Abstract
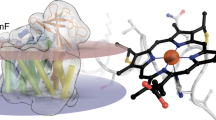
Cytochrome c oxidase is a member of the haem copper oxidase superfamily (HCO)1. HCOs function as the terminal enzymes in the respiratory chain of mitochondria and aerobic prokaryotes, coupling molecular oxygen reduction to transmembrane proton pumping. Integral to the enzyme’s function is the transfer of electrons from cytochrome c to the oxidase via a transient association of the two proteins. Electron entry and exit are proposed to occur from the same site on cytochrome c2,3,4. Here we report the crystal structure of the caa3-type cytochrome oxidase from Thermus thermophilus, which has a covalently tethered cytochrome c domain. Crystals were grown in a bicontinuous mesophase using a synthetic short-chain monoacylglycerol as the hosting lipid. From the electron density map, at 2.36 Å resolution, a novel integral membrane subunit and a native glycoglycerophospholipid embedded in the complex were identified. Contrary to previous electron transfer mechanisms observed for soluble cytochrome c, the structure reveals the architecture of the electron transfer complex for the fused cupredoxin/cytochrome c domain, which implicates different sites on cytochrome c for electron entry and exit. Support for an alternative to the classical proton gate characteristic of this HCO class is presented.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ferguson-Miller, S. & Babcock, G. T. Heme/copper terminal oxidases. Chem. Rev. 96, 2889–2908 (1996)
Muresanu, L. et al. The electron transfer complex between cytochrome c552 and the CuA domain of the Thermus thermophilus ba3 oxidase. A combined NMR and computational approach. J. Biol. Chem. 281, 14503–14513 (2006)
Maneg, O., Ludwig, B. & Malatesta, F. Different interaction modes of two cytochrome-c oxidase soluble CuA fragments with their substrates. J. Biol. Chem. 278, 46734–46740 (2003)
Maneg, O., Malatesta, F., Ludwig, B. & Drosou, V. Interaction of cytochrome c with cytochrome oxidase: two different docking scenarios. Biochim. Biophys. Acta 1655, 274–281 (2004)
Pereira, M. M., Santana, M. & Teixeira, M. A novel scenario for the evolution of haem-copper oxygen reductases. Biochim. Biophys. Acta 1505, 185–208 (2001)
Pereira, M. M., Sousa, F. L., Verissimo, A. F. & Teixeira, M. Looking for the minimum common denominator in haem-copper oxygen reductases: towards a unified catalytic mechanism. Biochim. Biophys. Acta 1777, 929–934 (2008)
Misquitta, L. V. et al. Membrane protein crystallization in lipidic mesophases with tailored bilayers. Structure 12, 2113–2124 (2004)
Caffrey, M., Lyons, J., Smyth, T. & Hart, D. J. in Current Topics in Membranes Vol. 63 (ed. DeLucas, L. J. ) Ch. 4 83–108 (Academic Press, 2009)
Höfer, N., Aragão, D., Lyons, J. A. & Caffrey, M. Membrane protein crystallization in lipidic mesophases. Hosting lipid effects on the crystallization and structure of a transmembrane peptide. Cryst. Growth Des. 11, 1182–1192 (2011)
Li, D., Lee, J. & Caffrey, M. Crystallizing membrane proteins in lipidic mesophases. A host lipid screen. Cryst. Growth Des. 11, 530–537 (2011)
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011)
Caffrey, M. Crystallizing membrane proteins for structure determination: use of lipidic mesophases. Annu. Rev. Biophys. 38, 29–51 (2009)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Fee, J. A., Choc, M. G., Findling, K. L., Lorence, R. & Yoshida, T. Properties of a copper-containing cytochrome c1aa3 complex: a terminal oxidase of the extreme thermophile Thermus thermophilus HB8. Proc. Natl Acad. Sci. USA 77, 147–151 (1980)
Steffens, G. C., Biewald, R. & Buse, G. Cytochrome c oxidase is a three-copper, two-heme-A protein. Eur. J. Biochem. 164, 295–300 (1987)
Mather, M. W., Springer, P., Hensel, S., Buse, G. & Fee, J. A. Cytochrome oxidase genes from Thermus thermophilus. Nucleotide sequence of the fused gene and analysis of the deduced primary structures for subunits I and III of cytochrome caa3 . J. Biol. Chem. 268, 5395–5408 (1993)
Mather, M. W., Springer, P. & Fee, J. A. Cytochrome oxidase genes from Thermus thermophilus. Nucleotide sequence and analysis of the deduced primary structure of subunit IIc of cytochrome caa3 . J. Biol. Chem. 266, 5025–5035 (1991)
Lübben, M. & Morand, K. Novel prenylated hemes as cofactors of cytochrome oxidases. Archaea have modified hemes A and O. J. Biol. Chem. 269, 21473–21479 (1994)
Backgren, C., Hummer, G., Wikstrom, M. & Puustinen, A. Proton translocation by cytochrome c oxidase can take place without the conserved glutamic acid in subunit I. Biochemistry 39, 7863–7867 (2000)
Pereira, M. M., Sousa, F. L., Teixeira, M., Nyquist, R. M. & Heberle, J. A tyrosine residue deprotonates during oxygen reduction by the caa3 reductase from Rhodothermus marinus. FEBS Lett. 580, 1350–1354 (2006)
Soares, C. M., Baptista, A. M., Pereira, M. M. & Teixeira, M. Investigation of protonatable residues in Rhodothermus marinus caa3 haem-copper oxygen reductase: comparison with Paracoccus denitrificans aa3 haem-copper oxygen reductase. J. Biol. Inorg. Chem. 9, 124–134 (2004)
Santana, M., Pereira, M. M., Elias, N. P., Soares, C. M. & Teixeira, M. Gene cluster of Rhodothermus marinus high-potential iron-sulfur protein: oxygen oxidoreductase, a caa(3)-type oxidase belonging to the superfamily of heme-copper oxidases. J. Bacteriol. 183, 687–699 (2001)
Srinivasan, V. et al. Structure at 1.3 Å resolution of Rhodothermus marinus caa3 cytochrome c domain. J. Mol. Biol. 345, 1047–1057 (2005)
Wienk, H. et al. Interaction of cytochrome c with cytochrome c oxidase: an NMR study on two soluble fragments derived from Paracoccus denitrificans. Biochemistry 42, 6005–6012 (2003)
Roberts, V. A. & Pique, M. E. Definition of the interaction domain for cytochrome c on cytochrome c oxidase. J. Biol. Chem. 274, 38051–38060 (1999)
Lange, C. & Hunte, C. Crystal structure of the yeast cytochrome bc1 complex with its bound substrate cytochrome c. Proc. Natl Acad. Sci. USA 99, 2800–2805 (2002)
Page, C. C., Moser, C. C. & Dutton, P. L. Mechanism for electron transfer within and between proteins. Curr. Opin. Chem. Biol. 7, 551–556 (2003)
Drosou, V., Malatesta, F. & Ludwig, B. Mutations in the docking site for cytochrome c on the Paracoccus heme aa3 oxidase. Electron entry and kinetic phases of the reaction. Eur. J. Biochem. 269, 2980–2988 (2002)
Drosou, V., Reincke, B., Schneider, M. & Ludwig, B. Specificity of the interaction between the Paracoccus denitrificans oxidase and its substrate cytochrome c: comparing the mitochondrial to the homologous bacterial cytochrome c552, and its truncated and site-directed mutants. Biochemistry 41, 10629–10634 (2002)
Witt, H., Malatesta, F., Nicoletti, F., Brunori, M. & Ludwig, B. Tryptophan 121 of subunit II is the electron entry site to cytochrome-c oxidase in Paracoccus denitrificans. Involvement of a hydrophobic patch in the docking reaction. J. Biol. Chem. 273, 5132–5136 (1998)
Lomize, A. L., Pogozheva, I. D., Lomize, M. A. & Mosberg, H. I. Positioning of proteins in membranes: a computational approach. Protein Sci. 15, 1318–1333 (2006)
Buse, G., Soulimane, T., Dewor, M., Meyer, H. E. & Bluggel, M. Evidence for a copper-coordinated histidine-tyrosine cross-link in the active site of cytochrome oxidase. Protein Sci. 8, 985–990 (1999)
Soulimane, T., Kiefersauer, R. & Than, M. E. in Membrane Protein Purification and Crystallization (Second Edition) (eds Hunte, C., von Jagow, G. & Schägger, H. ) Ch. 14 229–251 (Academic Press, 2003)
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nature Protocols 4, 706–731 (2009)
Cherezov, V., Peddi, A., Muthusubramaniam, L., Zheng, Y. F. & Caffrey, M. A robotic system for crystallizing membrane and soluble proteins in lipidic mesophases. Acta Crystallogr. D 60, 1795–1807 (2004)
Soulimane, T., Than, M. E., Dewor, M., Huber, R. & Buse, G. Primary structure of a novel subunit in ba3-cytochrome oxidase from Thermus thermophilus. Protein Sci. 9, 2068–2073 (2000)
Fischetti, R. F. et al. Mini-beam collimator enables microcrystallography experiments on standard beamlines. J. Synchrotron Radiat. 16, 217–225 (2009)
Cherezov, V. et al. Rastering strategy for screening and centring of microcrystal samples of human membrane proteins with a sub-10 micron size X-ray synchrotron beam. J. R. Soc. Interface 6, S587–S597 (2009)
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Cryst. 43, 186–190 (2010)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D 62, 72–82 (2006)
Collaborative Computational Project, number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007)
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Bashford, D. & Gerwert, K. Electrostatic calculations of the pKa values of ionizable groups in bacteriorhodopsin. J. Mol. Biol. 224, 473–486 (1992)
Zhang, L. & Hermans, J. Hydrophilicity of cavities in proteins. Proteins 24, 433–438 (1996)
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005)
MacKerell, A. D. et al. All atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998)
MacKerell, A. D., Feig, M. & Brooks, C. L. Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25, 1400–1415 (2004)
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010)
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010)
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996)
Ho, B. K. & Gruswitz, F. HOLLOW: generating accurate representations of channel and interior surfaces in molecular structures. BMC Struct. Biol. 8, 49 (2008)
Krissinel, E. & Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D 60, 2256–2268 (2004)
Šali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993)
Bond, C. S. & Schuttelkopf, A. W. ALINE: a WYSIWYG protein-sequence alignment editor for publication-quality alignments. Acta Crystallogr. D 65, 510–512 (2009)
Acknowledgements
We acknowledge support from Science Foundation Ireland Grants 07/IN.1/B1836 (M.C.) and BICF685 (T.S.), the National Institutes of Health Grants GM75915, P50GM073210, U54GM094599 (M.C.), and FP7 COST CM0902 (M.C.) and Marie Curie Actions PIEF-GA-2009-235612 (D.A.). We thank D. Hart, J. Lee and T. Smyth for help with 7.7 MAG synthesis, M. Dewor for amino acid sequencing, C. Soares for discussions regarding surface potential calculations, and G. Winter for help with data reduction from multiple crystals. The assistance and support of beamline scientists at the Advanced Photon Source (23-ID), Diamond Light Source (I24) and Swiss Light Source (PX1) are gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
J.A.L. synthesized 7.7 MAG, optimized crystallization conditions, collected and processed data, solved, refined and analysed the structure, and wrote the manuscript. D.A. collected crystals, collected and processed data, solved, refined and analysed the structure, and wrote the manuscript. O.S. performed initial protein production, purification, crystallization and data collection. A.V.P. carried out molecular dynamics simulations and pKa calculations. T.S. initiated the project, produced, purified and characterized protein, helped with crystallization, data collection and processing and structure analysis, and wrote the manuscript. M.C. was responsible for the overall project strategy and management and oversaw manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Text, Supplementary Figures 1-15, Supplementary Tables 1-3 and Supplementary references. (PDF 2701 kb)
Rights and permissions
About this article
Cite this article
Lyons, J., Aragão, D., Slattery, O. et al. Structural insights into electron transfer in caa3-type cytochrome oxidase. Nature 487, 514–518 (2012). https://doi.org/10.1038/nature11182
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11182
This article is cited by
-
Evaluation of Cyc1 protein stability in Acidithiobacillus ferrooxidans bacterium after E121D mutation by molecular dynamics simulation to improve electron transfer
Journal of Microbiology (2022)
-
Architecture of bacterial respiratory chains
Nature Reviews Microbiology (2021)
-
Copper signalling: causes and consequences
Cell Communication and Signaling (2018)
-
The quaternary structure of Thermus thermophilus aldehyde dehydrogenase is stabilized by an evolutionary distinct C-terminal arm extension
Scientific Reports (2018)
-
The cubicon method for concentrating membrane proteins in the cubic mesophase
Nature Protocols (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.