Abstract
A major challenge for the development of a highly effective AIDS vaccine is the identification of mechanisms of protective immunity. To address this question, we used a nonhuman primate challenge model with simian immunodeficiency virus (SIV). We show that antibodies to the SIV envelope are necessary and sufficient to prevent infection. Moreover, sequencing of viruses from breakthrough infections revealed selective pressure against neutralization-sensitive viruses; we identified a two-amino-acid signature that alters antigenicity and confers neutralization resistance. A similar signature confers resistance of human immunodeficiency virus (HIV)-1 to neutralization by monoclonal antibodies against variable regions 1 and 2 (V1V2), suggesting that SIV and HIV share a fundamental mechanism of immune escape from vaccine-elicited or naturally elicited antibodies. These analyses provide insight into the limited efficacy seen in HIV vaccine trials.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Rerks-Ngarm, S. et al. Vaccination with ALVAC and AIDSVAX to prevent HIV-1 infection in Thailand. N. Engl. J. Med. 361, 2209–2220 (2009)
Haynes, B. F. et al. Immune-correlates analysis of an HIV-1 vaccine efficacy trial. N. Engl. J. Med. 366, 1275–1286 (2012)
Tomaras, G. D. et al. Vaccine-induced plasma IgA specific for the C1 region of the HIV-1 envelope blocks binding and effector function of IgG. Proc. Natl Acad. Sci. USA 110, 9019–9024 (2013)
Rolland, M. et al. Increased HIV-1 vaccine efficacy against viruses with genetic signatures in Env V2. Nature 490, 417–420 (2012)
Liao, H. X. et al. Vaccine induction of antibodies against a structurally heterogeneous site of immune pressure within HIV-1 envelope protein variable regions 1 and 2. Immunity 38, 176–186 (2013)
Hammer, S. M. et al. Efficacy Trial of a DNA/rAd5 HIV-1 Preventive Vaccine. N. Engl. J. Med. 369, 2083–2092 (2013)
Keele, B. F. et al. Low-dose rectal inoculation of rhesus macaques by SIVsmE660 or SIVmac251 recapitulates human mucosal infection by HIV-1. J. Exp. Med. 206, 1117–1134 (2009)
Hidajat, R. et al. Correlation of vaccine-elicited systemic and mucosal nonneutralizing antibody activities with reduced acute viremia following intrarectal simian immunodeficiency virus SIVmac251 challenge of rhesus macaques. J. Virol. 83, 791–801 (2009)
Lai, L. et al. Prevention of infection by a granulocyte-macrophage colony-stimulating factor co-expressing DNA/modified vaccinia Ankara simian immunodeficiency virus vaccine. J. Infect. Dis. 204, 164–173 (2011)
Schell, J. B. et al. Significant protection against high-dose simian immunodeficiency virus challenge conferred by a new prime-boost vaccine regimen. J. Virol. 85, 5764–5772 (2011)
Barouch, D. H. et al. Vaccine protection against acquisition of neutralization-resistant SIV challenges in rhesus monkeys. Nature 482, 89–93 (2012)
Flatz, L. et al. Gene-based vaccination with a mismatched envelope protects against simian immunodeficiency virus infection in nonhuman primates. J. Virol. 86, 7760–7770 (2012)
Lai, L. et al. SIVmac239 MVA vaccine with and without a DNA prime, similar prevention of infection by a repeated dose SIVsmE660 challenge despite different immune responses. Vaccine 30, 1737–1745 (2012)
Letvin, N. L. et al. Immune and genetic correlates of vaccine protection against mucosal infection by SIV in monkeys. Sci. Transl. Med. 3, 81ra36 (2011)
Lopker, M. et al. Heterogeneity in neutralization sensitivities of viruses comprising the simian immunodeficiency virus SIVsmE660 isolate and vaccine challenge stock. J. Virol. 87, 5477–5492 (2013)
Fischer, W. et al. Distinct evolutionary pressures underlie diversity in simian immunodeficiency virus and human immunodeficiency virus lineages. J. Virol. 86, 13217–13231 (2012)
Fischer, W. et al. Polyvalent vaccines for optimal coverage of potential T-cell epitopes in global HIV-1 variants. Nature Med. 13, 100–106 (2007)
Hudgens, M. G. & Gilbert, P. B. Assessing vaccine effects in repeated low-dose challenge experiments. Biometrics 65, 1223–1232 (2009)
Hudgens, M. G. et al. Power to detect the effects of HIV vaccination in repeated low-dose challenge experiments. J. Infect. Dis. 200, 609–613 (2009)
Gray, E. S. et al. Isolation of a monoclonal antibody that targets the alpha-2 helix of gp120 and represents the initial autologous neutralizing-antibody response in an HIV-1 subtype C-infected individual. J. Virol. 85, 7719–7729 (2011)
Moore, P. L. et al. Limited neutralizing antibody specificities drive neutralization escape in early HIV-1 subtype C infection. PLoS Pathog. 5, e1000598 (2009)
Del Prete, G. Q. et al. Derivation and characterization of a simian immunodeficiency virus SIVmac239 variant with tropism for CXCR4. J. Virol. 83, 9911–9922 (2009)
Gray, R. T. A class of K-sample tests for comparing the cumulative incidence of a competing risk. Ann. Stat. 16, 1141–1154 (1988)
Lim, S. Y. et al. Contributions of Mamu-A*01 status and TRIM5 allele expression, but not CCL3L copy number variation, to the control of SIVmac251 replication in Indian-origin rhesus monkeys. PLoS Genet. 6, e1000997 (2010)
Lim, S. Y. et al. TRIM5alpha modulates immunodeficiency virus control in rhesus monkeys. PLoS Pathog. 6, e1000738 (2010)
Fenimore, P. W. et al. Designing and testing broadly-protective filoviral vaccines optimized for cytotoxic T-lymphocyte epitope coverage. PLoS ONE 7, e44769 (2012)
Yusim, K. et al. Genotype 1 and global hepatitis C T-cell vaccines designed to optimize coverage of genetic diversity. J. Gen. Virol. 91, 1194–1206 (2010)
Donaldson, M. M. et al. Optimization and qualification of an 8-color intracellular cytokine staining assay for quantifying T cell responses in rhesus macaques for pre-clinical vaccine studies. J. Immunol. Methods 386, 10–21 (2012)
Santra, S. et al. Heterologous prime/boost immunizations of rhesus monkeys using chimpanzee adenovirus vectors. Vaccine 27, 5837–5845 (2009)
Imholte, G. C. et al. A computational framework for the analysis of peptide microarray antibody binding data with application to HIV vaccine profiling. J. Immunol. Methods 395, 1–13 (2013)
Bolton, D. L. et al. Comparison of systemic and mucosal vaccination: impact on intravenous and rectal SIV challenge. Mucosal Immunol. 5, 41–52 (2012)
Tomaras, G. D. et al. Initial B-cell responses to transmitted human immunodeficiency virus type 1: virion-binding immunoglobulin M (IgM) and IgG antibodies followed by plasma anti-gp41 antibodies with ineffective control of initial viremia. J. Virol. 82, 12449–12463 (2008)
Alam, S. M. et al. Human immunodeficiency virus type 1 gp41 antibodies that mask membrane proximal region epitopes: antibody binding kinetics, induction, and potential for regulation in acute infection. J. Virol. 82, 115–125 (2008)
Shu, Y. et al. Efficient protein boosting after plasmid DNA or recombinant adenovirus immunization with HIV-1 vaccine constructs. Vaccine 25, 1398–1408 (2007)
Wu, L. et al. Enhanced exposure of the CD4-binding site to neutralizing antibodies by structural design of a membrane-anchored human immunodeficiency virus type 1 gp120 domain. J. Virol. 83, 5077–5086 (2009)
Li, M. et al. Human immunodeficiency virus type 1 env clones from acute and early subtype B infections for standardized assessments of vaccine-elicited neutralizing antibodies. J. Virol. 79, 10108–10125 (2005)
Zhao, J. et al. Preclinical studies of human immunodeficiency virus/AIDS vaccines: inverse correlation between avidity of anti-Env antibodies and peak postchallenge viremia. J. Virol. 83, 4102–4111 (2009)
Qin, L., Gilbert, P. B., Corey, L., McElrath, M. J. & Self, S. G. A framework for assessing immunological correlates of protection in vaccine trials. J. Infect. Dis. 196, 1304–1312 (2007)
Plotkin, S. A. & Gilbert, P. B. Nomenclature for immune correlates of protection after vaccination. Clin. Infect. Dis. 54, 1615–1617 (2012)
Acknowledgements
We thank the following individuals: W. Shi, L. Wu, S.-Y. Ko, L. Wang and W.-P. Kong for immunogen construction; M. M. Donaldson, S.-F. Kao, D. Quinn, J. Owuor, K. Denison, H. Balachandran, C. Luedemann, W. T. Williams, G. Overman, A. Deal, C. Brinkley and L. Racz for technical assistance with immunology assays; A. Ault for assistance managing NHP studies; S. O’Connor for deep-sequencing data; M. Seaman for providing plasmids encoding E660 envelopes; F. McCutchan, J. Overbaugh and J. Kim for HIV-1 strains; and P. Gilbert for advice with using the Aalen and Johansen model. This work was supported by the Intramural Research Program of the Vaccine Research Center, NIAID, NIH; by NIH contracts HHSN261200800001E (B.F.K., W.G.) and HHSN27201100016C (D.C.M.); by NIH grant AI100645 (B.T.K., W.F.); and by the Bill and Melinda Gates Foundation grant OPP1032317.
Author information
Authors and Affiliations
Contributions
M.R., R.A.S., R.A.K., G.J.N., N.L.L., S.S.R. and J.R.M. designed and supervised the study. W.F., Z.-Y.Y., B.T.K. and G.J.N. designed and manufactured immunogens. J.-P.M.T., N.L.L. and S.S.R. supervised nonhuman primate procedures. M.R., B.F.K., K.E.F., A.P.B., L.V.M., K.E.S., B.M.W., R.T.B., R.G., G.F., S.M.A., T.N.D., D.C.M., G.D.T., R.A.K. and J.R.M. supervised assays and performed primary data analysis. D.C.M. provided HIV-1 strains. S.D.S., R.D.M., H.C.W., C.L., M.K.L., L.V.M., L.S., K.E.S. and B.M.W. performed assays. M.R., W.G. and M.C.N. aggregated data and performed statistical analysis. M.R. and J.R.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Cellular immunogenicity of vaccines.
Gag- or Env-specific CD4 and CD8 T-cell responses measured by intracellular cytokine stimulation. Total T-cell responses were similar in all three active arms. a, Induction of T-cell responses are shown as the fraction of CD4 or CD8 memory T cells producing IFN-γ, IL2 or TNF in response to stimulation with overlapping peptides matched to the E660 challenge strain. Time points include peak post-DNA prime (week 10), pre-boost (week 25), peak post-rAd5 boost (week 32) and pre-challenge (week 52). b, The quality of the week 32 T-cell response is shown by the fraction of CD4 or CD8 cells responding to overlapping peptide pools matched to the E660 challenge swarm. There was no difference in the quality between any of the groups at any time point. c, Mosaic vaccination did not significantly improve the breadth of the T-cell response. Responses to pools of 10 overlapping peptides corresponding to mac239 Env or Gag, or the smE543 Env or Gag were tested for responses measured by ELISpot for the week 32 samples. Graphed is the number of positive pools (out of 23 for Env, and 13 for Gag) for each animal by group.
Extended Data Figure 2 Humoral immunogenicity of vaccines.
Mosaic immunization induced mildly lower humoral responses that were qualitatively different. a, b, Plasma IgG (a) or IgA (b) responses at week 32 were quantified against SIV envelope proteins derived from mac239, E660-CP3C, E660-CR54, or a mac239 V1V2 polypeptide expressed on a J08 scaffold. MFI, mean fluorescence intensity using a bead-based Luminex platform. AUC, area under the curve. c, CD4-binding site activity was measured by the ability of sera to cross-block CD4-Ig binding to mac239 or smE543 envelopes. d, Antibody-dependent cellular cytotoxicity mediated killing of SIV-infected target PBMC, shown as per cent specific killing. e, PBMC neutralization assay showing no substantial difference between immunization arms or time since vaccination. f, Neutralization by week 32 plasma was measured against three envelope-pseudotyped viruses. g, Neutralization of the E660 challenge stock using the TZM-bl indicator cell line. h, Week 32 plasma antibody binding to overlapping peptides spanning the SIV E543 envelope was quantified for the two envelope immunization arms. The mean response for all 20 animals in each arm (top) or the fraction of animals responding (bottom) is shown for peptides from the extracellular portion of the envelope. The arrow indicates an area near the V1V2 junction targeted by the mosaic but not the mac239 immunogen; several other areas, including C1, V3, C3 and V5, were better targeted by the mac239 immunogen.
Extended Data Figure 3 Viral pathogenesis and influence of TRIM5α alleles.
a, Viral load (VL) was measured weekly until 12 weeks post peak and then monthly thereafter. Curves are shown for all 74 infected animals and are synchronized by the peak VL. b, For each time point, the distribution of VL in each immunization arm was compared to the control arm. The mean difference (lower) and significance of the difference (Student’s t-test; upper) is graphed. c, The loss of CD4 cells following mucosal challenge is much more temperate than following intravenous challenge31. The most consistent measurable loss was for CD4 transitional memory cells (CD45RA–CCR7–CD28+); the change in the frequency of these cells relative to the pre-infection average is shown. Other CD4 subsets showed less dramatic depletion. d, e, All 80 animals were grouped according to predicted resistance based on TRIM5α allelism (resistant: TRIM5αQ/Q; sensitive: all other combinations). A significant effect of genetics on acquisition (d) and pathogenesis (e) was observed. Animals were randomized equally into the four immunization arms based on TRIM5α genotype (for all homozygous and heterozygous genotypes).
Extended Data Figure 4 Transmitted/founder (T/F) selection in any vaccine arm.
The number of T/F viruses with a variant from consensus was compared across all four arms for amino acid positions showing heterogeneity. A permutation test was used to compute the significance of a difference across all groups. The P values for positions 23, 45 and 47 remain significant after correction for multiple comparisons.
Extended Data Figure 5 Neutralization sensitivity of variant envelopes.
Nine envelope variants (Fig. 3) were evaluated for neutralization sensitivity by antisera from vaccinated animals (black or grey) and monoclonal antibodies to the CD4 binding site (brown) or the V1V2 loops (purple). a, The ICHM (reciprocal concentration of antisera resulting in 50% of maximum neutralization) for all neutralization experiments is summarized by animal (left) or monoclonal antibody (right) for the seven CP3C variants and the two CR54 variants. The range of ICHM across the viruses was less than twofold; that is, C1 sequence variations do not affect ICHM but only the fraction of neutralization-resistant virions within each virus preparation (Fig. 3e). b, In a separate experiment, sera from three vaccinated animals and five monoclonal antibodies were compared. Note that the V1V2 antibodies only neutralize ∼60% of the sensitive CP3C strain. c, Relative neutralization sensitivity was calculated by normalizing neutralization of each class of antibodies to 100% for CP3C.
Extended Data Figure 6 Pathogenesis of TR and A/K viruses.
Animals were divided into groups based on whether they were infected solely with TR viruses, A/K viruses, or both (that is, with multiple T/F per animal). Bars indicate the interquartile range of values. The peak and set point viral load did not differ according to which type of virus infected and replicated in the animal. In addition, no significant differences were observed when these data were split by vaccine arm.
Extended Data Figure 7 Immunological correlates of risk, plasma IgG.
a, b, Week 52 plasma IgG against the CP3C envelope is graphed against time to infection (uninfected animals were assigned a value of 13). Data are shown excluding A/K virus infections (a) or for all infections (b). Significant correlations are indicated by a linear least-squares regression line; statistics are nonparametric Spearman’s tests. c, d, Similar analyses using week 32 (peak) plasma, for all TR infections (c), or all viral infections (d). e, Avidity to SIV envelopes was measured by Biacore; for each KM analysis, animals were divided in two equal groups based on having lower than median disassociation rate (high avidity) vs higher (low avidity), for TR infections.
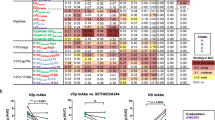
Extended Data Figure 8 Immunological correlates of risk, breadth of binding to linear peptides.
a, A multivariable regression of time to infection vs responses to each of the four regions shown in Fig. 4g was performed. All four regions provided independent predictive power. b, The binding activity to all four regions was summed; the total response to these four epitopes showed a high correlation with time to infection. c, The number of the four regions with positive responses within each animal was computed (no animal responded to all four). The line indicates a linear regression; statistics are based on a nonparametric Spearman’s test. d, The number of epitopes with positive responses across the entire envelope was computed for each animal. No correlation with protection (for all viruses or for only TR viruses) was seen with overall breadth. ns, P > 0.05. e, Average binding to the linear C3 peptides 119 and 120 correlates with time to infection for all animals, irrespective of virus. f, KM analysis comparing Env-immunized animals with a positive response to C3 peptides to those with a negative response.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-5. (PDF 625 kb)
Rights and permissions
About this article
Cite this article
Roederer, M., Keele, B., Schmidt, S. et al. Immunological and virological mechanisms of vaccine-mediated protection against SIV and HIV. Nature 505, 502–508 (2014). https://doi.org/10.1038/nature12893
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12893
This article is cited by
-
Differential V2-directed antibody responses in non-human primates infected with SHIVs or immunized with diverse HIV vaccines
Nature Communications (2022)
-
Glycopeptide epitope facilitates HIV-1 envelope specific humoral immune responses by eliciting T cell help
Nature Communications (2020)
-
The Antibodiome—Mapping the Humoral Immune Response to HIV
Current HIV/AIDS Reports (2019)
-
Recombinant HIV-1 vaccine candidates based on replication-defective flavivirus vector
Scientific Reports (2019)
-
Nonhuman primate models of human viral infections
Nature Reviews Immunology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.