Abstract
Cultivated bacteria such as actinomycetes are a highly useful source of biomedically important natural products. However, such ‘talented’ producers represent only a minute fraction of the entire, mostly uncultivated, prokaryotic diversity. The uncultured majority is generally perceived as a large, untapped resource of new drug candidates, but so far it is unknown whether taxa containing talented bacteria indeed exist. Here we report the single-cell- and metagenomics-based discovery of such producers. Two phylotypes of the candidate genus ‘Entotheonella’ with genomes of greater than 9 megabases and multiple, distinct biosynthetic gene clusters co-inhabit the chemically and microbially rich marine sponge Theonella swinhoei. Almost all bioactive polyketides and peptides known from this animal were attributed to a single phylotype. ‘Entotheonella’ spp. are widely distributed in sponges and belong to an environmental taxon proposed here as candidate phylum ‘Tectomicrobia’. The pronounced bioactivities and chemical uniqueness of ‘Entotheonella’ compounds provide significant opportunities for ecological studies and drug discovery.
Similar content being viewed by others
Main
More than half of the known natural products with antimicrobial, antitumour or antiviral activity are of bacterial origin1. Most of these compounds were isolated from cultivated representatives of only five bacterial groups: filamentous actinomycetes, Myxobacteria, Cyanobacteria, and members of the genera Pseudomonas and Bacillus. Uncultivated bacteria, which are proposed to form 70% of all known prokaryotic phyla2, represent a particularly promising source for new, chemically prolific taxa. However, except for individual biosynthetic pathways reported from environmental sources3,4, the true metabolic potential of these microbes remains unexplored. Two such pathways, involved in the production of onnamide- and theopederin-type polyketides5 and ribosomal peptides of the polytheonamide group6 (Fig. 1), were previously discovered in the marine sponge Theonella swinhoei. Like many other sponges, this animal harbours a massive consortium of uncultivated bacteria belonging to hundreds of distinct phylotypes7,8,9. T. swinhoei is the source of exceptionally diverse natural products and forms distinct chemotypes; samples of the sponge collected from different locations have largely non-overlapping metabolite profiles. From the onnamide and polytheonamide chemotype occurring at Hachijo Jima, Japan, here termed T. swinhoei Y (Y referring to the yellow interior of the sponge), in total more than 40 bioactive polyketides and modified peptides belonging to seven structural classes were isolated (Fig. 1)10. As previous work on onnamides and polytheonamides has produced only metagenomic DNA fragments lacking taxonomically diagnostic features, it was unknown which members of the bacterial community are the producers of these compounds.
Polytheonamides A and B differ in the stereochemistry of the sulphoxide moiety (polytheonamide A shows S chirality; polytheonamide B shows R chirality).
Attribution of metabolic genes to ‘Entotheonella’
Single-cell analysis has recently emerged as an efficient strategy to correlate the phylogenetic identity of environmental microorganisms with their functional gene repertoire11,12,13. To pinpoint producers in T. swinhoei Y, samples enriched in bacteria of different cell densities were prepared by differential centrifugation after sponge collection. When a fraction of higher density (Fig. 2a) was microscopically examined, we found that it contained a highly enriched population of large filamentous bacteria that fluoresce when excited with ultraviolet light (Fig. 2b). The bacteria were morphologically similar to the symbiont ‘Candidatus Entotheonella palauensis’ previously reported from a Palauan Theonella swinhoei chemotype and suspected as producer of antifungal peptides14,15. Scanning electron micrographs (Fig. 2c) revealed the presence of approximately 2- to 3-μm cells linked to each other. These bacteria, as well as those from the low-density fraction containing mostly unicellular bacteria, were sorted individually into 96-well plates by fluorescence-assisted cell sorting (FACS) (Extended Data Fig. 1a), resulting in filamentous and unicellular plates. Subsequently, multiple displacement amplification (MDA) of single bacterial genomes was performed on each well, resulting in DNA product sizes of approximately 10 kb (Extended Data Fig. 1b).
a, Differential interference contrast micrograph of the filamentous fraction after differential centrifugation (n = 3). b, Fluorescence micrograph of filamentous bacteria without (top) and with (bottom) ultraviolet excitation (n = 3). c, Scanning electron micrograph of a single filamentous bacterium (n = 3). d, Nested PCR of natural product gene clusters from whole-genome amplification samples of wells sorted with single filaments (n = 48). Wells showing positive amplification for ‘Entotheonella’ sp. 16S rRNA gene, onnamide (onn) and polytheonamide (poy) gene clusters (n = 3) were used for the identification of the other enzyme clusters (n = 1). Each lane represents a single well defined by the well identifier above the top row. cth, cyclotheonamide; ker, keramamide; kon, konbamide; naz, nazumamide. PC, metagenomic DNA from filamentous fraction; pts, unknown proteusin.
To detect wells containing DNA from the onnamide or polytheonamide producer, primers specific for onn and poy genes encoding the respective pathways were used in diagnostic PCRs. For both gene clusters, a large number of positive wells were detected among the filamentous plates (Fig. 2d and Extended Data Fig. 1c). Subsequent PCRs with eubacterial and ‘Entotheonella’-specific 16S ribosomal RNA gene primers showed that about half of the wells contained DNA originating from ‘Entotheonella’ phylotypes. Overall, from 48 wells of an analysed filamentous plate, 22 wells were positive for the onnamide, 34 for the polytheonamide, and 27 for the ‘Entotheonella’ sp. 16S rRNA gene, as confirmed by sequencing of each amplicon. Sixteen of the positive wells showed amplification for all three of the onn, poy and ‘Entotheonella’ sp. primer pairs in one or more out of three repetitive PCRs (Fig. 2d). For further analysis, wells positive for all three primer sets were subjected to PCR using eubacterial 16S rRNA gene primers. 16S rRNA genes from ‘Entotheonella’ sp. as well as Escherichia coli were identified. The E. coli amplicon was discarded, as it was also identified in MDA-treated wells that only contained water. Thus, the data suggested ‘Entotheonella’ as the source of both the onnamide-type compounds and polytheonamides.
Two chemically distinct ‘Entotheonella’ symbionts
As not all wells were positive for all three primer pairs and bacteria might have been overlooked owing to incomplete genome coverage during MDA16, we wished to validate further our results by metagenomic sequencing. On the filamentous bacterial cell sample, several rounds of Illumina, 454, PacBio, and Sanger sequencing were performed (Supplementary Table 1). Of the sequencing reads, 78.3% assembled to longer contigs, resulting in 18,093 contigs of at least 500 bp. The remaining reads did not show significant overlap, suggesting that the corresponding phylotypes were present only at low concentration. This hypothesis was backed by the observation of a high variance in coverage, ranging between 3.3- and 1,564.7-fold for contigs of at least 2 kb length. Basic Local Alignment Search Tool X (BLASTX) analysis of the contig and scaffold sequences followed by binning based on sequence depth and G + C content revealed two large populations of bacterial DNA with a G + C content around 55% (Supplementary Table 2). A third set of low coverage and low G + C contigs delivered hits against various eukaryotic genomes and was therefore excluded from further analyses (Extended Data Fig. 2a). A more detailed analysis of the filtered data set revealed for most bacterial genes the existence of two highly similar versions (approximately 85–91% nucleotide identity) that resided in virtually syntenous genomic environments encompassing over 4.5 Mb (Extended Data Fig. 3). The overlapping genomic regions included exactly two orthologues of 35 single-copy genes often used as bacterial phylogenic markers (Supplementary Table 3)17. These features suggested that the large majority of assembled bacterial sequences belonged to two closely related ‘Entotheonella’ variants, termed TSY1 and TSY2, with 97.6% identical 16S rRNA gene sequences and an average G + C content of 55.79% (Extended Data Fig. 2b). The identity of the 16S rRNA genes to that of ‘E. palauensis’ was about 97%. Depth analysis also suggested the presence of about 236 kb of DNA belonging to at least one large plasmid (G + C content: 55.11%). Coverage was 60.3-, 24.5- and 278.5-fold for the TSY1 and TSY2 chromosomes and the plasmid(s), respectively (corresponding to a ratio of 1:0.4:5), indicating that TSY1 is the dominant strain (Extended Data Fig. 2b). Both strains possess genomes of similar size that exceed 9 Mb, thus belonging to the largest known prokaryotic genomes (Supplementary Table 2). A remarkably large number of repetitive elements, some present in about 25 to 100 copies, as well as the high degree of similarity of the two genomes prohibited further assembly. To determine completeness of genomes, a core gene group analysis18 was performed, identifying 62 of 66 core groups for both TSY1 and TSY2. Thus we assume that the protein inventory of both strains was almost completely established.
The search for metabolic genes in this data set revealed complete sets of onn and poy genes on the plasmid-derived contigs. In addition, an unexpectedly high number of further gene clusters for polyketide and ribosomal or non-ribosomal peptide biosynthesis were identified on the chromosomal sequences. To allow for prediction of the corresponding metabolites, sequence gaps within most of these loci were filled by paired-end sequencing of 3- and 8-kb libraries and by combinatorial or targeted PCR, resulting in at least 28 biosynthetic gene clusters on 31 scaffolds (Extended Data Fig. 4 and Supplementary Table 4). For many non-ribosomal peptide synthetase (NRPS) clusters, bioinformatic predictions based on enzyme colinearity rules19, substrate recognition motifs20,21,22, and the presence of genes for non-proteinogenic amino acid biosynthesis (Supplementary Table 5), revealed known bioactive peptides from Japanese T. swinhoei as the best structural hits. Specifically, we identified virtually perfect matches for the cyclotheonamides, keramamides and nazumamide A. In addition, we identified a konbamide A-type23 cluster in which five of the six NRPS modules are present and colinear with the compound structure, but two ORF insertions disrupt the NRPS architecture, suggesting that the cluster is an inactive evolutionary relic. Consistent with this, members of the onnamide, polytheonamide, keramamide, and cyclotheonamide compound families were detected using high-resolution mass spectroscopy (HRMS) in extracts of our sponge specimens and enriched filamentous cell fractions, but we were unable to detect the konbamides (Supplementary Tables 6 and 7, and Extended Data Fig. 5). Taking together the combined bioinformatic and chemical analyses, candidate gene clusters existed for all known peptide and polyketide families including onnamides and polytheonamides, except for the aurantosides. In addition to these attributable genes, loci for at least 14 peptides of unknown identity were found (Extended Data Fig. 4). Notably, this also includes four further gene clusters for proteusins, a recently discovered new natural product family with polytheonamides as the only members known to date6,24. Tandom mass spectrometry (MS–MS)-based molecular networking25 suggested a high diversity of previously unknown metabolite families, indicating that at least some of these orphan pathways are likely to be active (Extended Data Fig. 6). The gene candidates for konbamides (kon), keramamides (ker), nazumamide A (naz), and an unknown non-ribosomal peptide formed a supercluster of 129 kb. The binning data suggested that this region, the putative cyclotheonamide (cth), and the two unassigned proteusin loci all belong to the chromosome of the dominant ‘Entotheonella’ sp. TSY1 (Extended Data Fig. 2b). The chromosome of TSY2 contained fewer (at least seven) metabolic gene clusters (two polyketide, at least two NRPS, and a further proteusin cluster) that could not be assigned to known compounds. Except for a small NRPS and a type III polyketide synthase (PKS) system present in both genomes, there was no overlap in the natural product gene repertoires of TSY1 and TSY2, indicating that significant chemical variation exists among members of ‘Entotheonella’, even within the same sponge individual. To validate further the source of the plasmid-based polytheonamide and onnamide genes, we conducted additional single-cell experiments (Fig. 2d). All MDA samples previously analysed positive for onn and poy genes were tested again with PCR primers for various genes of the kon, naz, cth, ker and one unknown proteusin pathway. For all cases, positive wells were identified, suggesting that TSY1 carries the plasmid and produces the entire set of metabolites.
Functional evidence for the identity of ‘Entotheonella’ gene clusters was obtained by biochemically characterizing gene products from several pathways. Two selected NRPS adenylation domains encoded within the putative cth and ker pathways were overproduced in E. coli and analysed using a γ-18O4-ATP pyrophosphate exchange assay26 to investigate their amino acid substrate specificity (Extended Data Fig. 7). For the cth NRPS, the adenylation domain of module 2 (CthA2) exhibited high selectivity for the rare amino acid 2,3-diaminopropionate (DAP), consistent with the cyclotheonamide structure (Extended Data Fig. 7). The incorporation of this building block is also supported by the presence of two genes in the cluster that encode homologues of SbnAB-type DAP synthases27. KerA5 showed greatest substrate specificity for Leu, in agreement with known keramamides and the bioinformatic prediction (Extended Data Fig. 7). Thus, taking the colinearity rule of NRPSs into account, the data support the proposed function of these gene clusters.
We also obtained functional support for a biosynthetic role of the unknown proteusin pathway TSY1_14 by co-expressing the putative nitrile-hydratase-like precursor peptide with a predicted lanthionine synthetase encoded directly adjacent to the precursor gene. Up to three dehydrations of the core peptide were observed by HRMS for the co-expression product compared to the unmodified peptide from expression of the precursor peptide alone. Subsequent alkylation of reduced cysteine residues and tandem MS–MS indicated for each dehydration, one lanthionine bridge was formed within the predicted core peptide (Extended Data Fig. 8 and Supplementary Table 8). These experiments demonstrated that the putative proteusin gene cluster TSY1_14 encodes a functional precursor peptide and modifying lanthionine synthetase. Considering the high complexity of the sponge microbiome, which contains hundreds of ribotypes, the accumulation of metabolic genes in two variants of ‘Entotheonella’ is remarkable. Owing to the extraordinary biosynthetic repertoire, we propose the name ‘Candidatus Entotheonella factor’ (latin, factor; the producer) for these bacteria.
‘Entotheonella’ species are ubiquitous
These findings raised the question whether ‘Entotheonella’ spp. also inhabit other sponges and could have a general role in natural product biosynthesis. It was previously shown that an enriched fraction of ‘E. palauensis’ from a Palauan chemotype of T. swinhoei contained high concentrations of the hybrid polyketide-peptide theopalauamide14,15. ‘Entotheonella’ members were also detected in another lithistid sponge, Discodermia dissoluta, that contains the anticancer polyketide discodermolide28. To analyse the distribution of ‘Entotheonella’ spp. in depth, 37 taxonomically diverse sponge species collected at 20 locations (Supplementary Table 9) were tested by PCR based on conserved, unique regions of ‘Entotheonella’ 16S rRNA genes. Of the 37 sponges, 28 yielded amplicons with sequences exhibiting 95.5–99.9% nucleotide identity to the homologues of ‘E. factor’ (Extended Data Fig. 9a, b). Thus, ‘Entotheonella’ spp. seem to be widely distributed in marine sponges from distant geographical regions. ‘Entotheonella’ amplicons were also obtained from various seawater samples; however, contamination from sponges growing nearby cannot be excluded. For further insights into the discovery potential and chemical variability of these bacteria, we initiated studies on another chemotype of T. swinhoei (type W1, referring to the white sponge interior) that contains the actin inhibitor misakinolide A (Extended Data Fig. 10b), a complex polyketide not present in the Y chemotype. PCR detection of PKS genes using the total sponge DNA generated exclusively amplicons that were phylogenetically attributed to the sup type (Extended Data Fig. 10c), a putative fatty acid synthase that is widespread and dominant in most sponge microbial consortia and not involved in the production of complex, bioactive polyketides29. In contrast, a highly enriched ‘Entotheonella’ fraction (Extended Data Fig. 10a) prepared from this sponge yielded a completely different set of amplicons consisting of six gene fragments all belonging to PKSs associated with complex polyketide production (Extended Data Fig. 10c). None of these had a close homologue in TSY1 or TSY2, thus further supporting a diverse chemistry of ‘Entotheonella’ phylotypes.
A new candidate phylum, ‘Tectomicrobia’
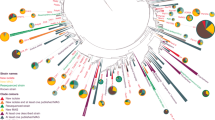
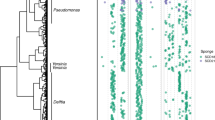
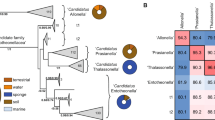
To obtain insights into the taxonomic position of ‘Entotheonella’, an initial 16S rRNA-based phylogenetic analysis was conducted (Extended Data Fig. 9c). Altogether, 243 16S rRNA gene sequences were analysed from marine sponges in this study and from public databases. As the 16S rRNA sequences were only 82% identical to representatives from known bacterial phyla and form a well-separated clade, we suggest the status of a new candidate phylum30. The name ‘Tectomicrobia’ (latin, tegere; to hide, to protect) was chosen to reflect their uncultured status as well as the capability to produce bioactive compounds that are likely used as chemical defence. The closest relatives to ‘Tectomicrobia’ are Nitrospina spp., which were recently proposed to belong to a new phylum, Nitrospinae31. The known sequences belonging to ‘Tectomicrobia’ comprise at least three discrete phylogenetic clades. The largest encompasses all ‘Entotheonella’ sequences sensu stricto, which were largely recovered from marine sponges but also seawater (138 sequences total, of which 107 sequences were produced in this study), a second clade includes related, non-‘Entotheonella’ 16S rRNA gene sequences from various marine sponges (36 sequences), and a third group contains 16S rRNA gene sequences from terrestrial soils (18 sequences). For further validation of the phylogenetic data, we calculated trees using up to 38 concatenated, universally conserved single-copy marker proteins17 of TSY1, TSY2, and 2,509 bacterial and archaeal taxa to determine the position of ‘Entotheonella’ in the tree of life. Recalculations with data sets from closely affiliated phyla (Fig. 3) supported ‘Entotheonella’ as belonging to a new sister phylum to Nitrospinae, in agreement with the 16S rRNA data.
RAxML inference of 991 taxa with 100 bootstrap iterations based on up to 38 marker genes. Sequences are collapsed on the phylum level and the number of collapsed sequences is shown for each clade. The two ‘Tectomicrobia’ variants TSY1 and TSY2 are highlighted in bold. Bootstrap support values of equal or greater than 70% are shown for each node. The scale bar represents 10% estimated sequence divergence. PV-1 and 3-11 are strain names; OP8 is the former name of the (then candidate) phylum Aminicenantes.
Conclusions
Owing to the high frequency of structurally distinct, bioactive metabolites in sponges, these animals have an important role in drug discovery. Compound localization studies suggested Bacteria as producers of individual metabolites14,15,32,33, but remained ambiguous owing to the possibility of sequestration or transport. The true source of sponge natural products has therefore been a long-standing and, with the exception of metagenomic data providing kingdom-level information5,6,34, unanswered question. Here we provide evidence that a single member of the highly diverse microbiome of T. swinhoei Y, ‘E. factor TSY1’, is the source of almost all polyketides and peptides that have been isolated from its sponge host. The bioinformatic assignment to known compounds is further supported by functional studies for polytheonamides6, onnamide-type compounds35,36, keramamides, cyclotheonamides and an orphan proteusin. Our data on TSY1, TSY2, and a highly enriched ‘Entotheonella’ preparation from a second T. swinhoei chemotype, indicate that members of this candidate genus contain producers with a rich and, so far, unique secondary metabolism. Reports on ‘Entotheonella’ spp. from two other chemically rich sponges15,28,37 and our detection of these bacteria in many additional species hint at their more widespread role in the chemistry of their hosts. This study adds the first uncultivated prokaryotes to the taxonomically limited canon of metabolically talented bacteria. ‘Entotheonella’ spp. exhibit interesting parallels to streptomycetes and some other well-known producer groups38,39,40,41,42; for example, expanded genome size, biosynthetic superclusters43 and multiple modular assembly lines, high metabolic variability among closely related organisms, and complex morphology. For ‘Entotheonella’ spp., complex morphology is particularly noteworthy, as it affords attractive opportunities to systematically study chemical interactions in marine symbioses and to exploit uncultivated bacteria in a targeted way for drug discovery.
Methods Summary
An adapted differential centrifugation protocol14 was used to sediment filamentous and unicellular bacteria from the sponge tissue. Single bacteria cells and filaments were sorted into micro-titre plates by flow cytometry with a BD FACSAria II cell sorter (BD Biosciences). Genomic DNA was amplified using an Illustra Genomiphi V2 DNA Amplification Kit (GE Healthcare) and subjected to PCR analysis. Sequence information was obtained using the GS-FLX (454) and MiSeq (Illumina) platforms, using whole-genome sequencing and long mate-pair libraries. Additional sequence reads were obtained by PacBio sequencing (GATC) and Sanger sequencing (IIT). Reads were assembled using the Newbler (v2.6) de novo assembler. Automated annotation was performed with Rapid Annotation and Subsystem Technology (RAST)44 and validated manually. PKS and NRPS domain architecture and substrate specificities were based on sequence alignments and prediction-based software22,45,46. Adenylation domains overexpressed in E. coli were characterized using a γ-18O4-ATP pyrophosphate exchange assay as previously described26. The TSY1_14 proteusin precursor peptide was overexpressed in E. coli with and without the putative modifying LanM-like lanthionine synthetase from the same gene cluster. The resulting peptide products were analysed by liquid chromatography (LC)–electrospray ionization (ESI)–HRMS after TCEP (tris-(2-carboxyethyl)-phosphine) treatment, tryptic digest and derivatization. Extracts of T. swinhoei and enriched ‘Entotheonella’ were analysed by ultra-performance liquid chromatography (UPLC) and nano-LC heated ESI (HESI)–HRMS followed by eMZed47 data analysis and molecular networking25.
Online Methods
Sponge collection
A list of analysed sponges and their collection sites is provided in Supplementary Table 6. Specimens were placed separately into plastic bags and brought to the surface. Immediately after collection, sponge tissues were cut into pieces and stored at −80 °C in a transportable liquid nitrogen freezer (Bahamas collection) or in 70% aqueous ethanol.
Isolation of bacteria
To prepare enriched bacterial fractions, a protocol adapted from a previous paper was used14. From freshly collected T. swinhoei, the thin red ectosome layer of a 500-g sponge piece was removed. The remaining portion was cut into smaller pieces, cleaned of other animals and processed in a National MJ-C28 juicer. Liquids and the solid residues were transferred into a 2-l graduated cylinder, and the volume was adjusted to 1.5 l with Ca- and Mg-free artificial sea water (CMF ASW)48. The mixture was incubated at room temperature for 15 min, while stirring every 2 min to dissociate sponge cells, then left undisturbed for additional 10 min to allow settling of particles. Supernatants were decanted into another graduated cylinder, and CMF ASW was added to the residues to a final volume of 1.5 l, followed by a second dissociation and settling period. The collected supernatants were passed through a 32-μm Nitex mesh and centrifuged at 1000g for 10 min to pellet filamentous cells. The supernatants were subsequently centrifuged at 4500g to pellet unicellular bacteria. Each bacterial fraction was washed once with 200 ml CMF ASW, pelleted again, resuspended in 200 ml CMF ASW and stored at 4 °C.
Flow-cytometric analysis and cell sorting
Prior to sorting, both the pellet and supernatant fraction were analysed by flow cytometry using the BD FACSAria II cell sorter (BD Bioscience). Size distributions of the bacteria within both the fractions were determined using size calibration beads (Life Technologies) based on the following sizes: 1 μm, 2 μm, 4 μm, 6 μm, 10 μm and 15 μm (Life Technologies). Samples used for sorting were diluted accordingly and size distribution was analysed at 500–1,000 events per sec. Flow-cytometry results were analysed using the FACSDiva software (BD Bioscience). Sterile 96-well plates, containing 1 μl of nuclease-free ultrapure water in each well were prepared and single cells or multicellular filaments were sorted accordingly. Confirmation of the successful cell sorting was conducted by observation of each drop under a fluorescence microscope. The resulting 96-well plates were stored at 4 °C for subsequent whole-genome amplification (WGA).
Whole-genome amplification
Single isolated bacterial cells or filaments attained from the cell sorting were disrupted by heat treatment at 95 °C, 3 min, and WGA was conducted based on the phi29 polymerase-mediated multiple displacement amplification (MDA) technique using the Illustra Genomiphi V2 DNA Amplification Kit (GE Healthcare). Each WGA reaction per well was optimized and conducted at 10-μl volumes as recommended by the manufacturer. Upon MDA, each well was diluted tenfold with nuclease-free ultrapure water before storage.
Detection of biosynthetic and 16S rRNA genes
For the detection of 16S rRNA genes and the biosynthetic gene clusters (onn, poy, kon, naz, cth, ker and pts), nested PCR was performed against wells containing MDA amplified genomic DNA from single-filament bacteria. PCR was conducted using the high-fidelity PrimeSTAR Max DNA Polymerase (TaKaRa) (first amplification) and the AmpliTaq Gold 360 Master Mix (Applied biosystems) (second amplification) based on the following conditions. First amplification: 98 °C, 5 min (initial denaturation); 98 °C, 10 s; 54 °C, 15 s; 72 °C, 1.5 min (for 35 cycles); 72 °C, 7 min (final extension). Second amplification: 95 °C, 10 min (initial denaturation); 98 °C, 30 s; 59 °C, 30 s; 72 °C, 30 s (for 30 cycles); 72 °C, 7 min (final extension). For the first amplification, 1 μl of template from the tenfold-diluted MDA samples was used, and 0.5 μl was used directly from the first PCR amplification product for the second amplification. All PCRs were conducted at 25 μl final volume. The primer sets used in the nested PCRs are summarized in Supplementary Table 10. Amplicon sizes were determined by agarose gel electrophoresis and confirmed by Sanger sequencing. Nested PCR was performed in triplicate for the ‘Entotheonella’ 16S rRNA gene, onn and poy gene clusters, each on different days. For the other biosynthetic clusters, nested PCR was only conducted once. To test for the presence of contaminating bacteria, the 16S rRNA gene was amplified using the 16SU27F and 16SU1492R primers from MDA-amplified wells containing or not containing single-filament bacteria, cloned, and 20 clones were randomly selected for sequencing.
Genome sequencing and annotation
Sequencing was performed on two platforms: first, a GS-FLX (454) using a whole-genome shotgun and a 3-kb-long paired-end library, both prepared according to the manufacturer’s instructions and sequenced using the Titanium chemistry; second, a MiSeq (Illumina) using a Nextera (Epicentre Biotechnologies) whole-genome shotgun paired-end and a 8-kb mate-pair library. The latter was prepared using a hybrid protocol by replacing the 454 adapters with Illumina paired-end adapters in the final steps of an 8-kb-long paired-end library construction. The resulting library fragments were size selected by gel electrophoresis and excision to a size of 400 bp. The library was sequenced in a 2× 151-bp paired-end run, and a 2× 251-bp run was performed for the 8-kb mate-pair library. Prior to assembly, the read pairs of the 8-kb mate-pair library were joined, excluding all reads without a perfect match in the overlapping region. The joined pairs were split at the 454 long paired end linker, excluding all reads without a perfect match and the two resulting reads were reverse complemented to simulate large insert Sanger reads. In addition, sequence information was obtained by PacBio sequencing (GATC) and Sanger sequencing (IIT). All reads were then assembled using the Newbler (v2.6) de novo assembler, non-GS-FLX reads were provided in FASTQ format, assembly was performed with default parameters except using 30 bp as minimum overlap match. Contig numbers for the TSY1 and the TSY2 genome are 1,774 and 3,270, respectively. Synteny analysis was performed with r2cat49. Automated annotation was performed with Rapid Annotation and Subsystem Technology (RAST)44. Manual identification and annotation of natural product biosynthetic gene clusters were based on similarity searches (BLAST) with validated biosynthetic genes as queries. Additional automated identification of natural product gene clusters was performed with antibiotics and Secondary Metabolite Analysis SHell (antiSMASH)45. Preliminary PKS and NRPS domain architecture and adenylation domain specificities were determined using freely available software22,45,46. All manual annotation and routine bioinformatic analysis was performed using Geneious version 6.0.6 created by Biomatters (available from http://www.geneious.com). Scaffold gaps were closed using PCR amplification with Phusion High-Fidelity or LongAmp Taq DNA polymerase (New England Biolabs) and sequencing (GATC Biotech).
PCR detection of ‘Entotheonella’ spp
DNA was isolated from sponge samples with the Fast DNA spin kit for soil (Q-Biogene) according to the manufacturer’s protocol. PCR amplification was performed with the eubacterial 16S rRNA gene specific primers 16SU27F and 16SU1492R as published previously50. The resulting PCR product was used as template in a following nested PCR using two different procedures. First, newly designed ‘Entotheonella’ 16S rRNA gene-specific primers Ento271F and Ento1290R. Conditions: 1 µl DMSO was added to 50 µl of PCR mix. An initial denaturing step for 2 min at 95 °C followed by 35 cycles of a denaturing step at 95 °C for 30 s, primer annealing at 63 °C for 30 s and elongation at 72 °C for 1 min. The program was completed with a final elongation step at 72 °C for 5 min. DreamTaq Polymerase (Fermentas) was used. Second, ‘Entotheonella’-specific primers Ento238F (ref. 15) and Ento1442R. Conditions: an initial denaturing step for 2 min at 98 °C followed by 35 cycles of a denaturing step at 95 °C for 10 s, primer annealing at 55 °C for 30 s and elongation at 72 °C for 1.5 min. TaKaRa Ex Taq polymerase was used. Different primer pairs were used to increase the diversity of detectable ‘Entotheonella’ 16S rRNA genes. The PCR products were purified, ligated into pGEM-T (Promega) and transformed into heat-competent E. coli Novablue cells. Using the vector primers SP6 and T7, colony PCRs on 20 clones of each sponge were performed. Double restriction digestion of these PCR products was performed using the enzymes HaeIII and MspI, and the insert of one representative per RFLP (restriction fragment length polymorphism) pattern was sequenced.
Detection and analysis of PKS genes in the misakinolide chemotype of T. swinhoei
Metagenomic DNA was prepared from Theonella swinhoei W1 as described previously34. Crude DNA was purified further by electrophoresis on low-melting-point agarose followed by gel extraction using the peqGOLD Gel Extraction Kit (PEQLAB). The filamentous cell fraction was prepared from T. swinhoei W1 as described above. Ketosynthase fragments were amplified from total sponge DNA and filamentous cells using the primers KSDPQQF and KSHGTGR51. Approximately 0.4 µl of the purified sponge metagenomic DNA or 0.5 µl of the rinsed cell pellet suspension was used as PCR template in a 25 µl PCR mixture that also contained 2.5 µl of 10× thermopol buffer (New England Biolabs), 0.5 µl of 10 mM dNTPs, 1 µl of 25 mM MgCl2, 2 µl each of 50 mM KSDPQQF and KSHGTGR primer, and 0.25 µl High Fidelity Hot Start Polymerase (Jena Bioscience GmbH) or 0.125 µl Taq DNA Polymerase High Fidelity (Invitrogen). The thermal cycle program was set up at 35 cycles and consisted of the following steps: lid heating at 105 °C, predenaturation at 95 °C for 2 min, denaturation at 95 °C for 1 min, annealing at different temperatures (55, 58, 61 °C), elongation at 74 °C for 1 min, and final elongation at 74 °C for 10 min. Subsequently the PCR products with the desired size (approximately 700 bp) were gel-purified and ligated into pBlueScript II (SK-) (Stratagene). Plasmids harbouring 700-bp inserts were sequenced.
Phylogenetic analysis
16S rRNA gene sequences and reference sequences identified by initial BLAST52 searches were automatically aligned to a SILVA reference alignment using the SINA Webaligner53 and merged into the SILVA version 106 database54,55. The alignment was then manually refined. A maximum parsimony tree with 1,000 bootstrap resamplings and a maximum likelihood tree with 100 bootstrap resamplings were calculated in ARB using long (≥1,200 bp) sequences only. Short sequences were added using the parsimony interactive tool in ARB without changing the tree topology.
For the whole-genome trees, both ‘Entotheonella’ variants TSY1 and TSY2 were scanned for homologues of a set of 38 universally conserved single-copy proteins present in Bacteria and Archaea17. The assemblies were translated into all six reading frames, and marker genes were detected and aligned with hmmsearch and hmmalign included in the HMMER3 package56 using HMM profiles obtained from phylosift (http://phylosift.wordpress.com/). Extracted marker protein sequences were used to build concatenated alignments of up 38 markers per genome. Phylogenetic inference methods used were the maximum likelihood based FastTree2 (ref. 57) and a custom RAxML bootstrap script originally provided by Christian Goll and Alexandros Stamatakis (Scientific Computing Group, Heidelberg Institute for Theoretical Studies) and modified by D. Jacobsen. The script requires two input files, the alignment file as PHYLIP format and a starting tree calculated by RAxML-Light58. The script workflow can be briefly summarized as follows. First RAxML version 7.3.5 (ref. 59) creates bootstrap replicates of the multiple sequence alignments and stepwise addition order parsimony trees as starting points for the maximum likelihood search, based on user defined rate heterogeneity and substitution models. Next RAxML-Light58 is run on every bootstrap replicate. After all RAxML-Light runs are finished the resulting replicate trees are fed into RAxML to calculate the bootstrap support values which are drawn upon the starting tree. The major advantage of this approach over simply running RAxML to sequentially perform the bootstrapping calculation is that multiple RAxML-Light instances can be used to evaluate several bootstrap replicates in parallel. In addition, the RAxML-Light implementation is both faster and more efficient than RAxML for large-scale phylogenetic inferences since it was specifically developed for use in high performance computing environments and has an efficient checkpointing and restart capability58. The rate heterogeneity and amino acid evolution models used were GAMMA and Le-Gascuel (LG) for the custom RAxML bootstrap script, and CAT approximation with 20 rate categories and Jones–Taylor–Thornton (JJT) for FastTree2. We first generated a phylogenetic tree with FastTree2 including 2,509 bacterial and archaeal taxa to verify the position of the ‘Entotheonella’ variants in the tree of life. Next we calculated phylogenies (Fasttree2, RAxML script) using a reduced data set of 991 taxa of closely affiliated phyla, including Proteobacteria, Acidobacteria, Nitrospirae, Deferribacteres, Chrysiogenetes, and the recently proposed phyla Aminincenantes and Nitrospinae17,31.
Adenylation-domain assays
Adenylation-domain regions of the ker (KerA5) and cth (CthA2) NRPS (non-ribosomal peptide synthetase) genes were PCR amplified from genomic DNA isolated from the T. swinhoei Y filamentous fraction using Phusion High-Fidelity DNA polymerase (New England Biolabs). Primers kerA5F and kerA5R were used for domain KerA5 and primers cthA2F and cthA2R for domain CthA2. The PCR products were ligated into the vectors pGEM-T Easy or pBluescript SK(+) and transformed into E. coli DH5α for subcloning into pET28b. Based on recent findings that many A domains are only active in vitro when coexpressed with a MbtH-like protein60,61, each of the His-tagged adenylation domains were coexpressed with KerL, a MbtH-like protein from the ker pathway. KerL was cloned into the co-expression vector pCDF-DUET. For overexpression of KerA5, 1-l cultures of E. coli BL21(DE3) containing the expression plasmids were grown in Terrific Broth medium with kanamycin (50 μg ml−1) and streptomycin (50 μg ml−1) selection to a D600 nm of 1.8 at 37 °C and 250 r.p.m. The cultures were then cooled to 16 °C before being induced with IPTG (Isopropyl β-D-1-thiogalactopyranoside; 1 mM) and grown for 20 h at 16 °C and 250 r.p.m. Overexpression of soluble CthA2 was greatly improved by transforming the expression plasmids into E. coli Bl21-A1 and growing the clones as above except a final concentration of 0.2% arabinose was added at when the D600 nm reached 0.4. The protein-purification and mass-exchange-based adenylation assays were performed as reported previously26,62,63.
UPLC high-resolution HESI–MS analysis of sponge and bacterial fractions
Whole sponge samples of Theonella swinhoei and enriched ‘Entotheonella’ cell fractions were extracted with ethanol, methanol, propanol and acetone. Whole sponge extracts and ‘Entotheonella’ extracts were subjected to ultra-performance liquid chromatography (UPLC) heated electrospray ionization (HESI)–high-resolution mass spectrometry (HRMS) and nano-LC HESI–HRMS analysis, followed by eMZed47 data analysis. HESI–HRMS data were collected on a Thermo Q Exactive coupled to a Dionex Ultimate 3000 UPLC system. For the standard analysis of ‘Entotheonella’ and Theonella extracts, solvent gradients (A = H20 + 0.1% formic acid, and B = acetonitrile + 0.1% formic acid with B at 5% for 0–2 min, 5–95% for 2–14 min and 95% for 14–17 min at a flowrate of 0.5 ml min−1) were used on a Phenomenex Kinetex 2.6 µm C18 100 Å (150 × 4.6 mm) column at 27 °C (A) to 30 °C (B). The MS was operated in positive ionization mode at a scan range of (A) 200–2,500 m/z (mass-to-charge ratio) or (B) 100–1,600 m/z, respectively, and a resolution of 70,000 or 140,000 at m/z 200. The spray voltage was set to 3.7 kV and the capillary temperature to 320 °C. For the identification of polytheonamides with the UPLC setup an isocratic elution with 45% n-propanol at a flow rate of 0.5 ml min−1 was used on a Phenomenex Kinetex 2.6 µm C18 100 Å (150 × 4.6 mm) column at 45 °C. The MS was operated in positive ionization mode at a scan range from 1,000–6,000 m/z and a resolution of 140,000 at m/z 200, the spray voltage was set to 3.7 kV and capillary temperature to 320 °C. For the detection of heterologously produced and TCEP (tris-(2-carboxyethyl)-phosphine)-treated proteusins, a solvent gradient (A = H20 + 0.1% formic acid, and B = acetonitrile + 0.1% formic acid with B at 5% for 0–2 min, 5–95% for 2–14 min and 95% for 14–17 min at a flowrate of 0.5 ml min−1) was used on a Phenomenex Aeris WIDEPORE 3.6 µm C4 200 Å column (50 × 2.1 mm) at 27 °C. The MS was operated in positive ionization mode at a scan range from 600–2,000 m/z and a resolution of 70,000 at m/z 200, the spray voltage was set to 3.7 kV and capillary temperature to 320 °C. MS experiments for iodoacetamide treated samples were adjusted to a scan range from 150–2,000 m/z. The Manual Xtract function of Thermo Protein Deconvolution software version 2.0.53.5 was used to obtain protein masses from the spectra, using charge states 5–50 (1–50 in tryptic digest) for mass calculation, thresholds were set to 3 for signal to noise, 3 for the minimum number of detected charge states (2 in tryptic digest), 0% for relative abundance, 25% for overlap remainder, and 1 for minimum intensity. The fit factor for isotopic pattern comparison was set to 80% and the expected intensity error to 3. MS–MS experiments on tryptic digested samples were carried out for a mass range of 400-1800 m/z at a stepwise normalized collision energy (NCE) of 24.5, 35, and 45.5. Targeted MS–MS was applied on the [M+H] ions [1051.8]3+, [1057.8]3+, [1076.54]3+, [1105.73]4+, [1110.0]4+, [1129.0]4+, [1148.0]4+, [1474.0]3+, [1480.0]3+, [1505.0]3+ and [1530.0]3+ within an isolation window of 2 m/z at a resolution of 70,000 at m/z 200. Peptide masses for MS–MS fragments were manually calculated from the observed [M+H]+ ions ([M+2H]2+ for the y37 ion). In addition, a Thermo Easy-nLC 1000 equipped with a Acclaim PepMap RSLC nano Viper column (2 µm, C18, 100 Å (15 cm × 50 µm) coupled to a Thermo Q Exactive was used increase detection sensitivity for trace compounds. The nano-LC was operated using the following solvent gradient: A = H2O + 0.1% formic acid, and B = 1-propanol + 0.1% formic acid with B at 10–80% for 0–60 min and 100% for 61–95 min at a flowrate of 0.3 ml min−1. For the identification of secondary metabolites produced by ‘Entotheonella’, as indicated by the presence of the respective gene cluster, Thermo raw files were converted into mzXML files using MSExport and further processed with the Python-based, open-source eMZed framework47. The processed data were compared to a list of known Theonella compounds and purified onnamide, cyclotheonamide and polytheonamide standards. To generate a mass-spectral molecular network25, combined data sets of data dependent nano-LC and UPLC high-resolution HESI–MS–MS experiments were used. The chromatographic separation was conducted using the aforementioned UPLC and nano-LC conditions. The top 10 most intense ions of each parent ion scan were subsequently fragmented with a resolution of 17,500 at m/z 200 and a normalized collision energy (NCE) of 35. The isolation width was set to 5 Da, the dynamic exclusion to 15 s, and the default charge state to 4 to enable fragmentation of high molecular mass secondary metabolites (for example, polytheonamides). Identical spectra were merged to form consensus spectra using MSCluster64. Cosine values were calculated for every possible pair of spectra (spectral alignment) and spectra obtained from solvent controls were removed using MATLAB scripts25,65. The cosine value threshold was set to 0.5, whereas a cosine value of one indicates identical spectra and a cosine value of 0 displays no spectral correlation. The resulting network was visualized in cytoscape66. Consensus spectra were annotated by comparison with a database of published secondary metabolites from T. swinhoei.
Functional studies on the putative proteusin gene cluster TSY1_14
The putative nitrile hydratase-like precursor gene from gene cluster TSY1_14 was PCR-amplified from genomic DNA from the T. swinhoei Y filamentous fraction using Phusion High-Fidelity DNA polymerase (New England Biolabs) and cloned into pET28b (Merck) using primers Prec-TSY1_14-F and Prec-TSY1_14-R (Supplementary Table 10). The resulting construct was digested with NcoI/HindIII and ligated into pETDuetI (Merck), producing the N-His-tagged precursor peptide expression construct pMH124. The putative LanM-like lanthionine synthetase gene from TSY1_14 was amplified using primers Lanth-TSY1_14-F and Lanth-TSY1_14-R and cloned into pCDFDuetI to obtain the untagged modifying enzyme expression construct pMH104. For functional analysis, pMH124 was transformed into E. coli BL21(DE3) alone and co-transformed with pMH104. Expression cultures (100 ml) of Terrific Broth were inoculated with 1 ml of overnight culture and incubated at 16 °C at 250 r.p.m. for 5 days after induction with IPTG. Cells were collected by centrifugation (3,220g for 10 min) and pellets frozen in liquid nitrogen. The cells were lysed by sonication in lysis buffer (50 mM potassium phosphate, pH 8.0, 300 mM NaCl, 20 mM imidazole, 10% (v/v) glycerol) and the supernatant was incubated with 300 µl Protino Ni-NTA resin (Macherey Nagel) for 1 h at 4 °C on a rocking platform. Resin was pelleted at 850g (20 min, 4°C) and applied to a Poly-Prep Chromatography column (Bio-Rad), washed with 5 ml lysis buffer, 5 ml wash buffer (50 mM potassium phosphate, pH 8.0, 300 mM NaCl, 40 mM imidazole, 10% (v/v) glycerol), and eluted with three 500-µl fractions of elution buffer (50 mM potassium phosphate, pH 8.0, 300 mM NaCl, 250 mM imidazole, 10% (v/v) glycerol). The elution fractions were adjusted to a protein concentration of 152 µM (2 µg µl−1 as measured by Roti-Nanoquant modified Bradford assay (Carl Roth)) and Tris(2-carboxyethyl)phosphine (TCEP) hydrochloride (Sigma Life Science, BioUltra) was added in 100-fold molar excess to reduce disulphide bond formation. After incubation at 25 °C for 30 min, the samples were subjected to UPLC HESI–HRMS analysis as described above. To determine the number of free thiols and putative lanthionine bridges the control and modified peptides were derivatized with iodoacetamide. In brief, the peptides were desalted (Vivaspin, 5 kDa MWCO, Sartorius) with 10 volumes of 100 mM ammonium bicarbonate (pH 7.86) and adjusted to 1 µg µl−1 (76 µM). After treatment with 9 mM TCEP and 0.09% SDS (25 min at 25 °C), the samples were treated with 16 mM Iodoacetamide (Sigma Life Science, BioUltra) in the absence of light for 30 min at 25 °C and analysed by UPLC HESI-HRMS. Sequencing grade trypsin (Promega) was added to the remainder at a trypsin:protein ratio of 1:25 and incubated for 2 h at 37 °C before HRMS analysis.
Accession codes
Accessions
GenBank/EMBL/DDBJ
Data deposits
Sequence data for 16S rRNA have been deposited in GenBank under accession numbers KF926701–KF926822. Sequence data for Whole Genome Shotgun projects have been deposited at DDBJ/EMBL/GenBank under the accession AZHW00000000 and AZHX00000000. The versions described in this paper are AZHW01000000 and AZHX01000000, respectively.
References
Berdy, J. Bioactive microbial metabolites — a personal view. J. Antibiot. 58, 1–26 (2005)
Achtman, M. & Wagner, M. Microbial diversity and the genetic nature of microbial species. Nature Rev. Microbiol. 6, 431–440 (2008)
Brady, S. F., Simmons, L., Kim, J. H. & Schmidt, E. W. Metagenomic approaches to natural products from free-living and symbiotic organisms. Nat. Prod. Rep. 26, 1488–1503 (2009)
Piel, J. Approaches to capturing and designing biologically active small molecules produced by uncultured microbes. Annu. Rev. Microbiol. 65, 431–453 (2011)
Piel, J. et al. Antitumor polyketide biosynthesis by an uncultivated bacterial symbiont of the marine sponge Theonella swinhoei . Proc. Natl. Acad. Sci. USA 101, 16222–16227 (2004)
Freeman, M. F. et al. Metagenome mining reveals polytheonamides as posttranslationally modified ribosomal peptides. Science 338, 387–390 (2012)
Hentschel, U. et al. Molecular evidence for a uniform microbial community in sponges from different oceans. Appl. Environ. Microbiol. 68, 4431–4440 (2002)
Taylor, M. W., Radax, R., Steger, D. & Wagner, M. Sponge-associated microorganisms: evolution, ecology, and biotechnological potential. Microbiol. Mol. Biol. Rev. 71, 295–347 (2007)
Hentschel, U., Piel, J., Degnan, S. M. & Taylor, M. W. Genomic insights into the marine sponge microbiome. Nature Rev. Microbiol. 10, 641–654 (2012)
Fusetani, N. & Matsunaga, S. Bioactive sponge peptides. Chem. Rev. 93, 1793–1806 (1993)
Binga, E. K., Lasken, R. S. & Neufeld, J. D. Something from (almost) nothing: the impact of multiple displacement amplification on microbial ecology. ISME J. 2, 233–241 (2008)
Siegl, A. et al. Single-cell genomics reveals the lifestyle of Poribacteria, a candidate phylum symbiotically associated with marine sponges. ISME J. 5, 61–70 (2011)
Grindberg, R. V. et al. Single cell genome amplification accelerates identification of the apratoxin biosynthetic pathway from a complex microbial assemblage. PLoS ONE 6, e18565 (2011)
Bewley, C. A., Holland, N. D. & Faulkner, D. J. Two classes of metabolites from Theonella swinhoei are localized in distinct populations of bacterial symbionts. Experientia 52, 716–722 (1996)
Schmidt, E. W., Obraztsova, A. Y., Davidson, S. K., Faulkner, D. J. & Haygood, M. G. Identification of the antifungal peptide-containing symbiont of the marine sponge Theonella swinhoei as a novel δ-proteobacterium, “Candidatus Entotheonella palauensis”. Mar. Biol. 136, 969–977 (2000)
Raghunathan, A. et al. Genomic DNA amplification from a single bacterium. Appl. Environ. Microbiol. 71, 3342–3347 (2005)
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013)
Chitsaz, H. et al. Efficient de novo assembly of single-cell bacterial genomes from short-read data sets. Nature Biotechnol. 29, 915–921 (2011)
Fischbach, M. A. & Walsh, C. T. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: Logic, machinery, and mechanisms. Chem. Rev. 106, 3468–3496 (2006)
Stachelhaus, T., Mootz, H. D. & Marahiel, M. A. The specificity-conferring code of adenylation domains in nonribosomal peptide synthetases. Chem. Biol. 6, 493–505 (1999)
Challis, G. L., Ravel, J. & Townsend, C. A. Predictive, structure-based model of amino acid recognition by nonribosomal peptide synthetase adenylation domains. Chem. Biol. 7, 211–224 (2000)
Rottig, M. et al. NRPSpredictor2—a web server for predicting NRPS adenylation domain specificity. Nucleic Acids Res. 39, W362–W367 (2011)
Kobayashi, J. & i et al. Konbamide, a novel peptide with calmoduiin antagonistic activity from the Okinawan marine sponge Theonella sp. J. Chem. Soc. Chem. Commun. 1050–1052 (1991)
Haft, D. H., Basu, M. K. & Mitchell, D. A. Expansion of ribosomally produced natural products: a nitrile hydratase-and Nif11-related precursor family. BMC Biol. 8, 70 (2010)
Watrous, J. et al. Mass spectral molecular networking of living microbial colonies. Proc. Natl. Acad. Sci. USA 109, E1743–E1752 (2012)
Phelan, V. V., Du, Y., McLean, J. A. & Bachmann, B. O. Adenylation enzyme characterization using γ-18O4-ATP pyrophosphate exchange. Chem. Biol. 16, 473–478 (2009)
Beasley, F. C., Cheung, J. & Heinrichs, D. E. Mutation of l-2,3-diaminopropionic acid synthase genes blocks staphyloferrin B synthesis in Staphylococcus aureus . BMC Microbiol. 11, 199 (2011)
Bruck, W. M., Sennett, S. H., Pomponi, S. A., Willenz, P. & McCarthy, P. J. Identification of the bacterial symbiont Entotheonella sp. in the mesohyl of the marine sponge Discodermia sp. ISME J. 2, 335–339 (2008)
Hochmuth, T. et al. Linking chemical and microbial diversity in marine sponges: possible role for poribacteria as producers of methyl-branched fatty acids. ChemBioChem 11, 2572–2578 (2010)
Hugenholtz, P., Goebel, B. M. & Pace, N. R. Impact of culture-independent studies on the emerging phylogenetic view of bacterial diversity. J. Bacteriol. 180, 4765–4774 (1998)
Lucker, S., Nowka, B., Rattei, T., Spieck, E. & Daims, H. The genome of Nitrospina gracilis illuminates the metabolism and evolution of the major marine nitrite oxidizer. Front. Microbiol. 4, 27 (2013)
Unson, M. D., Holland, N. D. & Faulkner, D. J. A brominated secondary metabolite synthesized by the cyanobacterial symbiont of a marine sponge and accumulation of the crystalline metabolite in the sponge tissue. Mar. Biol. 119, 1–11 (1994)
Flowers, A. E., Garson, M. J., Webb, R. I., Dumdei, E. J. & Charan, R. D. Cellular origin of chlorinated diketopiperazines in the dictyoceratid sponge Dysidea herbacea (Keller). Cell Tissue Res. 292, 597–607 (1998)
Fisch, K. M. et al. Polyketide assembly lines of uncultivated sponge symbionts from structure-based gene targeting. Nature Chem. Biol. 5, 494–501 (2009)
Zimmermann, K., Engeser, M., Blunt, J. W., Munro, M. H. & Piel, J. Pederin-type pathways of uncultivated bacterial symbionts: analysis of O-methyltransferases and generation of a biosynthetic hybrid. J. Am. Chem. Soc. 131, 2780–2781 (2009)
Pöplau, P., Frank, S., Morinaka, B. I. & Piel, J. An enzymatic domain for cyclic ether formation in complex polyketides. Angew. Chem. Int. Ed. 52, 13215–13218 (2013)
Schirmer, A. et al. Metagenomic analysis reveals diverse polyketide synthase gene clusters in microorganisms associated with the marine sponge Discodermia dissoluta . Appl. Environ. Microbiol. 71, 4840–4849 (2005)
Bentley, S. D. et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417, 141–147 (2002)
Omura, S. et al. Genome sequence of an industrial microorganism Streptomyces avermitilis: deducing the ability of producing secondary metabolites. Proc. Natl. Acad. Sci. USA 98, 12215–12220 (2001)
Schneiker, S. et al. Complete genome sequence of the myxobacterium Sorangium cellulosum . Nature Biotechnol. 25, 1281–1289 (2007)
Frangeul, L. et al. Highly plastic genome of Microcystis aeruginosa PCC 7806, a ubiquitous toxic freshwater cyanobacterium. BMC Genomics 9, 274 (2008)
Flores, E. & Herrero, A. Compartmentalized function through cell differentiation in filamentous cyanobacteria. Nature Rev. Microbiol. 8, 39–50 (2010)
Mast, Y. et al. Characterization of the 'pristinamycin supercluster' of Streptomyces pristinaespiralis . Microb. Biotechnol. 4, 192–206 (2011)
Aziz, R. K. et al. The RAST server: rapid annotations using subsystems technology. BMC Genomics 9, 75–89 (2008)
Medema, M. H. et al. antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. 39, W339–W346 (2011)
Bachmann, B. O. & Ravel, J. Chapter 8. Methods for in silico prediction of microbial polyketide and nonribosomal peptide biosynthetic pathways from DNA sequence data. Methods Enzymol. 458, 181–217 (2009)
Kiefer, P., Schmitt, U. & Vorholt, J. A. eMZed: an open source framework in Python for rapid and interactive development of LC/MS data analysis workflows. Bioinformatics 29, 963–964 (2013)
Spiegel, M. & Rubinstein, N. A. Synthesis of RNA by dissociated cells of the sea urchin embryo. Exp. Cell Res. 70, 423–430 (1972)
Husemann, P. & Stoye, J. r2cat: synteny plots and comparative assembly. Bioinformatics 26, 570–571 (2010)
Hentschel, U. et al. Isolation and phylogenetic analysis of bacteria with antimicrobial activities from the Mediterranean sponges Aplysina aerophoba and Aplysina cavernicola . FEMS Microbiol. Ecol. 35, 305–312 (2001)
Piel, J. A polyketide synthase-peptide synthetase gene cluster from an uncultured bacterial symbiont of Paederus beetles. Proc. Natl. Acad. Sci. USA 99, 14002–14007 (2002)
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990)
Pruesse, E., Peplies, J. & Glockner, F. O. SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28, 1823–1829 (2012)
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371 (2004)
Pruesse, E. et al. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 35, 7188–7196 (2007)
Eddy, S. R. Accelerated profile HMM searches. PLOS Comput. Biol. 7, e1002195 (2011)
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010)
Stamatakis, A. et al. RAxML-Light: a tool for computing terabyte phylogenies. Bioinformatics 28, 2064–2066 (2012)
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006)
Herbst, D. A., Boll, B., Zocher, G., Stehle, T. & Heide, L. Structural basis of the interaction of MbtH-like proteins, putative regulators of nonribosomal peptide biosynthesis, with adenylating enzymes. J. Biol. Chem. 288, 1991–2003 (2013)
Davidsen, J. M., Bartley, D. M. & Townsend, C. A. Non-ribosomal propeptide precursor in nocardicin A biosynthesis predicted from adenylation domain specificity dependent on the MbtH family protein NocI. J. Am. Chem. Soc. 135, 1749–1759 (2013)
Hofer, I. et al. Insights into the biosynthesis of hormaomycin, an exceptionally complex bacterial signaling metabolite. Chem. Biol. 18, 381–391 (2011)
Crüsemann, M., Kohlhaas, C. & Piel, J. Evolution-guided engineering of nonribosomal peptide synthetase adenylation domains. Chemical Sci. 4, 1041–1045 (2013)
Frank, A. M. et al. Clustering millions of tandem mass spectra. J. Proteome Res. 7, 113–122 (2008)
Bandeira, N., Tsur, D., Frank, A. & Pevzner, P. A. Protein identification by spectral networks analysis. Proc. Natl. Acad. Sci. USA 104, 6140–6145 (2007)
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003)
Acknowledgements
We thank R. Lasken and M. F. Freeman for discussion, R. W. M. van Soest, P. R. Bergquist, and Y. Ise for taxonomic identification of sponges, P. Kiefer for eMZed support, P. Dorrestein and J. Watrous for mass spectrometry networking support, A. Semeniuk and R. Meoded for experimental support, and T. Ravasi, P. Crews, Y. Kashman and M. Aknin for providing sponge specimens. This work was supported by the SNF (31003A_146992) to J.P., BMBF (GenBioCom: 0315581I to J.P. and 0315585J to J.K.), DFG (PI 430/1-3 and PI 430/9-1 to J.P., SFB 630-TP A5 to U.H.), the EU (BlueGenics to J.P.), MIWFT within the BIO.NRW initiative (280371902 to C. Rückert), the Grants-in-aid for Young Scientists (B), KAKENHI (23760755 to T.M.), JSPS to J.P., S.M. and H.T., Alexander von Humboldt Foundation to M.C.W., German National Academic Foundation to M.J.H. and E.J.N.H., and DAAD to A.R.U. The work conducted by the US Department of Energy Joint Genome Institute is supported by the Office of Science of the US Department of Energy under Contract no. DE-AC02-05CH11231.
Author information
Authors and Affiliations
Contributions
J.P., H.T., T.M. and J.K. conceived the study. S.M., T.W., K.T., I.A. and U.H. collected the sponges and determined the chemotypes. J.P. performed the cell separation and differential centrifugation, C.G. and A.R.U. isolated the DNA, T.M. and E.J.N.H. conducted the single-cell studies. C.Rü. and J.K. sequenced the metagenome, M.C.W., C. Rü., J.P., U.A.E.S., N.H., C.G., A.R.U. and M.J.H. analysed genomic data, K.T. and C.G. performed the distribution studies. M.C., M.K., and M.C.W. performed the adenylation domain assays, M.J.H. performed the proteusin studies, A.R.U. performed the studies on the misakinolide chemotype, C. Ri. and S.S. performed the phylogenetic analysis, and A.O.B. and E.J.N.H. performed HRMS experiments. All authors planned the experiments, analysed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Cell sorting of the sponge microbiome and single-cell or filament analysis.
a, Flow-cytometric analysis of the unicellular (left) and the filamentous (‘pellet’, right) fraction after differential centrifugation. Cell distribution was determined based on size gates that were set using calibration beads ranging from 1 μm to 15 μm in size. b, Whole-genome amplification (WGA) of single wells containing single-sorted cells (n = 48). Amplified genomic DNA was observed to be approximately 10 kb in size. Only five wells (A1–E1) are shown. M, GeneRuler 1 kb Plus DNA ladder (Fermentas). c, Nested PCR of the ‘Entotheonella’ 16S rRNA gene, the onnamide and polytheonamide gene clusters from wells sorted with single filament cells. WGA DNA was used as PCR templates. Each target gene PCR was performed in triplicate to prove consistency.
Extended Data Figure 2 Binning of ‘Entotheonella factor’ contigs and natural product biosynthetic genes.
a, Plot of G+C content versus relative frequency for all 18,093 contigs assembled from sequencing of the enriched filamentous bacterial fraction from Theonella swinhoei Y. BLASTX analysis was performed against the RefSeq database, and the taxonomic domain classification for the best hit was assigned to each contig. Scaffolds attributed to bacterial sources are shown in red, contigs attributed to Archaea are shown in blue, those exhibiting eukaryotic features in green, and unclassified contigs in black. b, Filtered scaffolds, in which contaminating non-bacterial contigs and bacterial contigs that could not be assembled into scaffolds were removed. Scaffolds containing ORFs predicted to code for natural product biosynthesis are indicated in blue. The Bacteria-associated contigs and scaffolds clustered into two main groups designated ‘Entotheonella factor TSY1’ for the dominant organism, ‘Entotheonella factor TSY2’ for the less-abundant organism, and pTSY for plasmid-associated scaffolds. Scaffolds containing biosynthetic genes associated with the known compound classes, cyclotheonamides (cth), konbamides (kon), keramamides (ker), and nazumamides (naz) all cluster with TSY1. The onnamide (onn) and polytheonamide (poy) biosynthetic genes are located on the plasmid, pTSY.
Extended Data Figure 3 Synteny analysis of TSY1 and TSY2.
All scaffolds of TSY2 were compared against all scaffolds of TSY1 using r2cat49. Blue line denotes syntenic scaffold alignments. Grey vertical and horizontal lines denote scaffold breaks. The horizontal bar at the bottom indicates coverage of the matches to the reference scaffold, TSY1, with maximal coverage indicated with black, fading to light grey with less coverage. Red indicates uncovered regions.
Extended Data Figure 4 Natural product biosynthetic gene loci from ‘Entotheonella factor TSY1’ and ‘Entotheonella factor TSY2’.
a–g, Natural product biosynthetic gene loci from ‘Entotheonella factor TSY1’ are shown in a–d, and those from ‘Entotheonella factor TSY2’ are shown in e–g. Biosynthetic gene clusters are numbered according to Supplementary Table 4 and open reading frames (ORFs) are colour-coded based on predicted function. Clusters associated with known compounds are indicated by name. PKS catalytic domains (spheres) are: ACP, acyl carrier protein; AT, acyltransferase; DH, dehydratase; EH, enoyl-CoA hydratase; ER, enoyl reductase; GN, GCN5-like N-acetyltransferase; KR, ketoreductase; KS, ketosynthase; MT, methyltransferase; PS, pyran synthase; TE, thioesterase. NRPS catalytic domains (squares) are: A, adenylation; C, condensation; E, epimerase; M, methyltransferase; NT, aminotransferase; T, thiolation; TE, thioesterase. Amino-acid residues based on adenylation domain specificity (black) or structure-based (red) predictions are indicated below the corresponding adenylation domain. Int, Integrase. Non-proteinogenic amino acids are: mLeu, methyl-leucine; EtGly, ethylglycine (2-aminobutyric acid); Dap, 2,3-diaminopropionate. The clusters are grouped as: non-ribosomal peptide, modular polyketide, and hybrid NRPS/PKS biosynthetic loci associated with TSY1 (a); proteusin gene clusters from TSY1 (b); type III PKS locus from TSY1 (c); ectoine locus from TSY1 (d); NRPS and PKS loci from TSY2 (e); type III PKS locus from TSY2 (f); putative proteusin pathway from TSY2 (g). Several ORFs could not be closed owing to assembly complexity, thus open ORFs are designated with three dashes.
Extended Data Figure 5 Mass spectra and retention times for polytheonamides A and B identified in extracts from T. swinhoei, and an enriched ‘Entotheonella’ cell fraction.
a–d, Monoisotopic [M+3H]3+ ion (calculated 1677.9181), retentions times (tR), and respective isotopic pattern for polytheonamides A and B (C219H376N60O72S) from authentic standards of polytheonamide A and B (a), a T. swinhoei extract (b) an enriched ‘Entotheonella’ cell extract (c), and a negative control (ethanol) (d).
Extended Data Figure 6 Mass-spectral molecular-network analysis of extracts from enriched ‘Entotheonella’ cell fractions and T. swinhoei.
Network analysis was performed as described previously25. Nodes represent consensus spectra and interconnecting edges indicate significant pairwise aligned spectra25. The width of an edge and the distance between two nodes correspond with the relatedness of the two consensus spectra. Related compounds cluster together to form structural families. Nodes were false-colour-coded according to the extract they were derived from: metabolites detected in both the enriched ‘Entotheonella’ fraction and the whole sponge extract are in blue, and metabolites only present in the whole sponge extract are in green. Nodes representing the polytheonamides, orbiculamide A, onnamide A and aurantosides were detected in both the whole sponge and the enriched ‘Entotheonella’ cell pellet. Theopederin D, nazumamide, and several onnamide and aurantoside congeners, as well as the keramamides, were only detected in the whole sponge extract (however, note that keramamides were detected in the ‘Entotheonella’ sample using a more sensitive mass spectrometry method; see Supplementary Table 6). The keramamides are clustering with orbiculamide A, thus suggesting the same biosynthetic origin. In addition, several structural families were detected to be derived from the ‘Entotheonella’ enriched fraction that could not be linked to any metabolites known from T. swinhoei. This observation suggests the presence of as yet unidentified metabolites produced by ‘Entotheonella’.
Extended Data Figure 7 Matrix-assisted laser desorption ionization (MALDI) mass-spectrometry detection of adenylation-domain substrate activation by ATP–pyrophosphate exchange assay.
a, Assay of CthA2 with the 20 proteinogenic amino acids and l-2,3-diaminopropionic acid. b, Assay of KerA5 with the 20 proteinogenic amino acids (n = 3; error bars, s.d.).
Extended Data Figure 8 Functional analysis of the orphan proteusin gene cluster TSY1_14.
a–d, The purified His-tagged precursor peptide of TSY1_14 was co-expressed in E. coli with a putative LanM-like lanthionine synthetase from the gene cluster (green, treated peptide) and compared to product of the TSY1_14 precursor overexpressed without putative modifying enzymes (grey, untreated control) by UPLC HESI–HRMS (see Supplementary Table 5 for deconvoluted protein masses). a, ESI–HRMS spectra of the full-length His-tagged precursor peptide after TCEP treatment. Three dehydrations were observed for the peptide co-expressed with the LanM-like enzyme. Mass shifts compared to expected masses may be due to cystine formation and prompted subsequent alkylation experiments. b, Mass spectra of the full-length His-tagged precursor peptide after addition of carbamidomethyl groups (CAM) to free thiols using iodoacetamide. Predominant peaks for the untreated control correspond to products with alkylation of five out of five cysteine residues, suggesting that all cysteines contained free thiol groups after reduction with TCEP. For the treated peptide products, two alkylations and three dehydrations were observed for the major peaks, whereas additional peaks suggest that the number of free thiols decreases for each observed dehydration of the precursor peptide, consistent with the formation of up to three lanthionine bridges. Probable α-N-gluconoylation was also observed (+Gluc). c, ESI–HRMS spectra of tryptic digests of the His-tagged and iodoacetamide-treated precursor peptides. Masses corresponding to the uncleaved peptide, Ma, are only observed for the coexpression sample, suggesting formation of a lanthionine bridge that spans the trypsin cleavage site. Addition of water to Ma as compared to the full peptide may be due to partial trypsin cleavage. d, MS–MS fragmentation of [Ma-3H2O+2CAM+3H]3+ maps all three dehydrations and both alkylations to the predicted core peptide (underlined).
Extended Data Figure 9 16S rRNA analysis and phylogeny of ‘Entotheonella’ spp. from various habitats.
a, Agarose gel of PCR products obtained with ‘Entotheonella’-specific primers from Japanese sponges. Marker (M), Pseudoceratina purpurea, Nakano-shima (lane 1), Stylissa carteri (lane 2), Penares aff. incrustans (lane 3), Hexadella sp. (lane 4), Penares sp. (lane 5), Asteropus simplex (lane 6), Topsentia sp. (lane 7), Amphimedon sp., Io-jima (lane 8), Erylus placenta (lane 9), Agelas nakamurai (lane 10), Aaptos ciliata (lane 11), Ceratopsion sp. (lane 12), Epipolasis sp., Hachijo-jima (lane 13), Dercitus simplex (lane 14), Pseudoceratina purpurea, Oshima-shinsone (lane 15), Haliclona digitata (lane 16), Petrosia volcano (lane 17), Epipolasis sp., Nagannu-jima (lane 18), Erylus nobilis (lane 19), Anthosigmella (Cliona) raromicrosclera, Mitsuke (lane 20), Mycale magellanica (lane 21), Amphimedon sp., Hachijo-jima (lane 22), Penares aff. incrustans (lane 23), Anthosigmella (Cliona) raromicrosclera, Kamikoshiki-jima (lane 24), Axinella sp. (lane 25), Discodermia calyx (lane 26), Theonella swinhoei (onnamide chemotype) (lane 27), Discodermia kiiensis (lane 28), negative control (N), positive control, filamentous bacteria from Theonella swinhoei W1 (P). From each PCR reaction (n = 1), 20 clones were sequenced. b, Agarose gel of ‘Entotheonella’ 16S rRNA PCR products from sponges from other locations. M, N and P, as above. Agelas dilatata (lane 1), Amphimedon compressa (lane 2), Aplysina aerophoba (lane 3), Cacospongia mycofijiensis (lane 4), Callyspongia vaginalis (lane 5), Dysidea avara (lane 6), Dysidea etheria (lane 7), Fascaplysinopsis sp. (lane 8), Ircinia felix (lane 9), Niphates digitalis (lane 10), Psammocinia aff. bulbosa (lane 11), Ptilocaulis sp. (lane 12), Stylissa carteri (lane 13), Xestospongia muta (lane 14), Xestospongia testudinaria (lane 15), Mediterranean sea water (lane 16), Florida sea water (lane 17), Red Sea sea water (lane 18). From each PCR reaction (n = 1), 20 clones were sequenced. c, 16S rRNA based maximum likelihood tree of the novel candidate phylum ‘Tectomicrobia’. The candidate phylum (grey box) consists of the ‘Entotheonella’ clade as well as an environmental cluster with sequences from marine and terrestrial environments and a cluster with only sponge-derived sequences. Names for other bacterial phyla, proteobacterial classes and deltaproteobacterial families are given. Numbers at nodes indicate bootstrap values as calculated by maximum likelihood and maximum parsimony analyses. Arrow, to outgroup. Scale bar indicates 10% sequence divergence.
Extended Data Figure 10 Analysis of PKS genes in the misakinolide chemotype T. swinhoei W1.
a, Light micrograph of the enriched ‘Entotheonella’ fraction (n = 3). b, Structure of misakinolide A. c, Phylogram of PKS amplicons generated from the total sponge DNA (blue labels) and the ‘Entotheonella’ fraction (red labels). Black labels belong to known PKS sequences that were retrieved from GenBank and belong to pikromycin (PikAIII), erythromycin (EryKS4), pimaricin (PimS3), myxothiazol (MtaD), barbamide (BarE), nodularin (NdaC), difficidin (DifKS14), onnamide (OnnKS11), bacillaene (BaeKS11) biosynthesis or to synthases putatively involved in production of methyl-branched lipids (SupA). KS numbers refer to the position of the KS in the PKS; that is, sequences were aligned with MUSCLE, and trees were inferred by the neighbour-joining method with 500 replicates, using the Jukes–Cantor correction model and the fatty acid synthase component FabH from E. coli BL21(DE3) as the outgroup. Bootstrap values larger than 50% are shown at the nodes.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-3, Supplementary Tables 1-10 and additional references. (PDF 1021 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-Non-Commercial-ShareAlike 3.0 Unported licence. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Wilson, M., Mori, T., Rückert, C. et al. An environmental bacterial taxon with a large and distinct metabolic repertoire. Nature 506, 58–62 (2014). https://doi.org/10.1038/nature12959
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12959
This article is cited by
-
Tools for microbial single-cell genomics for obtaining uncultured microbial genomes
Biophysical Reviews (2024)
-
Drug Discovery from Natural Sources
Current Pharmacology Reports (2023)
-
Microbiome profile of the Antarctic clam Laternula elliptica
Brazilian Journal of Microbiology (2023)
-
Lineage-specific energy and carbon metabolism of sponge symbionts and contributions to the host carbon pool
The ISME Journal (2022)
-
Ancient plant-like terpene biosynthesis in corals
Nature Chemical Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.