Abstract
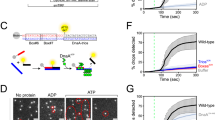
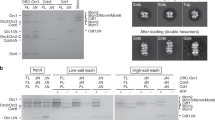
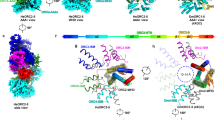
Many replication initiators form higher-order oligomers that process host replication origins to promote replisome formation. In addition to dedicated duplex-DNA-binding domains, cellular initiators possess AAA+ (ATPases associated with various cellular activities) elements that drive functions ranging from protein assembly to origin recognition. In bacteria, the AAA+ domain of the initiator DnaA has been proposed to assist in single-stranded DNA formation during origin melting. Here we show crystallographically and in solution that the ATP-dependent assembly of Aquifex aeolicus DnaA into a spiral oligomer creates a continuous surface that allows successive AAA+ domains to bind and extend single-stranded DNA segments. The mechanism of binding is unexpectedly similar to that of RecA, a homologous recombination factor, but it differs in that DnaA promotes a nucleic acid conformation that prevents pairing of a complementary strand. These findings, combined with strand-displacement assays, indicate that DnaA opens replication origins by a direct ATP-dependent stretching mechanism. Comparative studies reveal notable commonalities between the approach used by DnaA to engage DNA substrates and other, nucleic-acid-dependent, AAA+ systems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kornberg, A. & Baker, T. A. in DNA Replication (W. H. Freeman and Company, 1992)
Stillman, B. Origin recognition and the chromosome cycle. FEBS Lett. 579, 877–884 (2005)
Duderstadt, K. E. & Berger, J. M. AAA+ ATPases in the initiation of DNA replication. Crit. Rev. Biochem. Mol. Biol. 43, 163–187 (2008)
Lee, D. G. & Bell, S. P. ATPase switches controlling DNA replication initiation. Curr. Opin. Cell Biol. 12, 280–285 (2000)
Leipe, D. D., Koonin, E. V. & Aravind, L. Evolution and classification of P-loop kinases and related proteins. J. Mol. Biol. 333, 781–815 (2003)
Katayama, T., Ozaki, S., Keyamura, K. & Fujimitsu, K. Regulation of the replication cycle: conserved and diverse regulatory systems for DnaA and oriC. Nature Rev. Microbiol. 8, 163–170 (2010)
Kaguni, J. M. DnaA: controlling the initiation of bacterial DNA replication and more. Annu. Rev. Microbiol. 60, 351–371 (2006)
Leonard, A. C. & Grimwade, J. E. Regulating DnaA complex assembly: it is time to fill the gaps. Curr. Opin. Microbiol. 13, 766–772 (2010)
Fuller, R. S., Funnell, B. E. & Kornberg, A. The dnaA protein complex with the E. coli chromosomal replication origin (oriC) and other DNA sites. Cell 38, 889–900 (1984)
Funnell, B. E., Baker, T. A. & Kornberg, A. In vitro assembly of a prepriming complex at the origin of the Escherichia coli chromosome. J. Biol. Chem. 262, 10327–10334 (1987)
Crooke, E., Thresher, R., Hwang, D. S., Griffith, J. & Kornberg, A. Replicatively active complexes of DnaA protein and the Escherichia coli chromosomal origin observed in the electron microscope. J. Mol. Biol. 233, 16–24 (1993)
Bramhill, D. & Kornberg, A. Duplex opening by dnaA protein at novel sequences in initiation of replication at the origin of the E. coli chromosome. Cell 52, 743–755 (1988)
Kowalski, D. & Eddy, M. J. The DNA unwinding element: a novel, cis-acting component that facilitates opening of the Escherichia coli replication origin. EMBO J. 8, 4335–4344 (1989)
Erzberger, J. P., Mott, M. L. & Berger, J. M. Structural basis for ATP-dependent DnaA assembly and replication-origin remodeling. Nature Struct. Mol. Biol. 13, 676–683 (2006)
Ozaki, S. et al. A common mechanism for the ATP-DnaA-dependent formation of open complexes at the replication origin. J. Biol. Chem. 283, 8351–8362 (2008)
Speck, C. & Messer, W. Mechanism of origin unwinding: sequential binding of DnaA to double- and single-stranded DNA. EMBO J. 20, 1469–1476 (2001)
Sutton, M. D., Carr, K. M., Vicente, M. & Kaguni, J. M. Escherichia coli DnaA protein. The N-terminal domain and loading of DnaB helicase at the E. coli chromosomal origin. J. Biol. Chem. 273, 34255–34262 (1998)
Fang, L., Davey, M. J. & O’Donnell, M. Replisome assembly at oriC, the replication origin of E. coli, reveals an explanation for initiation sites outside an origin. Mol. Cell 4, 541–553 (1999)
Mott, M. L., Erzberger, J. P., Coons, M. M. & Berger, J. M. Structural synergy and molecular crosstalk between bacterial helicase loaders and replication initiators. Cell 135, 623–634 (2008)
Neuwald, A. F., Aravind, L., Spouge, J. L. & Koonin, E. V. AAA+: A class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes. Genome Res. 9, 27–43 (1999)
Ogura, T. & Wilkinson, A. J. AAA+ superfamily ATPases: common structure–diverse function. Genes Cells 6, 575–597 (2001)
Jeruzalmi, D., O’Donnell, M. & Kuriyan, J. Crystal structure of the processivity clamp loader gamma (γ) complex of E. coli DNA polymerase III. Cell 106, 429–441 (2001)
Speck, C., Chen, Z., Li, H. & Stillman, B. ATPase-dependent cooperative binding of ORC and Cdc6 to origin DNA. Nature Struct. Mol. Biol. 12, 965–971 (2005)
Clarey, M. G. et al. Nucleotide-dependent conformational changes in the DnaA-like core of the origin recognition complex. Nature Struct. Mol. Biol. 13, 684–690 (2006)
Duderstadt, K. E. et al. Origin remodeling and opening in bacteria rely on distinct assembly states of the DnaA initiator. J. Biol. Chem. 285, 28229–28239 (2010)
Dueber, E. L., Corn, J. E., Bell, S. D. & Berger, J. M. Replication origin recognition and deformation by a heterodimeric archaeal Orc1 complex. Science 317, 1210–1213 (2007)
Iyer, L. M., Leipe, D. D., Koonin, E. V. & Aravind, L. Evolutionary history and higher order classification of AAA+ ATPases. J. Struct. Biol. 146, 11–31 (2004)
Story, R. M., Weber, I. T. & Steitz, T. A. The structure of the E. coli recA protein monomer and polymer. Nature 355, 318–325 (1992)
Kowalczykowski, S. C. Initiation of genetic recombination and recombination-dependent replication. Trends Biochem. Sci. 25, 156–165 (2000)
Cox, M. M. Motoring along with the bacterial RecA protein. Nature Rev. Mol. Cell Biol. 8, 127–138 (2007)
Egelman, E. A common structural core in proteins active in DNA recombination and replication. Trends Biochem. Sci. 25, 179–182 (2000)
Conway, A. B. et al. Crystal structure of a Rad51 filament. Nature Struct. Mol. Biol. 11, 791–796 (2004)
Chen, Z., Yang, H. & Pavletich, N. P. Mechanism of homologous recombination from the RecA-ssDNA/dsDNA structures. Nature 453, 489–494 (2008)
Stasiak, A. & Di Capua, E. The helicity of DNA in complexes with recA protein. Nature 299, 185–186 (1982)
Galletto, R., Amitani, I., Baskin, R. J. & Kowalczykowski, S. C. Direct observation of individual RecA filaments assembling on single DNA molecules. Nature 443, 875–878 (2006)
Nishinaka, T., Ito, Y., Yokoyama, S. & Shibata, T. An extended DNA structure through deoxyribose-base stacking induced by RecA protein. Proc. Natl Acad. Sci. USA 94, 6623–6628 (1997)
Joo, C. et al. Real-time observation of RecA filament dynamics with single monomer resolution. Cell 126, 515–527 (2006)
Bianchi, M., Riboli, B. & Magni, G. E. coli recA protein possesses a strand separating activity on short duplex DNAs. EMBO J. 4, 3025–3030 (1985)
Thomsen, N. D. & Berger, J. M. Running in reverse: the structural basis for translocation polarity in hexameric helicases. Cell 139, 523–534 (2009)
Enemark, E. J. & Joshua-Tor, L. Mechanism of DNA translocation in a replicative hexameric helicase. Nature 442, 270–275 (2006)
Simonetta, K. R. et al. The mechanism of ATP-dependent primer-template recognition by a clamp loader complex. Cell 137, 659–671 (2009)
Gaudier, M., Schuwirth, B. S., Westcott, S. L. & Wigley, D. B. Structural basis of DNA replication origin recognition by an ORC protein. Science 317, 1213–1216 (2007)
Liu, X., Schuck, S. & Stenlund, A. Adjacent residues in the E1 initiator β-hairpin define different roles of the beta-hairpin in Ori melting, helicase loading, and helicase activity. Mol. Cell 25, 825–837 (2007)
MacDowell, A. A. et al. Suite of three protein crystallography beamlines with single superconducting bend magnet as the source. J. Synchrotron Radiat. 11, 447–455 (2004)
Otwinowski, Z. & Minor, W. in Methods in Enzymology 307–326 (Academic Press, 1997)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Terwilliger, T. C. Maximum-likelihood density modification. Acta Crystallogr. D 56, 965–972 (2000)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
DeLano, W. L. The PyMOL Molecular Graphics System (DeLano Scientific, 2002)
Ennifar, E., Carpentier, P., Ferrer, J. L., Walter, P. & Dumas, P. X-ray-induced debromination of nucleic acids at the Br K absorption edge and implications for MAD phasing. Acta Crystallogr. D 58, 1262–1268 (2002)
Felczak, M. M. & Kaguni, J. M. The box VII motif of Escherichia coli DnaA protein is required for DnaA oligomerization at the E. coli replication origin. J. Biol. Chem. 279, 51156–51162 (2004)
Mujumdar, R. B., Ernst, L. A., Mujumdar, S. R., Lewis, C. J. & Waggoner, A. S. Cyanine dye labeling reagents: sulfoindocyanine succinimidyl esters. Bioconjug. Chem. 4, 105–111 (1993)
Cantor, C. R. & Schimmel, P. R. in Biophysical Chemistry: Part II: Techniques for the study of biological structure and function 846 (W. H. Freeman and Company, 1980)
Magde, D., Wong, R. & Seybold, P. G. Fluorescence quantum yields and their relation to lifetimes of rhodamine 6G and fluorescein in nine solvents: improved absolute standards for quantum yields. Photochem. Photobiol. 75, 327–334 (2002)
Lakowicz, J. R. in Principles of Fluorescence Spectroscopy 2nd edn. (Kluwer/Plenum, 1999)
Clegg, R. M. Fluorescence resonance energy transfer and nucleic acids. Methods Enzymol. 211, 353–388 (1992)
Petsko, G. A. Chemistry and biology. Proc. Natl Acad. Sci. USA 97, 538–540 (2000)
Crisona, N. J. & Cozzarelli, N. R. Alteration of Escherichia coli topoisomerase IV conformation upon enzyme binding to positively supercoiled DNA. J. Biol. Chem. 281, 18927–18932 (2006)
Weber, K. & Osborn, M. The reliability of molecular weight determinations by dodecyl sulfate-polyacrylamide gel electrophoresis. J. Biol. Chem. 244, 4406–4412 (1969)
Acknowledgements
We would like to thank K. Drlica, J. Keck, T. Murray and the Berger laboratory for helpful comments, and M. M. Cox for his contribution of RecA protein. This work was supported by the NIGMS (GM071747) and the National Institute of Health Molecular Biophysics Training Grant T32 GM008295.
Author information
Authors and Affiliations
Contributions
K.E.D. and J.M.B. designed the experiments, analysed the data and wrote the paper. Protein purification, crystallization and ssDNA binding assays were performed by K.C. and K.E.D. K.E.D. performed the other experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Text, Supplementary References, Supplementary Tables 1-6 and Supplementary Figures 1-10 with legends. (PDF 16494 kb)
Rights and permissions
About this article
Cite this article
Duderstadt, K., Chuang, K. & Berger, J. DNA stretching by bacterial initiators promotes replication origin opening. Nature 478, 209–213 (2011). https://doi.org/10.1038/nature10455
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10455
This article is cited by
-
Archaeal Orc1 protein interacts with T-rich single-stranded DNA
BMC Research Notes (2021)
-
CcrZ is a pneumococcal spatiotemporal cell cycle regulator that interacts with FtsZ and controls DNA replication by modulating the activity of DnaA
Nature Microbiology (2021)
-
Structures, functions, and mechanisms of filament forming enzymes: a renaissance of enzyme filamentation
Biophysical Reviews (2019)
-
The mechanism of eukaryotic CMG helicase activation
Nature (2018)
-
The bacterial DnaA-trio replication origin element specifies single-stranded DNA initiator binding
Nature (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.