Abstract
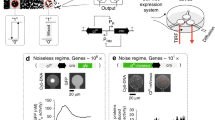
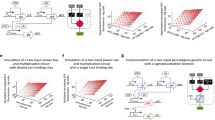
Computation underlies the organization of cells into higher-order structures, for example during development or the spatial association of bacteria in a biofilm1,2,3. Each cell performs a simple computational operation, but when combined with cell–cell communication, intricate patterns emerge. Here we study this process by combining a simple genetic circuit with quorum sensing to produce more complex computations in space. We construct a simple NOR logic gate in Escherichia coli by arranging two tandem promoters that function as inputs to drive the transcription of a repressor. The repressor inactivates a promoter that serves as the output. Individual colonies of E. coli carry the same NOR gate, but the inputs and outputs are wired to different orthogonal quorum-sensing ‘sender’ and ‘receiver’ devices4,5. The quorum molecules form the wires between gates. By arranging the colonies in different spatial configurations, all possible two-input gates are produced, including the difficult XOR and EQUALS functions. The response is strong and robust, with 5- to >300-fold changes between the ‘on’ and ‘off’ states. This work helps elucidate the design rules by which simple logic can be harnessed to produce diverse and complex calculations by rewiring communication between cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Neumann, J. V. The General and Logical Theory of Automata (Wiley, 1951)
Turing, A. M. The chemical basis of morphogenesis. 1953. Bull. Math. Biol. 52, 119–152 (discussion), 153–197 (1990)
Wolfram, S. A New Kind of Science 23–113 (Wolfram Media, 2002)
Brenner, K., Karig, D. K., Weiss, R. & Arnold, F. H. Engineered bidirectional communication mediates a consensus in a microbial biofilm consortium. Proc. Natl Acad. Sci. USA 104, 17300–17304 (2007)
Basu, S., Gerchman, Y., Collins, C. H., Arnold, F. H. & Weiss, R. A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005)
Li, F., Long, T., Lu, Y., Ouyang, Q. & Tang, C. The yeast cell-cycle network is robustly designed. Proc. Natl Acad. Sci. USA 101, 4781–4786 (2004)
Niklas, K. J. The bio-logic and machinery of plant morphogenesis. Am. J. Bot. 90, 515–525 (2003)
Morris, M. K., Saez-Rodriguez, J., Sorger, P. K. & Lauffenburger, D. A. Logic-based models for the analysis of cell signaling networks. Biochemistry 49, 3216–3224 (2010)
Mayo, A. E., Setty, Y., Shavit, S., Zaslaver, A. & Alon, U. Plasticity of the cis-regulatory input function of a gene. PLoS Biol. 4, e45 (2006)
Anderson, J. C., Voigt, C. A. & Arkin, A. P. Environmental signal integration by a modular AND gate. Mol. Syst. Biol. 3, 133 (2007)
Guet, C. C., Elowitz, M. B., Hsing, W. & Leibler, S. Combinatorial synthesis of genetic networks. Science 296, 1466–1470 (2002)
Rinaudo, K. et al. A universal RNAi-based logic evaluator that operates in mammalian cells. Nature Biotechnol. 25, 795–801 (2007)
Weber, W. et al. A synthetic mammalian gene circuit reveals antituberculosis compounds. Proc. Natl Acad. Sci. USA 105, 9994–9998 (2008)
Ellis, T., Wang, X. & Collins, J. J. Diversity-based, model-guided construction of synthetic gene networks with predicted functions. Nature Biotechnol. 27, 465–471 (2009)
Lou, C. et al. Synthesizing a novel genetic sequential logic circuit: a push-on push-off switch. Mol. Syst. Biol. 6, 350 (2010)
Tabor, J. J. et al. A synthetic genetic edge detection program. Cell 137, 1272–1281 (2009)
Friedland, A. E. et al. Synthetic gene networks that count. Science 324, 1199–1202 (2009)
Tan, C., Marguet, P. & You, L. Emergent bistability by a growth-modulating positive feedback circuit. Nature Chem. Biol. 5, 842–848 (2009)
Scharle, T. W. Axiomatization of propositional calculus with Sheffer functors. Notre Dame J. Formal Logic 6, 209–217 (1965)
Yokobayashi, Y., Weiss, R. & Arnold, F. H. Directed evolution of a genetic circuit. Proc. Natl Acad. Sci. USA 99, 16587–16591 (2002)
Sneppen, K. et al. A mathematical model for transcriptional interference by RNA polymerase traffic in Escherichia coli . J. Mol. Biol. 346, 399–409 (2005)
Pesci, E. C., Pearson, J. P., Seed, P. C. & Iglewski, B. H. Regulation of las and rhl quorum sensing in Pseudomonas aeruginosa. J. Bacteriol. 179, 3127–3132 (1997)
Karig, D. & Weiss, R. Signal-amplifying genetic circuit enables in vivo observation of weak promoter activation in the Rhl quorum sensing system. Biotechnol. Bioeng. 89, 709–718 (2005)
Rosenfeld, N., Young, J. W., Alon, U., Swain, P. S. & Elowitz, M. B. Gene regulation at the single-cell level. Science 307, 1962–1965 (2005)
Pedraza, J. M. & van Oudenaarden, A. Noise propagation in gene networks. Science 307, 1965–1969 (2005)
Katz, R. H. & Borriello, G. Contemporary Logic Design 141–146 (Prentice Hall, 1994)
Danino, T., Mondragon-Palomino, O., Tsimring, L. & Hasty, J. A synchronized quorum of genetic clocks. Nature 463, 326–330 (2010)
Clancy, K. & Voigt, C. A. Programming cells: towards an automated ‘genetic compiler’. Curr. Opin. Biotechnol. 21, 572–581 (2010)
Ilachinski, A. Cellular Automata: A Discrete Universe 1–18 (World Scientific, 2001)
Raju, B. S. & Mullick, S. K. Programmable cellular arrays. Int. J. Control 14, 1041–1061 (1971)
Acknowledgements
We thank W. Mulyasasmita and K. Temme for critical discussions. This work was supported by the National Science Foundation (SynBERC, NSF#0943385 and NSF Sandpit CCF-0943385) and the Office of Naval Research.
Author information
Authors and Affiliations
Contributions
A.T. designed and performed the experiments, analysed the data and wrote the manuscript. J.J.T. designed experiments and edited the manuscript. C.A.V. designed experiments, analysed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1-S11 with legends, Supplementary Table S1-S5, Supplementary Discussions, a List of Strains, Plasmid Maps, and Supplementary References. (PDF 1665 kb)
Rights and permissions
About this article
Cite this article
Tamsir, A., Tabor, J. & Voigt, C. Robust multicellular computing using genetically encoded NOR gates and chemical ‘wires’. Nature 469, 212–215 (2011). https://doi.org/10.1038/nature09565
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09565
This article is cited by
-
Dynamic cybergenetic control of bacterial co-culture composition via optogenetic feedback
Nature Communications (2022)
-
Deduction of signaling mechanisms from cellular responses to multiple cues
npj Systems Biology and Applications (2022)
-
Transcriptional programming in a Bacteroides consortium
Nature Communications (2022)
-
Synthetic neuromorphic computing in living cells
Nature Communications (2022)
-
Distributed computation with continual population growth
Distributed Computing (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.