Abstract
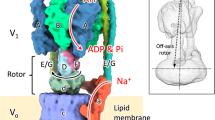
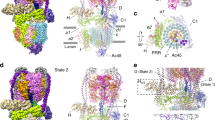
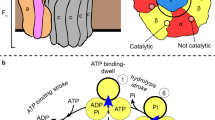
In various cellular membrane systems, vacuolar ATPases (V-ATPases) function as proton pumps, which are involved in many processes such as bone resorption and cancer metastasis, and these membrane proteins represent attractive drug targets for osteoporosis and cancer1. The hydrophilic V1 portion is known as a rotary motor, in which a central axis DF complex rotates inside a hexagonally arranged catalytic A3B3 complex using ATP hydrolysis energy, but the molecular mechanism is not well defined owing to a lack of high-resolution structural information. We previously reported on the in vitro expression, purification and reconstitution of Enterococcus hirae V1-ATPase from the A3B3 and DF complexes2,3. Here we report the asymmetric structures of the nucleotide-free (2.8 Å) and nucleotide-bound (3.4 Å) A3B3 complex that demonstrate conformational changes induced by nucleotide binding, suggesting a binding order in the right-handed rotational orientation in a cooperative manner. The crystal structures of the nucleotide-free (2.2 Å) and nucleotide-bound (2.7 Å) V1-ATPase are also reported. The more tightly packed nucleotide-binding site seems to be induced by DF binding, and ATP hydrolysis seems to be stimulated by the approach of a conserved arginine residue. To our knowledge, these asymmetric structures represent the first high-resolution view of the rotational mechanism of V1-ATPase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
Atomic coordinates and structure factors for the A3B3 and V1-ATPase complexes have been deposited in the Protein Data Bank under the accession codes 3VR2 (nucleotide-free A3B3 at 2.83), 3VR3 (nucleotide-bound A3B3 at 3.43), 3VR4 (nucleotide-free V1-ATPase at 2.23), 3VR5 (nucleotide-free V1-ATPase at 3.93) and 3VR6 (nucleotide-bound V1-ATPase at 2.73).
Change history
30 January 2013
The structure in Fig. 1e was corrected.
References
Forgac, M. Vacuolar ATPases: rotary proton pumps in physiology and pathophysiology. Nature Rev. Mol. Cell Biol. 8, 917–929 (2007)
Arai, S. et al. Reconstitution in vitro of the catalytic portion (NtpA3-B3-D-G complex) of Enterococcus hirae V-type Na+-ATPase. Biochem. Biophys. Res. Commun. 390, 698–702 (2009)
Saijo, S. et al. Crystal structure of the central axis DF complex of the prokaryotic V-ATPase. Proc. Natl Acad. Sci. USA 108, 19955–19960 (2011)
Walker, J. E. ATP synthesis by rotary catalysis (Nobel Lecture). Angew. Chem. Int. Edn Engl. 37, 2308–2319 (1998)
Mulkidjanian, A. Y., Makarova, K. S., Galperin, M. Y. & Koonin, E. V. Inventing the dynamo machine: the evolution of the F-type and V-type ATPases. Nature Rev. Microbiol. 5, 892–899 (2007)
Abrahams, J. P., Leslie, A. G., Lutter, R. & Walker, J. E. Structure at 2.8 Å resolution of F1-ATPase from bovine heart mitochondria. Nature 370, 621–628 (1994)
Menz, R. I., Walker, J. E. & Leslie, A. G. Structure of bovine mitochondrial F1-ATPase with nucleotide bound to all three catalytic sites: implications for the mechanism of rotary catalysis. Cell 106, 331–341 (2001)
Kagawa, R., Montgomery, M. G., Braig, K., Leslie, A. G. W. & Walker, J. E. The structure of bovine F1-ATPase inhibited by ADP and beryllium fluoride. EMBO J. 23, 2734–2744 (2004)
Bowler, M. W., Montgomery, M. G., Leslie, A. G. W. & Walker, J. E. Ground state structure of F1-ATPase from bovine heart mitochondria at 1.9 Å resolution. J. Biol. Chem. 282, 14238–14242 (2007)
Kabaleeswaran, V., Puri, N., Walker, J. E., Leslie, A. G. W. & Mueller, D. M. Novel features of the rotary catalytic mechanism revealed in the structure of yeast F1 ATPase. EMBO J. 25, 5433–5442 (2006)
Kabaleeswaran, V. et al. Asymmetric structure of the yeast F1 ATPase in the absence of bound nucleotides. J. Biol. Chem. 284, 10546–10551 (2009)
Stock, D., Leslie, A. G. & Walker, J. E. Molecular architecture of the rotary motor in ATP synthase. Science 286, 1700–1705 (1999)
Shirakihara, Y. et al. The crystal structure of the nucleotide-free α3β3 subcomplex of F1-ATPase from the thermophilic Bacillus PS3 is a symmetric trimer. Structure 5, 825–836 (1997)
Cingolani, G. & Duncan, T. M. Structure of the ATP synthase catalytic complex (F1) from Escherichia coli in an autoinhibited conformation. Nature Struct. Mol. Biol. 18, 701–707 (2011)
Noji, H., Yasuda, R., Yoshida, M. & Kinosita, K. Direct observation of the rotation of F1-ATPase. Nature 386, 299–302 (1997)
Yasuda, R., Noji, H., Yoshida, M., Kinosita, K. & Itoh, H. Resolution of distinct rotational substeps by submillisecond kinetic analysis of F1-ATPase. Nature 410, 898–904 (2001)
Adachi, K. et al. Coupling of rotation and catalysis in F1-ATPase revealed by single-molecule imaging and manipulation. Cell 130, 309–321 (2007)
Toei, M. et al. Dodecamer rotor ring defines H+/ATP ratio for ATP synthesis of prokaryotic V-ATPase from Thermus thermophilus. Proc. Natl Acad. Sci. USA 104, 20256–20261 (2007)
Maher, M. J. et al. Crystal structure of A3B3 complex of V-ATPase from Thermus thermophilus. EMBO J. 28, 3771–3779 (2009)
Numoto, N., Hasegawa, Y., Takeda, K. & Miki, K. Inter-subunit interaction and quaternary rearrangement defined by the central stalk of prokaryotic V1-ATPase. EMBO Rep. 10, 1228–1234 (2009)
Imamura, H. et al. Rotation scheme of V1-motor is different from that of F1-motor. Proc. Natl Acad. Sci. USA 102, 17929–17933 (2005)
Murata, T., Igarashi, K., Kakinuma, Y. & Yamato, I. Na+ binding of V-type Na+-ATPase in Enterococcus hirae. J. Biol. Chem. 275, 13415–13419 (2000)
Murata, T., Yamato, I., Kakinuma, Y., Leslie, A. G. W. & Walker, J. E. Structure of the rotor of the V-Type Na+-ATPase from Enterococcus hirae. Science 308, 654–659 (2005)
Murata, T. et al. Ion binding and selectivity of the rotor ring of the Na+-transporting V-ATPase. Proc. Natl Acad. Sci. USA 105, 8607–8612 (2008)
Mizutani, K. et al. Structure of the rotor ring modified with N,N-dicyclohexylcarbodiimide of the Na+-transporting vacuolar ATPase. Proc. Natl Acad. Sci. USA 108, 13474–13479 (2011)
Murata, T., Yamato, I. & Kakinuma, Y. Structure and mechanism of vacuolar Na+-translocating ATPase from Enterococcus hirae. J. Bioenerg. Biomembr. 37, 411–413 (2005)
Yamamoto, M. et al. Interaction and stoichiometry of the peripheral stalk subunits NtpE and NtpF and the N-terminal hydrophilic domain of NtpI of Enterococcus hirae V-ATPase. J. Biol. Chem. 283, 19422–19431 (2008)
Zhou, M. et al. Mass spectrometry of intact V-type ATPases reveals bound lipids and the effects of nucleotide binding. Science 334, 380–385 (2011)
Liu, Q. et al. Site-directed mutagenesis of the yeast V-ATPase A subunit. J. Biol. Chem. 272, 11750–11756 (1997)
Dittrich, M., Hayashi, S. & Schulten, K. On the mechanism of ATP hydrolysis in F1-ATPase. Biophys. J. 85, 2253–2266 (2003)
Kigawa, T. et al. Preparation of Escherichia coli cell extract for highly productive cell-free protein expression. J. Struct. Funct. Genomics 5, 63–68 (2004)
Deluca, M. & McElroy, W. D. Purification and properties of firefly luciferase. Methods Enzymol. 57, 3–15 (1978)
Battye, T. G. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. W. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D 67, 271–281 (2011)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D 66, 22–25 (2010)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Laskowski, R. A., MacArthur, M. W. & Thornton, J. M. Validation of protein models derived from experiment. Curr. Opin. Struct. Biol. 8, 631–639 (1998)
Lovell, S. C. et al. Structure validation by Cα geometry: φ, ψ and Cβ deviation. Proteins 50, 437–450 (2003)
Acknowledgements
We thank J. E. Walker for his suggestions, especially through the structural studies of F1-ATPase. The synchrotron radiation experiments were performed at SPring-8 and Photon Factory (proposals 2008S2-001, 2011S2-005, 2009G660, 2009B1031 and 2012G132). We also thank the beamline staff at BL41XU of SPring-8 (Harima, Japan) and NE3A, NW12A and BL1A of Photon Factory (Tsukuba, Japan) for help during data collection. This work was supported by the Targeted Proteins Research Program, grants-in-aid (23370047, 23118705), Special Coordination Funds for Promoting Science and Technology from the Ministry of Education, Culture, Sports, Science and Technology of the Japanese government.
Author information
Authors and Affiliations
Contributions
T.M. designed the study. S.A., Y.K. and N.O. constructed DNAs. Y.I.-K., T.T. and M.S. expressed and purified the proteins. K.S. and T.M. crystallized the proteins. S.A., S.S., K.S., K.M. and T.M. collected X-ray data. S.A., S.S. and K.M. processed and refined X-ray data. S.A. and K.S. performed functional analysis. S.A., S.S., I.Y. and T.M. analysed the results. S.A. and S.S. prepared figures and videos. T.M. wrote the paper. All authors discussed the results and commented on the manuscript. The study was managed by S.Y., S.I., I.Y. and T.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-17, Supplementary Tables 1-5, a Supplementary Discussion and Supplementary References. This file was replaced on 21 January 2013 as the original file posted online had corrupted. (PDF 12825 kb)
conformational changes of A3B3 complex induced by AMP-PNP:Mg binding
The colouring and viewing are consistent with Fig. 1. The video was generated by morphing between X-ray crystal structures of nucleotide-free (eA3B3) and nucleotide-bound (bA3B3) A3B3 complexes using PyMOL. (MOV 9918 kb)
conformational changes of nucleotide-free eA3B3 complex induced by DF binding
The colouring and viewing are consistent with Fig. 2. The video was generated by morphing between X-ray crystal structures of nucleotide-free A3B3 (eA3B3) and A3B3DF (eV1) complexes using PyMOL. (MOV 12726 kb)
conformational changes of nucleotide-bound A3B3 complex induced by DF binding
The colouring and viewing are consistent with Fig. 2. The video was generated by morphing between X-ray crystal structures of nucleotide-bound A3B3 (bA3B3) and A3B3DF (bV1) complexes using PyMOL. (MOV 13168 kb)
conformational difference of the nucleotide-binding sites of nucleotide-bound V1-ATPase
The colouring and viewing are consistent with Fig. 3. The video was generated by morphing between X-ray crystal structures of the ACBO’ pair (bound form) and ACRBCR pair (tight form) in nucleotide-bound A3B3DF (bV1) complex using PyMOL. (MOV 2162 kb)
Rights and permissions
About this article
Cite this article
Arai, S., Saijo, S., Suzuki, K. et al. Rotation mechanism of Enterococcus hirae V1-ATPase based on asymmetric crystal structures. Nature 493, 703–707 (2013). https://doi.org/10.1038/nature11778
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11778
This article is cited by
-
Design of allosteric sites into rotary motor V1-ATPase by restoring lost function of pseudo-active sites
Nature Chemistry (2023)
-
Downregulated ATP6V1B1 expression acidifies the intracellular environment of cancer cells leading to resistance to antibody-dependent cellular cytotoxicity
Cancer Immunology, Immunotherapy (2021)
-
Cryo-EM structures of intact V-ATPase from bovine brain
Nature Communications (2020)
-
High resolution structure of hexameric herpesvirus DNA-packaging motor elucidates revolving mechanism and ends 20-year fervent debate
Protein & Cell (2020)
-
Translation of the long-term fundamental studies on viral DNA packaging motors into nanotechnology and nanomedicine
Science China Life Sciences (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.