Abstract
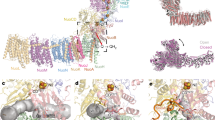
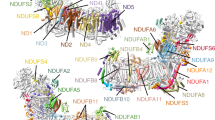
Complex I is the first enzyme of the respiratory chain and has a central role in cellular energy production, coupling electron transfer between NADH and quinone to proton translocation by an unknown mechanism. Dysfunction of complex I has been implicated in many human neurodegenerative diseases. We have determined the structure of its hydrophilic domain previously. Here, we report the α-helical structure of the membrane domain of complex I from Escherichia coli at 3.9 Å resolution. The antiporter-like subunits NuoL/M/N each contain 14 conserved transmembrane (TM) helices. Two of them are discontinuous, as in some transporters. Unexpectedly, subunit NuoL also contains a 110-Å long amphipathic α-helix, spanning almost the entire length of the domain. Furthermore, we have determined the structure of the entire complex I from Thermus thermophilus at 4.5 Å resolution. The L-shaped assembly consists of the α-helical model for the membrane domain, with 63 TM helices, and the known structure of the hydrophilic domain. The architecture of the complex provides strong clues about the coupling mechanism: the conformational changes at the interface of the two main domains may drive the long amphipathic α-helix of NuoL in a piston-like motion, tilting nearby discontinuous TM helices, resulting in proton translocation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Walker, J. E. The NADH:ubiquinone oxidoreductase (complex I) of respiratory chains. Q. Rev. Biophys. 25, 253–324 (1992)
Yagi, T. & Matsuno-Yagi, A. The proton-translocating NADH-quinone oxidoreductase in the respiratory chain: the secret unlocked. Biochemistry 42, 2266–2274 (2003)
Brandt, U. Energy converting NADH:quinone oxidoreductase (complex I). Annu. Rev. Biochem. 75, 69–92 (2006)
Ohnishi, T. Iron-sulfur clusters/semiquinones in complex I. Biochim. Biophys. Acta 1364, 186–206 (1998)
Sazanov, L. A. Respiratory complex I: mechanistic and structural insights provided by the crystal structure of the hydrophilic domain. Biochemistry 46, 2275–2288 (2007)
Schapira, A. H. Human complex I defects in neurodegenerative diseases. Biochim. Biophys. Acta 1364, 261–270 (1998)
Dawson, T. M. & Dawson, V. L. Molecular pathways of neurodegeneration in Parkinson’s disease. Science 302, 819–822 (2003)
Balaban, R. S., Nemoto, S. & Finkel, T. Mitochondria, oxidants, and aging. Cell 120, 483–495 (2005)
Carroll, J. et al. Bovine complex I is a complex of 45 different subunits. J. Biol. Chem. 281, 32724–32727 (2006)
Clason, T. et al. The structure of eukaryotic and prokaryotic complex I. J. Struct. Biol. 169, 81–88 (2010)
Morgan, D. J. & Sazanov, L. A. Three-dimensional structure of respiratory complex I from Escherichia coli in ice in the presence of nucleotides. Biochim. Biophys. Acta 1777, 711–718 (2008)
Sazanov, L. A. & Hinchliffe, P. Structure of the hydrophilic domain of respiratory complex I from Thermus thermophilus . Science 311, 1430–1436 (2006)
Berrisford, J. M. & Sazanov, L. A. Structural basis for the mechanism of respiratory complex I. J. Biol. Chem. 284, 29773–29783 (2009)
Fearnley, I. M. & Walker, J. E. Conservation of sequences of subunits of mitochondrial complex I and their relationships with other proteins. Biochim. Biophys. Acta 1140, 105–134 (1992)
Mathiesen, C. & Hagerhall, C. Transmembrane topology of the NuoL, M and N subunits of NADH:quinone oxidoreductase and their homologues among membrane-bound hydrogenases and bona fide antiporters. Biochim. Biophys. Acta 1556, 121–132 (2002)
Friedrich, T. Complex I: a chimaera of a redox and conformation-driven proton pump? J. Bioenerg. Biomembr. 33, 169–177 (2001)
Sazanov, L. A., Carroll, J., Holt, P., Toime, L. & Fearnley, I. M. A role for native lipids in the stabilization and two-dimensional crystallization of the Escherichia coli NADH-ubiquinone oxidoreductase (complex I). J. Biol. Chem. 278, 19483–19491 (2003)
Guénebaut, V., Schlitt, A., Weiss, H., Leonard, K. & Friedrich, T. Consistent structure between bacterial and mitochondrial NADH:ubiquinone oxidoreductase (complex I). J. Mol. Biol. 276, 105–112 (1998)
Baranova, E. A., Holt, P. J. & Sazanov, L. A. Projection structure of the membrane domain of Escherichia coli respiratory complex I at 8 Å resolution. J. Mol. Biol. 366, 140–154 (2007)
Kao, M. C., Di Bernardo, S., Matsuno-Yagi, A. & Yagi, T. Characterization of the membrane domain Nqo11 subunit of the proton-translocating NADH-quinone oxidoreductase of Paracoccus denitrificans . Biochemistry 41, 4377–4384 (2002)
Kao, M. C., Di Bernardo, S., Matsuno-Yagi, A. & Yagi, T. Characterization and topology of the membrane domain Nqo10 subunit of the proton-translocating NADH-quinone oxidoreductase of Paracoccus denitrificans . Biochemistry 42, 4534–4543 (2003)
Bernardo, S. D., Yano, T. & Yagi, T. Exploring the membrane domain of the reduced nicotinamide adenine dinucleotide-quinone oxidoreductase of Paracoccus denitrificans: characterization of the NQO7 subunit. Biochemistry 39, 9411–9418 (2000)
Mamedova, A. A., Holt, P. J., Carroll, J. & Sazanov, L. A. Substrate-induced conformational change in bacterial complex I. J. Biol. Chem. 279, 23830–23836 (2004)
Screpanti, E. & Hunte, C. Discontinuous membrane helices in transport proteins and their correlation with function. J. Struct. Biol. 159, 261–267 (2007)
Torres-Bacete, J., Sinha, P. K., Castro-Guerrero, N., Matsuno-Yagi, A. & Yagi, T. Features of subunit NuoM (ND4) in Escherichia coli NDH-1: topology and implication of conserved Glu144 for coupling site 1. J. Biol. Chem. 284, 33062–33069 (2009)
Holt, P. J., Morgan, D. J. & Sazanov, L. A. The location of NuoL and NuoM subunits in the membrane domain of the Escherichia coli complex I: implications for the mechanism of proton pumping. J. Biol. Chem. 278, 43114–43120 (2003)
Baranova, E. A., Morgan, D. J. & Sazanov, L. A. Single particle analysis confirms distal location of subunits NuoL and NuoM in Escherichia coli complex I. J. Struct. Biol. 159, 238–242 (2007)
Kao, M. C., Nakamaru-Ogiso, E., Matsuno-Yagi, A. & Yagi, T. Characterization of the membrane domain subunit NuoK (ND4L) of the NADH-quinone oxidoreductase from Escherichia coli . Biochemistry 44, 9545–9554 (2005)
Roth, R. & Hagerhall, C. Transmembrane orientation and topology of the NADH:quinone oxidoreductase putative quinone binding subunit NuoH. Biochim. Biophys. Acta 1504, 352–362 (2001)
Kao, M. C., Matsuno-Yagi, A. & Yagi, T. Subunit proximity in the H+-translocating NADH-quinone oxidoreductase probed by zero-length cross-linking. Biochemistry 43, 3750–3755 (2004)
Murai, M., Sekiguchi, K., Nishioka, T. & Miyoshi, H. Characterization of the inhibitor binding site in mitochondrial NADH-ubiquinone oxidoreductase by photoaffinity labeling using a quinazoline-type inhibitor. Biochemistry 48, 688–698 (2009)
Sekiguchi, K., Murai, M. & Miyoshi, H. Exploring the binding site of acetogenin in the ND1 subunit of bovine mitochondrial complex I. Biochim. Biophys. Acta 1787, 1106–1111 (2009)
Page, C. C., Moser, C. C., Chen, X. & Dutton, P. L. Natural engineering principles of electron tunnelling in biological oxidation–reduction. Nature 402, 47–52 (1999)
Yano, T., Dunham, W. R. & Ohnishi, T. Characterization of the ΔμH+-sensitive ubisemiquinone species (SQNf) and the interaction with cluster N2: new insight into the energy-coupled electron transfer in complex I. Biochemistry 44, 1744–1754 (2005)
Nakamaru-Ogiso, E., Sakamoto, K., Matsuno-Yagi, A., Miyoshi, H. & Yagi, T. The ND5 subunit was labeled by a photoaffinity analogue of fenpyroximate in bovine mitochondrial complex I. Biochemistry 42, 746–754 (2003)
Gong, X. et al. The ubiquinone-binding site in NADH:ubiquinone oxidoreductase from Escherichia coli . J. Biol. Chem. 278, 25731–25737 (2003)
Berrisford, J. M., Thompson, C. J. & Sazanov, L. A. Chemical and NADH-induced, ROS-dependent, cross-linking between subunits of complex I from Escherichia coli and Thermus thermophilus . Biochemistry 47, 10262–10270 (2008)
Belogrudov, G. & Hatefi, Y. Catalytic sector of complex I (NADH:ubiquinone oxidoreductase): subunit stoichiometry and substrate-induced conformation changes. Biochemistry 33, 4571–4576 (1994)
Gondal, J. A. & Anderson, W. M. The molecular morphology of bovine heart mitochondrial NADH-ubiquinone reductase. Native disulfide-linked subunits and rotenone-induced conformational changes. J. Biol. Chem. 260, 12690–12694 (1985)
Verkhovskaya, M. L., Belevich, N., Euro, L., Wikstrom, M. & Verkhovsky, M. I. Real-time electron transfer in respiratory complex I. Proc. Natl Acad. Sci. USA 105, 3763–3767 (2008)
Euro, L., Belevich, G., Verkhovsky, M. I., Wikstrom, M. & Verkhovskaya, M. Conserved lysine residues of the membrane subunit NuoM are involved in energy conversion by the proton-pumping NADH:ubiquinone oxidoreductase (Complex I). Biochim. Biophys. Acta 1777, 1166–1172 (2008)
Torres-Bacete, J., Nakamaru-Ogiso, E., Matsuno-Yagi, A. & Yagi, T. Characterization of the NuoM (ND4) subunit in Escherichia coli NDH-1: conserved charged residues essential for energy-coupled activities. J. Biol. Chem. 282, 36914–36922 (2007)
Kervinen, M., Patsi, J., Finel, M. & Hassinen, I. E. A pair of membrane-embedded acidic residues in the NuoK subunit of Escherichia coli NDH-1, a counterpart of the ND4L subunit of the mitochondrial complex I, are required for high ubiquinone reductase activity. Biochemistry 43, 773–781 (2004)
Kajiyama, Y., Otagiri, M., Sekiguchi, J., Kudo, T. & Kosono, S. The MrpA, MrpB and MrpD subunits of the Mrp antiporter complex in Bacillus subtilis contain membrane-embedded and essential acidic residues. Microbiology 155, 2137–2147 (2009)
Sazanov, L. A., Carroll, J., Holt, P., Toime, L. & Fearnley, I. M. A role for native lipids in the stabilization and two-dimensional crystallization of the Escherichia coli NADH-ubiquinone oxidoreductase (Complex I). J. Biol. Chem. 278, 19483–19491 (2003)
Hinchliffe, P., Carroll, J. & Sazanov, L. A. Identification of a novel subunit of respiratory complex I from Thermus thermophilus . Biochemistry 45, 4413–4420 (2006)
CCP4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Schneider, T. R. & Sheldrick, G. M. Substructure solution with SHELXD . Acta Crystallogr. D 58, 1772–1779 (2002)
de La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007)
Jones, T. A. & Kjeldgaard, M. Electron-density map interpretation. Methods Enzymol. 277, 173–208 (1997)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Rudolph, M. G., Wingren, C., Crowley, M. P., Chien, Y. H. & Wilson, I. A. Combined pseudo-merohedral twinning, non-crystallographic symmetry and pseudo-translation in a monoclinic crystal form of the γδ T-cell ligand T10. Acta Crystallogr. D 60, 656–664 (2004)
Berrisford, J. M. & Sazanov, L. A. Structural basis for the mechanism of respiratory complex I. J. Biol. Chem. 284, 29773–29783 (2009)
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007)
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001)
McGuffin, L. J., Bryson, K. & Jones, D. T. The PSIPRED protein structure prediction server. Bioinformatics 16, 404–405 (2000)
Rost, B., Yachdav, G. & Liu, J. The PredictProtein server. Nucleic Acids Res. 32, W321–W326 (2004)
Acknowledgements
This work was funded by the Medical Research Council. We thank the European Synchrotron Radiation Facility and the Swiss Light Source for provision of synchrotron radiation facilities. We are grateful to the staff of beamlines ID23, ID29 (ESRF, Grenoble) and X06SA (Swiss Light Source, Villigen) for assistance. We thank J. E. Walker and A. Leslie for discussions of the manuscript.
Author information
Authors and Affiliations
Contributions
R.G.E. purified and crystallized the membrane domain of E. coli complex I; R.B. purified and crystallized intact T. thermophilus complex I; R.G.E. and R.B. collected and analysed X-ray data; L.A.S. designed the project, analysed data and wrote the manuscript, with contributions from R.G.E. and R.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
This file contains Supplementary Figures 1-7 with legends, Supplementary Tables 1-2 and References. (PDF 2039 kb)
Rights and permissions
About this article
Cite this article
Efremov, R., Baradaran, R. & Sazanov, L. The architecture of respiratory complex I. Nature 465, 441–445 (2010). https://doi.org/10.1038/nature09066
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature09066
This article is cited by
-
N1-methylation of adenosine (m1A) in ND5 mRNA leads to complex I dysfunction in Alzheimer’s disease
Molecular Psychiatry (2024)
-
Strategies of chemolithoautotrophs adapting to high temperature and extremely acidic conditions in a shallow hydrothermal ecosystem
Microbiome (2023)
-
Doping of molecular semiconductors through proton-coupled electron transfer
Nature (2023)
-
The coupling mechanism of mammalian mitochondrial complex I
Nature Structural & Molecular Biology (2022)
-
Mechanisms of membrane protein crystallization in ‘bicelles’
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.