Abstract
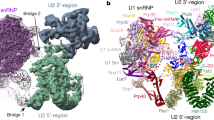
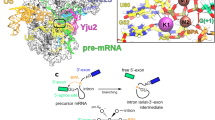
The active centre of the spliceosome consists of an intricate network formed by U5, U2 and U6 small nuclear RNAs, and a pre-messenger-RNA substrate. Prp8, a component of the U5 small nuclear ribonucleoprotein particle, crosslinks extensively with this RNA catalytic core. Here we present the crystal structure of yeast Prp8 (residues 885–2413) in complex with Aar2, a U5 small nuclear ribonucleoprotein particle assembly factor. The structure reveals tightly associated domains of Prp8 resembling a bacterial group II intron reverse transcriptase and a type II restriction endonuclease. Suppressors of splice-site mutations, and an intron branch-point crosslink, map to a large cavity formed by the reverse transcriptase thumb, and the endonuclease-like and RNaseH-like domains. This cavity is large enough to accommodate the catalytic core of group II intron RNA. The structure provides crucial insights into the architecture of the spliceosome active site, and reinforces the notion that nuclear pre-mRNA splicing and group II intron splicing have a common origin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Wahl, M. C., Will, C. L. & Lührmann, R. The spliceosome: design principles of a dynamic RNP machine. Cell 136, 701–718 (2009)
Wassarman, D. A. & Steitz, J. A. Interactions of small nuclear RNA’s with precursor messenger RNA during in vitro splicing. Science 257, 1918–1925 (1992)
Madhani, H. D. & Guthrie, C. A novel base-pairing interaction between U2 and U6 snRNAs suggests a mechanism for the catalytic activation of the spliceosome. Cell 71, 803–817 (1992)
Kandels-Lewis, S. & Seraphin, B. Involvement of U6 snRNA in 5′ splice site selection. Science 262, 2035–2039 (1993)
Lesser, C. F. & Guthrie, C. Mutations in U6 snRNA that alter splice site specificity: Implications for the active site. Science 262, 1982–1988 (1993)
Sun, J. S. & Manley, J. L. A novel U2–U6 snRNA structure is necessary for mammalian mRNA splicing. Genes Dev. 9, 843–854 (1995)
Yean, S.-L., Wuenschell, G., Termini, J. & Lin, R. J. Metal-ion coordination by U6 small nuclear RNA contributes to catalysis in the spliceosome. Nature 408, 881–884 (2000)
Steitz, T. A. & Steitz, J. A. A general two-metal-ion mechanism for catalytic RNA. Proc. Natl Acad. Sci. USA 90, 6498–6502 (1993)
Newman, A. J. & Norman, C. U5 snRNA interacts with exon sequences at 5′ and 3′ splice sites. Cell 68, 743–754 (1992)
Sontheimer, E. J. & Steitz, J. A. The U5 and U6 small nuclear RNAs as active site components of the spliceosome. Science 262, 1989–1996 (1993)
O'Keefe, R. T., Norman, C. & Newman, A. J. The invariant U5 snRNA loop 1 sequence is dispensable for the first catalytic step of pre-mRNA splicing in yeast. Cell 86, 679–689 (1996)
Achsel, T., Ahrens, K., Brahms, H., Teigelkamp, S. & Lührmann, R. The human U5-220kD protein (hPrp8) forms a stable RNA-free complex with several U5-specific proteins, including an unwindase, a homologue of ribosomal elongation factor EF-2, and a novel WD-40 protein. Mol. Cell. Biol. 18, 6756–6766 (1998)
Bartels, C., Urlaub, H., Lührmann, R. & Fabrizio, P. Mutagenesis suggests several roles of Snu114p in pre-mRNA splicing. J. Biol. Chem. 278, 28324–28334 (2003)
Small, E. C., Leggett, S. R., Winans, A. A. & Staley, J. P. The EF-G-like GTPase Snu114p regulates spliceosome dynamics mediated by Brr2p, a DExD/H box ATPase. Mol. Cell 23, 389–399 (2006)
Teigelkamp, S., Newman, A. J. & Beggs, J. D. Extensive interactions of PRP8 protein with the 5′ and 3′ splice sites during splicing suggest a role in stabilization of exon alignment by U5 snRNA. EMBO J. 14, 2602–2612 (1995)
Dix, I., Russell, C. S., O’Keefe, R. T., Newman, A. J. & Beggs, J. D. Protein-RNA interactions in the U5 snRNP of Saccharomyces cerevisiae. RNA 4, 1239–1250 (1998)
Vidal, V. P., Verdone, L., Mayes, A. E. & Beggs, J. D. Characterization of U6 snRNA-protein interactions. RNA 5, 1470–1481 (1999)
Reyes, J. L., Gustafson, E. H., Luo, H. R., Moore, M. J. & Konarska, M. M. The C-terminal region of hPrp8 interacts with the conserved GU dinucleotide at the 5′ splice site. RNA 5, 167–179 (1999)
MacMillan, A. M. et al. Dynamic association of proteins with the pre-mRNA branch region. Genes Dev. 8, 3008–3020 (1994)
Turner, I. A., Norman, C. M., Churcher, M. J. & Newman, A. J. Dissection of Prp8 protein defines multiple interactions with crucial RNA sequences in the catalytic core of the spliceosome. RNA 12, 375–386 (2006)
Grainger, R. J. & Beggs, J. D. Prp8 protein: at the heart of the spliceosome. RNA 11, 533–557 (2005)
Pena, V., Rozov, A., Fabrizio, P., Lührmann, R. & Wahl, M. C. Structure and function of an RNase H domain at the heart of the spliceosome. EMBO J. 27, 2929–2940 (2008)
Ritchie, D. B. et al. Structural elucidation of a PRP8 core domain from the heart of the spliceosome. Nature Struct. Mol. Biol. 15, 1199–1205 (2008)
Yang, K., Zhang, L., Xu, T., Heroux, A. & Zhao, R. Crystal structure of the β-finger domain of Prp8 reveals analogy to ribosomal proteins. Proc. Natl Acad. Sci. USA 105, 13817–13822 (2008)
Pena, V., Liu, S., Bujnicki, J. M., Lührmann, R. & Wahl, M. C. Structure of a multipartite protein-protein interaction domain in splicing factor Prp8 and its link to Retinitis pigmentosa. Mol. Cell 25, 615–624 (2007)
Zhang, L. et al. Crystal structure of the C-terminal domain of splicing factor Prp8 carrying retinitis pigmentosa mutants. Protein Sci. 16, 1024–1031 (2007)
Dlakić, M. & Mushegian, A. Prp8, the pivotal protein of the spliceosomal catalytic center, evolved from a retroelement-encoded reverse transcriptase. RNA 17, 799–808 (2011)
Boon, K. L. et al. Prp8 mutations that cause human retinitis pigmentosa lead to a U5 snRNP maturation defect in yeast. Nature Struct. Mol. Biol. 14, 1077–1083 (2007)
Weber, G. et al. Mechanism for Aar2p function as a U5 snRNP assembly factor. Genes Dev. 25, 1601–1612 (2011)
Joyce, C. M. & Steitz, T. A. Function and structure relationships in DNA polymerase. Annu. Rev. Biochem. 63, 777–822 (1994)
Dias, A. et al. The cap-snatching endonuclease of influenza virus polymerase resides in the PA subunit. Nature 458, 914–918 (2009)
Yuan, P. et al. Crystal structure of an avian influenza polymerase PAN reveals an endonuclease active site. Nature 458, 909–913 (2009)
Wyatt, J. R., Sontheimer, E. J. & Steitz, J. A. Site-specific cross-linking of mammalian U5 snRNP to the 5′ splice site before the first step of pre-mRNA splicing. Genes Dev. 6, 2542–2553 (1992)
Urlaub, H., Hartmuth, K., Kostka, S., Grelle, G. & Lührmann, R. A general approach for identification of RNA-protein cross-linking sites within native human spliceosomal small nuclear ribonucleoproteins (snRNPs). J. Biol. Chem. 275, 41458–41468 (2000)
Query, C. C. & Konarska, M. M. Suppression of multiple substrate mutations by spliceosomal prp8 alleles suggests functional correlations with ribosomal ambiguity mutants. Mol. Cell 14, 343–354 (2004)
Umen, J. G. & Guthrie, C. Mutagenesis of the yeast gene PRP8 reveals domains governing the specificity and fidelity of 3′ splice site selection. Genetics 143, 723–739 (1996)
Kuhn, A. N. & Brow, D. A. Suppressors of a cold-sensitive mutation in yeast U4 RNA define five domains in the splicing factor Prp8 that influence spliceosome activation. Genetics 155, 1667–1682 (2000)
Kuhn, A. N., Reichl, E. M. & Brow, D. A. Distinct domains of splicing factor Prp8 mediate different aspects of spliceosome activation. Proc. Natl Acad. Sci. USA 99, 9145–9149 (2002)
Sharp, P. A. On the origin of RNA splicing and introns. Cell 42, 397–400 (1985)
Cech, T. R. The generality of self-splicing RNA: relationship to nuclear mRNA splicing. Cell 44, 207–210 (1986)
Michel, F., Umesono, K. & Ozeki, H. Comparative and functional anatomy of group II catalytic introns—a review. Gene 82, 5–30 (1989)
Sharp, P. A. Five easy pieces. Science 254, 663 (1991)
Lambowitz, A. M. & Zimmerly, S. Mobile group II introns. Annu. Rev. Genet. 38, 1–35 (2004)
Pyle, A. M. & Lambowitz, A. M. in The RNA World 3rd edn (eds Gesteland, R. F., Cech, T. R. & Atkins, J. F. ) 469–505 (Cold Spring Harbor Laboratory Press, 2006)
Qui, Y.-L. & Palmer, J. D. Many different origins of trans splicing in a plant mitochondrial group II intron. J. Mol. Evol. 59, 80–89 (2004)
Toor, N. et al. Tertiary architecture of the Oceanobacillus iheyensis group II intron. RNA 16, 57–69 (2010)
Marcia, M. & Pyle, A. M. Visualizing group II intron catalysis through the stages of splicing. Cell 151, 497–507 (2012)
Matsuura, M., Noah, J. W. & Lambowitz, A. M. Mechanism of maturase-promoted group II intron splicing. EMBO J. 20, 7259–7270 (2001)
Rambo, R. P. & Doudna, J. A. Assembly of an active group II intron-maturase complex by protein dimerization. Biochemistry 43, 6486–6497 (2004)
Gu, S. Q. et al. Genetic identification of potential RNA-binding regions in a group II intron-encoded reverse transcriptase. RNA 16, 732–747 (2010)
Wagenbach, M. et al. Synthesis of wild type and mutant human hemoglobins in Saccharomyces cerevisiae. Biotechnology (N Y) 9, 57–61 (1991)
Christianson, T. W., Sikorski, R. S., Dante, M., Shero, J. H. & Hieter, P. Multifunctional yeast high-copy-number shuttle vectors. Gene 110, 119–122 (1992)
Leslie, A. G. W. & Powell, H. R. Processing diffraction data with Mosflm. Evolv. Methods Macromol. Crystallograph. 245, 41–51 (2007)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D 62, 72–82 (2006)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D 66, 22–25 (2010)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Sheldrick, G. M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D 66, 479–485 (2010)
Langer, G., Cohen, S. X., Lamzin, V. S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nature Protocols 3, 1171–1179 (2008)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Krissinel, E. & Henrick, K. Secondary-structure matching (PDBeFold), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D 60, 2256–2268 (2004)
Holm, L. & Park, J. DaliLite workbench for protein structure comparison. Bioinformatics 16, 566–567 (2000)
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001)
Ashkenazy, H., Erez, E., Martz, E., Pupko, T. & Ben-Tal, N. ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res. 38, 529–533 (2010)
Larkin, M. A. et al. ClustalW and ClustalX version 2. Bioinformatics 23, 2947–2948 (2007)
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983)
Heinig, M. & Frishman, D. STRIDE: a Web server for secondary structure assignment from known atomic coordinates of proteins. Nucleic Acids Res. 32, W500–W502 (2004)
Sikorski, R. S. & Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19–27 (1989)
Acknowledgements
We thank G. Murshudov for his help and guidance with crystallography; T.H.D. Nguyen, Y. Kondo, M. van Roon, J. Hardin, J. Li, C. Norman, A. Andreeva, A. Murzin and M. Yu for discussion and help; T. Ignjatovic and H. Oshikane for their contributions at the early stage of the project; L. Passmore and L. Jovine for reading of the manuscript; and M. Ikura for the gift of a calmodulin clone. We are grateful to the beamline staff at Diamond Light Source and European Synchrotron Radiation Facility for their help, and to E. Stephens and the LMB mass spectrometry facility for their help. W.P.G. thanks the Cambridge European Trust and Downing College for scholarships. This project was funded by the UK Medical Research Council.
Author information
Authors and Affiliations
Contributions
A.J.N. and K.N. initiated the project and worked on protein expression and purification for many years. Co-expression of Prp8 and Aar2 by A.J.N. was a crucial step of the project. W.P.G. successfully identified and expressed a stable large fragment of Prp8, crystallized the Prp8–Aar2 complex and solved and refined the structure almost single-handedly with practical support from K.N. and A.J.N. C.O. analysed the mercury derivative data and refined the structure of the P212121 crystal form. W.P.G. and K.N. analysed the structure and wrote the paper with important input from A.J.N. and C.O.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-18, Supplementary Tables 1-7 and Supplementary References. (PDF 3044 kb)
Rights and permissions
About this article
Cite this article
Galej, W., Oubridge, C., Newman, A. et al. Crystal structure of Prp8 reveals active site cavity of the spliceosome. Nature 493, 638–643 (2013). https://doi.org/10.1038/nature11843
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11843
This article is cited by
-
Structural Analysis of Monomeric RNA-Dependent Polymerases Revisited
Journal of Molecular Evolution (2022)
-
PRP8 Intein in Onygenales: Distribution and Phylogenetic Aspects
Mycopathologia (2019)
-
Formation of chimeric genes with essential functions at the origin of eukaryotes
BMC Biology (2018)
-
Phosphorylation by Prp4 kinase releases the self-inhibition of FgPrp31 in Fusarium graminearum
Current Genetics (2018)
-
Using bioinformatic and phylogenetic approaches to classify transposable elements and understand their complex evolutionary histories
Mobile DNA (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.