Abstract
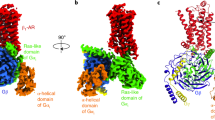
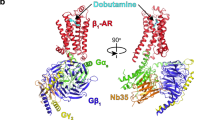
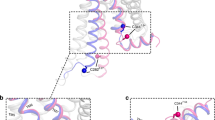
G-protein-coupled receptors are the largest class of cell-surface receptors, and these membrane proteins exist in equilibrium between inactive and active states1,2,3,4,5,6,7,8,9,10,11,12,13. Conformational changes induced by extracellular ligands binding to G-protein-coupled receptors result in a cellular response through the activation of G proteins. The A2A adenosine receptor (A2AAR) is responsible for regulating blood flow to the cardiac muscle and is important in the regulation of glutamate and dopamine release in the brain14. Here we report the raising of a mouse monoclonal antibody against human A2AAR that prevents agonist but not antagonist binding to the extracellular ligand-binding pocket, and describe the structure of A2AAR in complex with the antibody Fab fragment (Fab2838). This structure reveals that Fab2838 recognizes the intracellular surface of A2AAR and that its complementarity-determining region, CDR-H3, penetrates into the receptor. CDR-H3 is located in a similar position to the G-protein carboxy-terminal fragment in the active opsin structure1 and to CDR-3 of the nanobody in the active β2-adrenergic receptor structure2, but locks A2AAR in an inactive conformation. These results suggest a new strategy to modulate the activity of G-protein-coupled receptors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Scheerer, P. et al. Crystal structure of opsin in its G-protein-interacting conformation. Nature 455, 497–502 (2008)
Rasmussen, S. G. F. et al. Structure of a nanobody-stabilized active state of the β2 adrenoceptor. Nature 469, 175–180 (2011)
Palczewski, K. et al. Crystal structure of rhodopsin: a G protein-coupled receptor. Science 289, 739–745 (2000)
Shimamura, T. et al. Crystal structure of squid rhodopsin with intracellularly extended cytoplasmic region. J. Biol. Chem. 283, 17753–17756 (2008)
Murakami, M. & Kouyama, T. Crystal structure of squid rhodopsin. Nature 453, 363–367 (2008)
Warne, T. et al. Structure of a β1-adrenergic G-protein-coupled receptor. Nature 454, 486–491 (2008)
Rasmussen, S. G. F. et al. Crystal structure of the human β2 adrenergic G-protein-coupled receptor. Nature 450, 383–387 (2007)
Cherezov, V. et al. High-resolution crystal structure of an engineered human β2-adrenergic G protein-coupled receptor. Science 318, 1258–1265 (2007)
Jaakola, V.-P. et al. The 2.6 angstrom crystal structure of a human A2A adenosine receptor bound to an antagonist. Science 322, 1211–1217 (2008)
Wu, B. et al. Structures of the CXCR4 chemokine GPCR with small-molecule and cyclic peptide antagonists. Science 330, 1066–1071 (2010)
Shimamura, T. et al. Structure of the human histamine H1 receptor complex with doxepin. Nature 475, 65–70 (2011)
Chien, E. Y. T. et al. Structure of the human dopamine D3 receptor in complex with a D2/D3 selective antagonist. Science 330, 1091–1095 (2010)
Rasmussen, S. G. F. et al. Crystal structure of the β2 adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011)
Fredholm, B. B., Chen, J. F., Masino, S. A. & Vaugeois, J. M. Actions of adenosine at its receptors in the CNS: insights from knockouts and drugs. Annu. Rev. Pharmacol. Toxicol. 45, 385–412 (2005)
Müller, C. E. & Jacobson, K. A. Recent developments in adenosine receptor ligands and their potential as novel drugs. Biochim. Biophys. Acta 1808, 1290–1308 (2011)
Xu, F. et al. Structure of an agonist-bound human A2A adenosine receptor. Science 332, 322–327 (2011)
Ballesteros, J. A. &. Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods Neurosci. 25, 366–428 (1995)
Lebon, G. et al. Agonist-bound adenosine A2A receptor structures reveal common features of GPCR activation. Nature 474, 521–525 (2011)
Doré, A. S. et al. Structure of the adenosine A2A receptor in complex with ZM241385 and the xanthines XAC and caffeine. Structure 19, 1283–1293 (2011)
Chung, K. Y. et al. Conformational changes in the G protein Gs induced by the β2 adrenergic receptor. Nature 477, 611–615 (2011)
Yao, X. J. et al. The effect of ligand efficacy on the formation and stability of a GPCR-G protein complex. Proc. Natl Acad. Sci. USA 106, 9501–9506 (2009)
Warne, T. et al. The structural basis for agonist and partial agonist action on a β1-adrenergic receptor. Nature 469, 241–244 (2011)
Rosenbaum, D. M. et al. Structure and function of an irreversible agonist-β2 adrenoceptor complex. Nature 469, 236–240 (2011)
Yurugi-Kobayashi, T. et al. Comparison of functional non-glycosylated GPCRs expression in Pichia pastoris. Biochem. Biophys. Res. Commun. 380, 271–276 (2009)
Köhler, G. & Milstein, C. Continuous cultures of fused cells secreting antibody of predefined specificity. Nature 256, 495–497 (1975)
Warne, T., Chirnside, J. & Schertler, G. F. X. Expression and purification of truncated, non-glycosylated turkey β-adrenergic receptors for crystallization. Biochim. Biophys. Acta 1610, 133–140 (2003)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Afonine, P. V., Grosse-Kunstleve, R. W. & Adams, P. D. A robust bulk-solvent correction and anisotropic scaling procedure. Acta Crystallogr. D 61, 850–855 (2005)
Laskowski, R. A., MacArthur, M. W. & Thornton, J. M. Validation of protein models derived from experiment. Curr. Opin. Struct. Biol. 8, 631–639 (1998)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Yang, Z. R., Thomson, R., McNeil, P. & Esnouf, R. M. RONN: the bio-basis function neural network technique applied to the detection of natively disordered regions in proteins. Bioinformatics 21, 3369–3376 (2005)
DeLano, W. L. The PyMOL Molecular Graphics System 〈http://www.pymol.org〉 (2002)
Acknowledgements
This work was supported by the ERATO Human Receptor Crystallography Project of the Japan Science and Technology Agency (S.I.), by the Targeted Proteins Research Program of MEXT, Japan (S.I.), and by Development of New Functional Antibody Technologies (New Energy and Industrial Technology Development Organization, Japan) (T. Hamakubo). It was also partly funded by the Biotechnology and Biological Sciences Research Council (BB/G023425/1) (S.I.). The work was partly performed in the Membrane Protein Laboratory funded by the Wellcome Trust (grant 062164/Z/00/Z) (S.I.) at the Diamond Light Source, UK. Data were collected at Diamond Light Source beamline I24 with the assistance of G. Evans, R. Owen, D. Axford and J. Aishima.
Author information
Authors and Affiliations
Contributions
S.I. and T.M. designed the original research project. T.Y.-K. and T.K. established the A2AAR expression and purification protocols. T. Hino and C.I.-S. expressed, purified and characterized the receptor. H.I., Y.N.-N., O.K.-A. and T. Hamakubo performed the immunization, selection and isolation of antibodies. T. Hino, T.A. and C.I.-S. purified and characterized antibodies. N.N. sequenced antibodies. T. Hino, T.A. and T.S. purified and crystallized the receptor/Fab-fragment complex. S.W., A.D.C. and S.I. performed data collection. T. Hino solved and refined the structure. T. Hino, S.I. and T.M. wrote the manuscript and all authors provide editorial input. The project was managed by T.K., T. Hamakubo, S.I. and T.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-11 with legends, Supplementary Tables 1-2 and a Supplementary Discussion. (PDF 3397 kb)
Rights and permissions
About this article
Cite this article
Hino, T., Arakawa, T., Iwanari, H. et al. G-protein-coupled receptor inactivation by an allosteric inverse-agonist antibody. Nature 482, 237–240 (2012). https://doi.org/10.1038/nature10750
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10750
This article is cited by
-
Current achievements, strategies, obstacles, and overcoming the challenges of the protein engineering in Pichia pastoris expression system
World Journal of Microbiology and Biotechnology (2024)
-
Crystal structure of adenosine A2A receptor in complex with clinical candidate Etrumadenant reveals unprecedented antagonist interaction
Communications Chemistry (2023)
-
Anti-nanodisc antibodies specifically capture nanodiscs and facilitate molecular interaction kinetics studies for membrane protein
Scientific Reports (2023)
-
Nanobody-enabled monitoring of kappa opioid receptor states
Nature Communications (2020)
-
Haloperidol bound D2 dopamine receptor structure inspired the discovery of subtype selective ligands
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.