Abstract
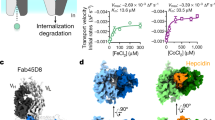
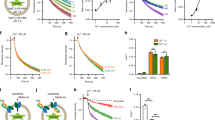
Cobalamin (Cbl, vitamin B12) is a bacterial organic compound and an essential coenzyme in mammals, which take it up from the diet. This occurs by the combined action of the gastric intrinsic factor (IF) and the ileal endocytic cubam receptor formed by the 460-kilodalton (kDa) protein cubilin and the 45-kDa transmembrane protein amnionless1,2. Loss of function of any of these proteins ultimately leads to Cbl deficiency in man3,4. Here we present the crystal structure of the complex between IF–Cbl and the cubilin IF–Cbl-binding-region (CUB5–8)5 determined at 3.3 Å resolution. The structure provides insight into how several CUB (for ‘complement C1r/C1s, Uegf, Bmp1’) domains collectively function as modular ligand-binding regions, and how two distant CUB domains embrace the Cbl molecule by binding the two IF domains in a Ca2+-dependent manner. This dual-point model provides a probable explanation of how Cbl indirectly induces ligand–receptor coupling. Finally, the comparison of Ca2+-binding CUB domains and the low-density lipoprotein (LDL) receptor-type A modules suggests that the electrostatic pairing of a basic ligand arginine/lysine residue with Ca2+-coordinating acidic aspartates/glutamates is a common theme of Ca2+-dependent ligand–receptor interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Birn, H. et al. Characterization of an epithelial approximately 460-kDa protein that facilitates endocytosis of intrinsic factor-vitamin B12 and binds receptor-associated protein. J. Biol. Chem. 272, 26497–26504 (1997)
Fyfe, J. C. et al. The functional cobalamin (vitamin B12)-intrinsic factor receptor is a novel complex of cubilin and amnionless. Blood 103, 1573–1579 (2004)
Tanner, S. M. et al. Genetically heterogeneous selective intestinal malabsorption of vitamin B12: founder effects, consanguinity, and high clinical awareness explain aggregations in Scandinavia and the Middle East. Hum. Mutat. 23, 327–333 (2004)
Tanner, S. M. et al. Hereditary juvenile cobalamin deficiency caused by mutations in the intrinsic factor gene. Proc. Natl Acad. Sci. USA 102, 4130–4133 (2005)
Kristiansen, M. et al. Molecular dissection of the intrinsic factor-vitamin B12 receptor, cubilin, discloses regions important for membrane association and ligand binding. J. Biol. Chem. 274, 20540–20544 (1999)
Christensen, E. I., Verroust, P. J. & Nielsen, R. Receptor-mediated endocytosis in renal proximal tubule. Pflugers Arch. 458, 1039–1048 (2009)
Mathews, F. S. et al. Crystal structure of human intrinsic factor: cobalamin complex at 2.6-A resolution. Proc. Natl Acad. Sci. USA 104, 17311–17316 (2007)
Fedosov, S. N. et al. Assembly of the intrinsic factor domains and oligomerization of the protein in the presence of cobalamin. Biochemistry 43, 15095–15102 (2004)
Kristiansen, M. et al. Cubilin P1297L mutation associated with hereditary megaloblastic anemia 1 causes impaired recognition of intrinsic factor-vitamin B12 by cubilin. Blood 96, 405–409 (2000)
Aminoff, M. et al. Mutations in CUBN, encoding the intrinsic factor-vitamin B12 receptor, cubilin, cause hereditary megaloblastic anaemia 1. Nature Genet. 21, 309–313 (1999)
Lindblom, A. et al. The intrinsic factor-vitamin B12 receptor, cubilin, is assembled into trimers via a coiled-coil α-helix. J. Biol. Chem. 274, 6374–6380 (1999)
Blanc, G. et al. Insights into how CUB domains can exert specific functions while sharing a common fold: conserved and specific features of the CUB1 domain contribute to the molecular basis of procollagen C-proteinase enhancer-1 activity. J. Biol. Chem. 282, 16924–16933 (2007)
Gregory, L. A., Thielens, N. M., Arlaud, G. J., Fontecilla-Camps, J. C. & Gaboriaud, C. X-ray structure of the Ca2+-binding interaction domain of C1s. Insights into the assembly of the C1 complex of complement. J. Biol. Chem. 278, 32157–32164 (2003)
Teillet, F. et al. Crystal structure of the CUB1-EGF-CUB2 domain of human MASP-1/3 and identification of its interaction sites with mannan-binding lectin and ficolins. J. Biol. Chem. 283, 25715–25724 (2008)
Bally, I. et al. Identification of the C1q binding sites of human C1r and C1s. A refined three-dimensional model of the C1 complex of complement. J. Biol. Chem. 284, 19340–19348 (2009)
Kristiansen, M. et al. Identification of the haemoglobin scavenger receptor. Nature 409, 198–201 (2001)
Blacklow, S. C. Versatility in ligand recognition by LDL receptor family proteins: advances and frontiers. Curr. Opin. Struct. Biol. 17, 419–426 (2007)
Quadros, E. V., Nakayama, Y. & Sequeira, J. M. The protein and the gene encoding the receptor for the cellular uptake of transcobalamin-bound cobalamin. Blood 113, 186–192 (2009)
Fisher, C., Beglova, N. & Blacklow, S. C. Structure of an LDLR-RAP complex reveals a general mode for ligand recognition by lipoprotein receptors. Mol. Cell 22, 277–283 (2006)
Gordon, M. M., Hu, C., Chokshi, H., Hewitt, J. E. & Alpers, D. H. Glycosylation is not required for ligand or receptor binding by expressed rat intrinsic factor. Am. J. Physiol. 260, G736–G742 (1991)
Gregory, L. A. et al. The X-ray structure of human mannan-binding lectin-associated protein 19 (MAp19) and its interaction site with mannan-binding lectin and L-ficolin. J. Biol. Chem. 279, 29391–29397 (2004)
Appleton, B. A. et al. Structural studies of neuropilin/antibody complexes provide insights into semaphorin and VEGF binding. EMBO J. 26, 4902–4912 (2007)
Bates, P. A., Kelley, L. A., MacCallum, R. M. & Sternberg, M. J. Enhancement of protein modeling by human intervention in applying the automatic programs 3D-JIGSAW and 3D-PSSM. Proteins 45, 39–46 (2001)
Kozyraki, R. et al. The human intrinsic factor-vitamin B12 receptor, cubilin: molecular characterization and chromosomal mapping of the gene to 10p within the autosomal recessive megaloblastic anemia (MGA1) region. Blood 91, 3593–3600 (1998)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Kleywegt, G. J. & Jones, T. A. xdlMAPMAN and xdlDATAMAN – programs for reformatting, analysis and manipulation of biomacromolecular electron-density maps and reflection data sets. Acta Crystallogr. D 52, 826–828 (1996)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Kleywegt, G. J. & Jones, T. A. Halloween... Masks and Bones 〈http://xray.bmc.uu.se/gerard/papers/halloween.html〉.
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007)
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993)
DeLano, W. L. The PyMOL Molecular Graphics System (DeLano Scientific) 〈http://www.pymol.org/〉 (2002)
Bond, C. S. & Schuttelkopf, A. W. ALINE: a WYSIWYG protein-sequence alignment editor for publication-quality alignments. Acta Crystallogr. D 65, 510–512 (2009)
Acknowledgements
We are grateful to G. Biller, G. Ratz and G. Hartvigsen for technical assistance, the staff at MAX-lab and SLS beamlines for help with data collection, and S. Fedosov for advice on expression. The study was essentially supported by a grant to S.K.M. from the Lundbeck foundation. G.R.A. was supported by the Danish Science Research Council and a Hallas-Møller stipend from the Novo-Nordisk foundation. M.M. was supported by the Novo Nordisk Foundation.
Author Contributions C.B.F.A.: cloning, expression, purification, crystallization, data collection, structure determination and analysis, manuscript preparation; M.M.: cloning, expression, purification, size-exclusion chromatographic analyses, manuscript preparation; T.S.: cloning, mutagenesis, expression, size-exclusion chromatographic analyses, cross-linkage experiments; G.R.A. and S.K.M.: study design and manuscript preparation.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1-13 with legends, Supplementary Tables 1-4 and Supplementary References. (PDF 8731 kb)
Rights and permissions
About this article
Cite this article
Andersen, C., Madsen, M., Storm, T. et al. Structural basis for receptor recognition of vitamin-B12–intrinsic factor complexes. Nature 464, 445–448 (2010). https://doi.org/10.1038/nature08874
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08874
This article is cited by
-
CUBN gene mutations may cause focal segmental glomerulosclerosis (FSGS) in children
BMC Nephrology (2022)
-
Deficiency of vitamin B12 and its relation with neurological disorders: a critical review
The Journal of Basic and Applied Zoology (2020)
-
Structural basis for adhesion G protein-coupled receptor Gpr126 function
Nature Communications (2020)
-
Systemically Administered Plant Recombinant Holo-Intrinsic Factor Targets the Liver and is not Affected by Endogenous B12 levels
Scientific Reports (2019)
-
Structural assembly of the megadalton-sized receptor for intestinal vitamin B12 uptake and kidney protein reabsorption
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.