Abstract
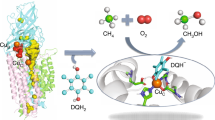
Vast world reserves of methane gas are underutilized as a feedstock for the production of liquid fuels and chemicals owing to the lack of economical and sustainable strategies for the selective oxidation of methane to methanol1. Current processes to activate the strong C–H bond (104 kcal mol-1) in methane require high temperatures, are costly and inefficient, and produce waste2. In nature, methanotrophic bacteria perform this reaction under ambient conditions using metalloenzymes called methane monooxygenases (MMOs). MMOs thus provide the optimal model for an efficient, environmentally sound catalyst3. There are two types of MMO. Soluble MMO (sMMO) is expressed by several strains of methanotroph under copper-limited conditions and oxidizes methane with a well-characterized catalytic di-iron centre4. Particulate MMO (pMMO) is an integral membrane metalloenzyme produced by all methanotrophs and is composed of three subunits, pmoA, pmoB and pmoC, arranged in a trimeric α3β3γ3 complex5. Despite 20 years of research and the availability of two crystal structures, the metal composition and location of the pMMO metal active site are not known. Here we show that pMMO activity is dependent on copper, not iron, and that the copper active site is located in the soluble domains of the pmoB subunit rather than within the membrane. Recombinant soluble fragments of pmoB (spmoB) bind copper and have propylene and methane oxidation activities. Disruption of each copper centre in spmoB by mutagenesis indicates that the active site is a dicopper centre. These findings help resolve the pMMO controversy and provide a promising new approach to developing environmentally friendly C–H oxidation catalysts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Hermans, I., Spier, E. S., Neuenschwander, U., Turra, N. & Baiker, A. Selective oxidation catalysis: opportunities and challenges. Top. Catal. 52, 1162–1174 (2009)
Arakawa, H. et al. Catalysis research of relevance to carbon management: progress, challenges, and opportunities. Chem. Rev. 101, 953–996 (2001)
Que, L. & Tolman, W. B. Biologically inspired oxidation catalysis. Nature 455, 333–340 (2008)
Merkx, M. et al. Dioxygen activation and methane hydroxylation by soluble methane monooxygenase: a tale of two irons and three proteins. Angew. Chem. Int. Ed. 40, 2782–2807 (2001)
Hakemian, A. S. & Rosenzweig, A. C. The biochemistry of methane oxidation. Annu. Rev. Biochem. 76, 223–241 (2007)
Lieberman, R. L. & Rosenzweig, A. C. Crystal structure of a membrane-bound metalloenzyme that catalyses the biological oxidation of methane. Nature 434, 177–182 (2005)
Balasubramanian, R. & Rosenzweig, A. C. Structural and mechanistic insights into methane oxidation by particulate methane monooxygenase. Acc. Chem. Res. 40, 573–580 (2007)
Lieberman, R. L. et al. Characterization of the particulate methane monooxygenase metal centers in multiple redox states by X-ray absorption spectroscopy. Inorg. Chem. 45, 8372–8381 (2006)
Lieberman, R. L. et al. Purified particulate methane monooxygenase from Methylococcus capsulatus (Bath) is a dimer with both mononuclear copper and a copper-containing cluster. Proc. Natl Acad. Sci. USA 100, 3820–3825 (2003)
Hakemian, A. S. et al. The metal centers of particulate methane monooxygenase from Methylosinus trichosporium OB3b. Biochemistry 47, 6793–6801 (2008)
Rosenzweig, A. C. The metal centres of particulate methane monooxygenase. Biochem. Soc. Trans. 36, 1134–1137 (2008)
Chan, S. I. & Yu, S. S. F. Controlled oxidation of hydrocarbons by the membrane-bound methane monooxygenase: the case for a tricopper cluster. Acc. Chem. Res. 41, 969–979 (2008)
Chen, K. H.-C. et al. The copper clusters in the particulate methane monooxygenase (pMMO) from Methylococcus capsulatus (Bath). J. Chin. Chem. Soc. 51, 1081–1098 (2004)
Nguyen, H.-H. T. et al. X-ray absorption and EPR studies on the copper ions associated with the particulate methane monooxygenase from Methylococcus capsulatus (Bath). Cu(I) ions and their implications. J. Am. Chem. Soc. 118, 12766–12776 (1996)
Chan, S. I. et al. Redox potentiometry studies of particulate methane monooxygenase: support for a trinuclear copper cluster active site. Angew. Chem. Int. Ed. 46, 1992–1994 (2007)
Yu, S. S. F. et al. The C-terminal aqueous-exposed domain of the 45 kDa subunit of the particulate methane monooxygenase in Methylococcus capsulatus (Bath) is a Cu(I) sponge. Biochemistry 46, 13762–13774 (2007)
Martinho, M. et al. Mössbauer studies of the membrane-associated methane monooxygenase from Methylococcus capsulatus Bath: evidence for a diiron center. J. Am. Chem. Soc. 129, 15783–15785 (2007)
Choi, D. W. et al. The membrane-associated methane monooxygenase pMMO and pMMO-NADH:quinone oxidoreductase complex from Methylococcus capsulatus Bath. J. Bacteriol. 185, 5755–5764 (2003)
Kitmitto, A., Myronova, N., Basu, P. & Dalton, H. Characterization and structural analysis of an active particulate methane monooxygenase trimer from Methylococcus capsulatus (Bath). Biochemistry 44, 10954–10965 (2005)
Yu, S. S.-F. et al. Production of high-quality particulate methane monooxygenase in high yields from Methylococcus capsulatus (Bath) with a hollow-fiber membrane bioreactor. J. Bacteriol. 185, 5915–5924 (2003)
Miyaji, A., Suzuki, M., Baba, T., Kamachi, T. & Okura, I. Hydrogen peroxide as an effecter on the inactivation of particulate methane monooxygenase under aerobic conditions. J. Mol. Catal. B 57, 211–215 (2009)
Kau, L.-S., Spira-Solomon, D. J., Penner-Hahn, J. E., Hodgson, K. O. & Solomon, E. I. X-ray absorption edge determination of the oxidation state and coordination number of copper. Application to the type 3 site in Rhus vernicifera laccase and its reaction with oxygen. J. Am. Chem. Soc. 109, 6433–6442 (1987)
Yatsunyk, L. A. & Rosenzweig, A. C. Copper(I) binding and transfer by the N-terminus of the Wilson disease protein. J. Biol. Chem. 282, 8622–8631 (2007)
Prior, S. D. & Dalton, H. Acetylene as a suicide substrate and active site probe for methane monooxygenase from Methylococcus capsulatus (Bath). FEMS Microbiol. Lett. 29, 105–109 (1985)
Zahn, J. A. & DiSpirito, A. A. Membrane-associated methane monooxygenase from Methylococcus capsulatus (Bath). J. Bacteriol. 178, 1018–1029 (1996)
Shiota, Y. & Yoshizawa, K. Comparison of the reactivity of bis(μ-oxo)CuIICuIII and CuIIICuIII species to methane. Inorg. Chem. 48, 838–845 (2009)
Woertink, J. S. et al. A [Cu2O]2+ core in Cu-ZSM-5, the active site in the oxidation of methane to methanol. Proc. Natl Acad. Sci. USA 106, 18908–18913 (2009)
Miller, K. W., Hammond, L. & Porter, E. G. The solubility of hydrocarbon gases in lipid bilayers. Chem. Phys. Lipids 20, 229–241 (1977)
Batliwala, H., Somasundaram, T., Uzgiris, E. E. & Makowski, L. Methane-induced hemolysis of human erythrocytes. Biochem. J. 307, 433–438 (1995)
Myronova, N., Kitmitto, A., Collins, R. F., Miyaji, A. & Dalton, H. Three-dimensional structure determination of a protein supercomplex that oxidizes methane to formaldehyde in Methylococcus capsulatus (Bath). Biochemistry 45, 11905–11914 (2006)
Stookey, L. L. Ferrozine - a new spectrophotometric reagent for iron. Anal. Chem. 42, 779–781 (1970)
George, G. N., George, S. J. & Pickering, I. J. EXAFSPAK 〈http://www-ssrl.slac.stanford.edu/~george/exafspak/exafs.htm〉 (2001)
Ankudinov, A. L. & Rehr, J. J. Relativistic calculations of spin-dependent X-ray absorption spectra. Phys. Rev. B 56, R1712–R1716 (1997)
Riggs-Gelasco, P. J., Stemmler, T. L. & Penner-Hahn, J. E. XAFS of dinuclear metal sites in proteins and model compounds. Coord. Chem. Rev. 144, 245–286 (1995)
Acknowledgements
This work was supported by US National Institutes of Health (NIH) grants GM070473 (A.C.R.) and DK068139 (T.L.S.). Portions of this research were carried out at the Stanford Synchrotron Radiation Lightsource (SSRL). The SSRL is a national user facility operated by Stanford University on behalf of the US Department of Energy, Office of Basic Energy Sciences. The SSRL Structural Molecular Biology Program is supported by the Department of Energy, Office of Biological and Environmental Research, and by the NIH, National Center for Research Resources, Biomedical Technology Program.
Author Contributions R.B., S.M.S., S.R. and L.A.Y. performed experiments. R.B., S.M.S., S.R., T.L.S. and A.C.R. contributed to experimental design, data analysis and manuscript preparation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables S1-S3 and Supplementary Figures S1-S5 with legends. (PDF 1970 kb)
Rights and permissions
About this article
Cite this article
Balasubramanian, R., Smith, S., Rawat, S. et al. Oxidation of methane by a biological dicopper centre. Nature 465, 115–119 (2010). https://doi.org/10.1038/nature08992
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08992
This article is cited by
-
Regulating atomic Fe–Rh site distance for efficient oxygen reduction reaction
Science China Chemistry (2024)
-
In-situ X-ray absorption spectroscopy study on the formation of superoxides on CuxSny composite catalysts enables the direct synthesis of catechol
Reaction Kinetics, Mechanisms and Catalysis (2023)
-
Microbial oxidation of atmospheric trace gases
Nature Reviews Microbiology (2022)
-
Selective Oxidation of Methane into Methanol Under Mild Conditions
Chemical Research in Chinese Universities (2022)
-
The effect of Zn doping on active Cu species and its location of Cu-exchanged mordenite for the stepwise oxidation of methane to methanol
Korean Journal of Chemical Engineering (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.