of 3D-SIM (see Supplementary Videos 2–7). c, In cells with two DSe foci and a RecA bundle, 47% of RecA–GFP bundles were associated with one DSe focus and 44% were associated with both DSe foci. C, FRAP of RecA bundles. Fluorescence intensities of bleached (green curve) and non-bleached regions (blue curve) of RecA bundles were monitored up to 1,500 s after bleaching. No fluorescence recovery was observed throughout this period, as shown by the indicative time-lapse images below. The green and blue boxes indicate the bleached and non-bleached regions of the bundle, respectively (time in s). Initial fluorescence was normalized to 1 (Supplementary Video 3). D, RecA bundle formation after UV irradiation. The ratio of cells with RecA bundle(s) after irradiation with UV light (50 J m−2) was measured, as was the total fluorescence intensity of RecA–GFP. The level of the SOS response, induced in response to DNA damage, was reported by the expression of the m-cherry gene controlled by the PsulA SOS-inducible promoter. Time-lapse images of RecA–GFP after UV irradiation confirms rapid bundle formation after UV treatment and snapshot images show multiple bundles present in most cells 45 min after UV irradiation (Supplementary Video 8). E, Relative frequencies of RecA bundles and RecA spots 60 min after DSB induction at codA (356.6 kb) or ydeO (1,581.7 kb) cut sites. ydeO and codA loci have different replication–segregation timings as manifested by the fractions of two-focus cells (8% and 69%, respectively). Images below show the morphology of RecA bundles observed in those two cell populations (see Supplementary Discussion).
Extended Data Figure 4 Movement of loci after DSB induction and recruitment of recombination proteins.
A, a, Mobility analysis of uncut loci. b, Comparative mobilities of long-travelling and the short-travelling sister foci during DSB-induced sister pairing. c, Mean square displacement values (MSDs), and MSD scaling parameters (α); α = 0.5 for the uncut locus compared with 0.69 for the long-travelling sister and 0.41 for the short-travelling one. This compares with α = 1 for diffusive movement and α = 2 for processive movement44. d, Summary showing the diffusion coefficients Dapp, the directionality of movement and the scaling factor values for uncut loci, and long- and short-travelling loci. B, Survival after UV irradiation of cells expressing RecN–YPet, PriA–YPet and RecG–YPet fluorescent proteins expressed from their endogenous promoters in their normal chromosomal position. All four fusion proteins were recombination proficient, as demonstrated by resistance to UV irradiation. C, Histogram of the fraction of cells harbouring RecN–YPet, RecG–YPet or PriA–YPet fusions showing DSB-induced focus formation in Rec+, but not in recB cells. Averages and standard deviations are calculated form snapshot images of three independent experiments. Bottom, the percentage of RecN–YPet, PriA–YPet and RecG–YPet foci that colocalized (<0.5 μm apart) with a DSe focus is shown with indicative images. Residence times (s) of these foci, estimated by time-lapse analysis, are indicated (see Supplementary Discussion).
Extended Data Figure 5 RecA bundles are not artefacts and require that RecA can bind ATP.
a, Immunocytochemistry using anti-RecA antibodies and Alexa-594-conjugated secondary antibodies. Left, bundles formed by wild-type RecA 60 min after DSB induction (in >25% of analysable cells) or 10 min after UV induction (in >80% of analysable cells). In the recAwt/recA–gfp diploid strain, bundles were visualized both by immunocytochemistry and by direct observation of RecA–GFP. b, Visualization of RecA803–RFP upon DNA damage induction. Merge of phase contrast and RFP images acquired with conventional fluorescent microscopy are shown. RecA803–RFP appeared diffuse in the cell before damage induction, and showed bundle formation similar to that observed with the RecA–GFP fusion after DSB induction (same conditions as in Figs 1 and 2) or UV irradiation (10 J m−2). Histogram presents the fractions of cells with RecA803–RFP bundles in corresponding conditions. c, Survival of cells carrying different recA alleles after DSB induction and UV irradiation (10 J m−2), estimated by a plating assay. d, 3D-SIM images of RecA–GFP, RecA(K72A)–GFP and RecA(K72R)–GFP fusions after DSB induction (same conditions as in Figs 1 and 2) or UV induction (10 J m−2). Histograms present the fraction of cells with diffuse fluorescence, cells with RecA spots or RecA bundles (‘others’ corresponds to cells with saturated signal or aberrant morphologies).
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-3, Supplementary Discussion and Supplementary References. (PDF 258 kb)
Time-lapse of nucleoid degradation after DSB-induction in recA cells.
From the left: Merge, Phase-contrast, DAPI, FM4-64 (3 min/frame over 190 min). Cell-by-cell differences are due to the fact that induction of DSBs is asynchronous in the population. (MP4 1628 kb)
Time-lapse of RecA bundle assembly (left) and directional disassembly (right) with corresponding fluorescence density dynamic plot (30 s/frame over 30 min)
Time-lapse of RecA bundle assembly (left) and directional disassembly (right) with corresponding fluorescence density dynamic plot (30 s/frame over 30 min). (MP4 1093 kb)
Fluorescence Recovery After Photobleaching of RecA bundles.
Three experiments are shown, from 10 min before bleaching to 25 min after bleaching. (MP4 2378 kb)
41586_2014_BFnature12868_MOESM11_ESM.mpg
4D rendering of time-lapse 3D-SIM of RecA bundles (2s/Frame), with orthogonal view showing the rapid movement of the bundle extension 4D rendering of time-lapse 3D-SIM of RecA bundles (2s/Frame), with orthogonal view showing the rapid movement of the bundle extension. (MPG 7366 kb)
Rights and permissions
About this article
Cite this article
Lesterlin, C., Ball, G., Schermelleh, L. et al. RecA bundles mediate homology pairing between distant sisters during DNA break repair. Nature 506, 249–253 (2014). https://doi.org/10.1038/nature12868
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12868
This article is cited by
-
In vivo tracking of functionally tagged Rad51 unveils a robust strategy of homology search
Nature Structural & Molecular Biology (2023)
-
Three-dimensional structured illumination microscopy with enhanced axial resolution
Nature Biotechnology (2023)
-
Isolation, Characterization, and Comparative Genomic Analysis of vB_Pd_C23, a Novel Bacteriophage of Pantoea dispersa
Current Microbiology (2023)
-
RecA finds homologous DNA by reduced dimensionality search
Nature (2021)
-
Two-step chromosome segregation in the stalked budding bacterium Hyphomonas neptunium
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.