Abstract
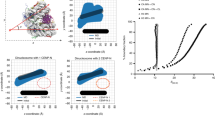
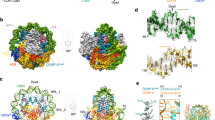
In eukaryotes, accurate chromosome segregation during mitosis and meiosis is coordinated by kinetochores, which are unique chromosomal sites for microtubule attachment1,2. Centromeres specify the kinetochore formation sites on individual chromosomes, and are epigenetically marked by the assembly of nucleosomes containing the centromere-specific histone H3 variant, CENP-A3,4,5,6,7,8,9,10,11,12. Although the underlying mechanism is unclear, centromere inheritance is probably dictated by the architecture of the centromeric nucleosome. Here we report the crystal structure of the human centromeric nucleosome containing CENP-A and its cognate α-satellite DNA derivative (147 base pairs). In the human CENP-A nucleosome, the DNA is wrapped around the histone octamer, consisting of two each of histones H2A, H2B, H4 and CENP-A, in a left-handed orientation. However, unlike the canonical H3 nucleosome, only the central 121 base pairs of the DNA are visible. The thirteen base pairs from both ends of the DNA are invisible in the crystal structure, and the αN helix of CENP-A is shorter than that of H3, which is known to be important for the orientation of the DNA ends in the canonical H3 nucleosome13. A structural comparison of the CENP-A and H3 nucleosomes revealed that CENP-A contains two extra amino acid residues (Arg 80 and Gly 81) in the loop 1 region, which is completely exposed to the solvent. Mutations of the CENP-A loop 1 residues reduced CENP-A retention at the centromeres in human cells. Therefore, the CENP-A loop 1 may function in stabilizing the centromeric chromatin containing CENP-A, possibly by providing a binding site for trans-acting factors. The structure provides the first atomic-resolution picture of the centromere-specific nucleosome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cheeseman, I. M. & Desai, A. Molecular architecture of the kinetochore-microtubule interface. Nature Rev. Mol. Cell Biol. 9, 33–46 (2008)
Santaguida, S. & Musacchio, A. The life and miracles of kinetocores. EMBO J. 28, 2511–2531 (2009)
Palmer, D. K., O'Day, K., Wener, M. H., Andrews, B. S. & Margolis, R. L. A 17-kD centromere protein (CENP-A) copurifies with nucleosome core particles and with histones. J. Cell Biol. 104, 805–815 (1987)
Stoler, S., Keith, K. C., Curnick, K. E. & Fitzgerald-Hayes, M. A mutation in CSE4, an essential gene encoding a novel chromatin-associated protein in yeast, causes chromosome nondisjunction and cell cycle arrest at mitosis. Genes Dev. 9, 573–586 (1995)
Meluh, P. B., Yang, P., Glowczewski, L., Koshland, D. & Smith, M. M. Cse4p is a component of the core centromere of Saccharomyces cerevisiae . Cell 94, 607–613 (1998)
Buchwitz, B. J., Ahmad, K., Moore, L. L., Roth, M. B. & Henikoff, S. A histone-H3-like protein in C. elegans . Nature 401, 547–548 (1999)
Henikoff, S., Ahmad, K., Platero, J. S. & van Steensel, B. Heterochromatic deposition of centromeric histone H3-like proteins. Proc. Natl Acad. Sci. USA 97, 716–721 (2000)
Howman, E. V. et al. Early disruption of centromeric chromatin organization in centromere protein A (Cenpa) null mice. Proc. Natl Acad. Sci. USA 97, 1148–1153 (2000)
Takahashi, K., Chen, E. S. & Yanagida, M. Requirement of Mis6 centromere connector for localizing a CENP-A-like protein in fission yeast. Science 288, 2215–2219 (2000)
Blower, M. D. & Karpen, G. H. The role of Drosophila CID in kinetochore formation, cell-cycle progression and heterochromatin interactions. Nature Cell Biol. 3, 730–739 (2001)
Oegema, K., Desai, A., Rybina, S., Kirkham, M. & Hyman, A. A. Functional analysis of kinetochore assembly in Caenorhabditis elegans . J. Cell Biol. 153, 1209–1226 (2001)
Régnier, V. et al. CENP-A is required for accurate chromosome segregation and sustained kinetochore association of BubR1. Mol. Cell. Biol. 25, 3967–3981 (2005)
Luger, K., Mäder, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 (1997)
Talbert, P. B. & Henikoff, S. Histone variants—ancient wrap artists of the epigenome. Nature Rev. Mol. Cell Biol. 11, 264–275 (2010)
Yoda, K. et al. Human centromere protein A (CENP-A) can replace histone H3 in nucleosome reconstitution in vitro . Proc. Natl Acad. Sci. USA 97, 7266–7271 (2000)
Tanaka, Y. et al. Human centromere protein B induces translational positioning of nucleosomes on alpha-satellite sequences. J. Biol. Chem. 280, 41609–41618 (2005)
Camahort, R. et al. Cse4 is part of an octameric nucleosome in budding yeast. Mol. Cell 35, 794–805 (2009)
Dalal, Y., Wang, H., Lindsay, S. & Henikoff, S. Tetrameric structure of centromeric nucleosomes in interphase Drosophila cells. PLoS Biol. 5, e218 (2007)
Dalal, Y., Furuyama, T., Vermaak, D. & Henikoff, S. Structure, dynamics, and evolution of centromeric nucleosomes. Proc. Natl Acad. Sci. USA 104, 15974–15981 (2007)
Furuyama, T. & Henikoff, S. Centromeric nucleosomes induce positive DNA supercoils. Cell 138, 104–113 (2009)
Tanaka, Y. et al. Expression and purification of recombinant human histones. Methods 33, 3–11 (2004)
Tsunaka, Y., Kajimura, N., Tate, S. & Morikawa, K. Alteration of the nucleosomal DNA path in the crystal structure of a human nucleosome core particle. Nucleic Acids Res. 33, 3424–3434 (2005)
Tachiwana, H. et al. Structural basis of instability of the nucleosome containing a testis-specific histone variant, human H3T. Proc. Natl Acad. Sci. USA 107, 10454–10459 (2010)
Tachiwana, H., Osakabe, A., Kimura, H. & Kurumizaka, H. Nucleosome formation with the testis-specific histone H3 variant, H3t, by human nucleosome assembly proteins in vitro . Nucleic Acids Res. 36, 2208–2218 (2008)
Osakabe, A. et al. Nucleosome formation activity of human somatic nuclear autoantigenic sperm protein (sNASP). J. Biol. Chem. 285, 11913–11921 (2010)
Sekulic, N., Bassett, E. A., Rogers, D. J. & Black, B. E. The structure of (CENP-A–H4)2 reveals physical features that mark centromeres. Nature 467, 347–351 (2010)
Conde e Silva, N. et al. CENP-A-containing nucleosomes: easier disassembly versus exclusive centromeric localization. J. Mol. Biol. 370, 555–573 (2007)
Kingston, I. J., Yung, J. S. & Singleton, M. R. Biophysical characterisation of the centromere-specific nucleosome from budding yeast. J. Biol. Chem. 286, 4021–4026 (2011)
Schalch, T., Duda, S., Sargent, D. F. & Richmond, T. J. X-ray structure of a tetranucleosome and its implications for the chromatin fibre. Nature 436, 138–141 (2005)
Masumoto, H., Masukata, H., Muro, Y., Nozaki, N. & Okazaki, T. A human centromere antigen (CENP-B) interacts with a short specific sequence in alphoid DNA, a human centromeric satellite. J. Cell Biol. 109, 1963–1973 (1989)
Dyer, P. N. et al. Reconstitution of nucleosome core particles from recombinant histones and DNA. Methods Enzymol. 375, 23–44 (2003)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Cryst. 30, 1022–1025 (1997)
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot . Acta Crystallogr. D 66, 486–501 (2010)
Fujisawa, T. et al. Small-angle X-ray scattering station at the SPring-8 RIKEN beamline. J. Appl. Cryst. 33, 797–800 (2000)
Svergun, D. I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Cryst. 25, 495–503 (1992)
Glatter, O. & Kratky, O. Small-angle X-ray Scattering (Academic Press, 1982)
Ando, S. et al. CENP-A, -B, and -C chromatin complex that contains the I-type α-satellite array constitutes the prekinetochore in HeLa cells. Mol. Cell. Biol. 22, 2229–2241 (2002)
Acknowledgements
We thank the beamline scientists, N. Shimizu, Y. Kawano, M. Makino and T. Hikima, for their assistance with data collection at the BL41XU and BL45XU beamlines of SPring-8. We also thank R. Matsumoto for technical assistance, K. Yoda for anti-CENP-C, and T. Fukagawa and Y. Hiraoka for discussions. This work was supported in part by Grants-in-Aid from the Japanese Society for the Promotion of Science (JSPS), and the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan. H.Ku. was also supported by the Waseda Research Institute for Science and Engineering.
Author information
Authors and Affiliations
Contributions
H.T., T.S., A.O. and Y.M. purified the histones and CENP-A, crystallized the CENP-A nucleosome, and performed biochemical analyses. H.T., W.K., K.S. and T.S. collected X-ray diffraction data, and H.T., W.K., and S.-Y.P. performed the structural analysis of the CENP-A nucleosome. H.T., A.O., Y.H.-T. and H.Ki. performed the cell biological experiments. T.O., H.T., W.K. and M.S. performed SAXS analysis. H.Ku. conceived, designed and supervised all of the work, and H.Ku., W.K. and H.T. wrote the paper. All of the authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-13 with legends, Supplementary Table 1 and additional references. (PDF 2314 kb)
Rights and permissions
About this article
Cite this article
Tachiwana, H., Kagawa, W., Shiga, T. et al. Crystal structure of the human centromeric nucleosome containing CENP-A. Nature 476, 232–235 (2011). https://doi.org/10.1038/nature10258
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10258
This article is cited by
-
In vitro co-expression chromatin assembly and remodeling platform for plant histone variants
Scientific Reports (2024)
-
CENP-A and CENP-B collaborate to create an open centromeric chromatin state
Nature Communications (2023)
-
Are extraordinary nucleosome structures more ordinary than we thought?
Chromosoma (2023)
-
CENP-N promotes the compaction of centromeric chromatin
Nature Structural & Molecular Biology (2022)
-
Chromatin, stacked at the centromere
Nature Structural & Molecular Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.