Abstract
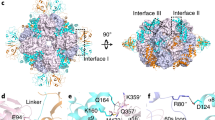
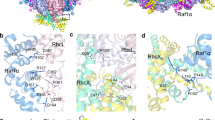
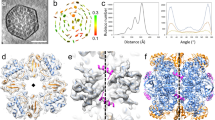
Form I Rubisco (ribulose 1,5-bisphosphate carboxylase/oxygenase), a complex of eight large (RbcL) and eight small (RbcS) subunits, catalyses the fixation of atmospheric CO2 in photosynthesis. The limited catalytic efficiency of Rubisco has sparked extensive efforts to re-engineer the enzyme with the goal of enhancing agricultural productivity. To facilitate such efforts we analysed the formation of cyanobacterial form I Rubisco by in vitro reconstitution and cryo-electron microscopy. We show that RbcL subunit folding by the GroEL/GroES chaperonin is tightly coupled with assembly mediated by the chaperone RbcX2. RbcL monomers remain partially unstable and retain high affinity for GroEL until captured by RbcX2. As revealed by the structure of a RbcL8–(RbcX2)8 assembly intermediate, RbcX2 acts as a molecular staple in stabilizing the RbcL subunits as dimers and facilitates RbcL8 core assembly. Finally, addition of RbcS results in RbcX2 release and holoenzyme formation. Specific assembly chaperones may be required more generally in the formation of complex oligomeric structures when folding is closely coupled to assembly.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Gutteridge, S. & Gatenby, A. A. Rubisco synthesis, assembly, mechanism, and regulation. Plant Cell 7, 809–819 (1995)
Andersson, I. & Taylor, T. C. Structural framework for catalysis and regulation in ribulose-1,5-bisphosphate carboxylase/oxygenase. Arch. Biochem. Biophys. 414, 130–140 (2003)
Tabita, F. R. Microbial ribulose 1,5-bisphosphate carboxylase/oxygenase: a different perspective. Photosynth. Res. 60, 1–28 (1999)
Newman, J., Branden, C. I. & Jones, T. A. Structure determination and refinement of ribulose 1,5-bisphosphate carboxylase/oxygenase from Synechococcus pcc6301. Acta Crystallogr. Sect D: Biol. Crystallogr. 49, 548–560 (1993)
Newman, J. & Gutteridge, S. The X-ray structure of Synechococcus ribulose-bisphosphate carboxylase/oxygenase-activated quaternary complex at 2.2 Å resolution. J. Biol. Chem. 268, 25876–25886 (1993)
Schneider, G., Lindqvist, Y. & Lundqvist, T. Crystallographic refinement and structure of ribulose-1,5-bisphosphate carboxylase from Rhodospirillum rubrum at 1.7 Å resolution. J. Mol. Biol. 211, 989–1008 (1990)
Andrews, T. J. & Lorimer, G. H. in The Biochemistry of plants: a comprehensive treatise (eds Hatch, M D. & Boardman, N K.) 10, 131–218 (1987)
Andersson, I. Catalysis and regulation in Rubisco. J. Exp. Bot. 59, 1555–1568 (2008)
Spreitzer, R. J. & Salvucci, M. E. Rubisco: structure, regulatory interactions, and possibilities for a better enzyme. Annu. Rev. Plant Biol. 53, 449–475 (2002)
Andrews, T. J. & Whitney, S. M. Manipulating ribulose bisphosphate carboxylase/oxygenase in the chloroplasts of higher plants. Arch. Biochem. Biophys. 414, 159–169 (2003)
Spreitzer, R. J., Peddi, S. R. & Satagopan, S. Phylogenetic engineering at an interface between large and small subunits imparts land-plant kinetic properties to algal Rubisco. Proc. Natl Acad. Sci. USA 102, 17225–17230 (2005)
Goloubinoff, P., Christeller, J. T., Gatenby, A. A. & Lorimer, G. H. Reconstitution of active dimeric ribulose bisphosphate carboxylase from an unfolded state depends on two chaperonin proteins and MgATP. Nature 342, 884–889 (1989)
Brinker, A. et al. Dual function of protein confinement in chaperonin-assisted protein folding. Cell 107, 223–233 (2001)
Hartl, F. U. & Hayer-Hartl, M. Converging concepts of protein folding in vitro and in vivo . Nature Struct. Mol. Biol. 16, 574–581 (2009)
Whitney, S. M. & Sharwood, R. E. Construction of a tobacco master line to improve Rubisco engineering in chloroplasts. J. Exp. Bot. 59, 1909–1921 (2008)
Saschenbrecker, S. et al. Structure and function of RbcX, an assembly chaperone for hexadecameric Rubisco. Cell 129, 1189–1200 (2007)
Tanaka, S., Sawaya, M. R., Kerfeld, C. A. & Yeates, T. O. Structure of the RuBisCO chaperone RbcX from Synechocystis sp PCC6803. Acta Crystallogr. Sect D. Biol. Crystallogr. 63, 1109–1112 (2007)
Tarnawski, M., Gubernator, B., Kolesinski, P. & Szczepaniak, A. Heterologous expression and initial characterization of recombinant RbcX protein from Thermosynechococcus elongatus BP-1 and the role of RbcX in RuBisCO assembly. Acta Biochim. Pol. 55, 777–785 (2008)
van der Vies, S. M., Bradley, D. & Gatenby, A. A. Assembly of cyanobacterial and higher plant ribulose bisphosphate carboxylase subunits into functional homologous and heterologous enzyme molecules in Escherichia coli . EMBO J. 5, 2439–2444 (1986)
Andrews, T. J. Catalysis by cyanobacterial ribulose-bisphosphate carboxylase large subunits in the complete absence of small subunits. J. Biol. Chem. 263, 12213–12219 (1988)
Goloubinoff, P., Gatenby, A. A. & Lorimer, G. H. GroE heat-shock proteins promote assembly of foreign prokaryotic ribulose bisphosphate carboxylase oligomers in Escherichia coli . Nature 337, 44–47 (1989)
Li, L.-A. & Tabita, F. R. Maximum activity of recombinant ribulose 1,5-bisphosphate carboxylase/oxygenase of Anabaena sp. strain CA requires the product of the rbcX gene. J. Bacteriol. 179, 3793–3796 (1997)
Onizuka, T. et al. The rbcX gene product promotes the production and assembly of ribulose-1,5-bisphosphate carboxylase/oxygenase of Synechococcus sp. PCC7002 in Escherichia coli . Plant Cell Physiol. 45, 1390–1395 (2004)
Emlyn-Jones, D., Woodger, F. J., Price, G. D. & Whitney, S. M. RbcX can function as a Rubisco chaperonin, but is non-essential in Synechococcus PCC7942. Plant Cell Physiol. 47, 1630–1640 (2006)
van der Vies, S. et al. Conformational states of ribulose bisphosphate carboxylase and their interaction with chaperonin 60. Biochem. 31, 3635–3644 (1992)
Weissman, J. S. et al. Characterization of the active intermediate of a GroEL-GroES-mediated protein folding reaction. Cell 84, 481–490 (1996)
Langer, T. et al. Chaperonin-mediated protein folding: GroES binds to one end of the GroEL cylinder, which accommodates the protein substrate within its central cavity. EMBO J. 11, 4757–4765 (1992)
Martin, J., Mayhew, M., Langer, T. & Hartl, F. U. The reaction cycle of GroEL and GroES in chaperonin-assisted protein folding. Nature 366, 228–233 (1993)
Martin, J. et al. Chaperonin-mediated protein folding at the surface of GroEL through a ‘molten globule’-like intermediate. Nature 352, 36–42 (1991)
Thow, G., Zhu, G. H. & Spreitzer, R. J. Complementing substitutions within loop regions 2 and 3 of the α/β-barrel active site influence the CO2/O2 specificity of chloroplast ribulose-1,5-bisphosphate carboxylase/oxygenase. Biochem. 33, 5109–5114 (1994)
Tabita, F. R. et al. Function, structure, and evolution of the RubisCO-like proteins and their RubisCO homologs. Microbiol. Mol. Biol. Rev. 71, 576–599 (2007)
Barraclough, R. & Ellis, R. J. Protein synthesis in chloroplasts. IX. Assembly of newly-synthesized large subunits into ribulose bisphosphate carboxylase in isolated intact pea chloroplasts. Biochim. Biophys. Acta 608, 19–31 (1980)
Hayer-Hartl, M. K., Weber, F. & Hartl, F. U. Mechanism of chaperonin action: GroES binding and release can drive GroEL-mediated protein folding in the absence of ATP hydrolysis. EMBO J. 15, 6111–6121 (1996)
Roessner, D. & Kulicke, W. M. On-line coupling of flow field-flow fractionation and multi-angle laser light scattering. J. Chromatogr. A 687, 249–258 (1994)
Tang, Y. C. et al. Structural features of the GroEL-GroES nano-cage required for rapid folding of encapsulated protein. Cell 125, 903–914 (2006)
Chin, J. W. & Schultz, P. G. In vivo photocrosslinking with unnatural amino acid mutagenesis. ChemBioChem 3, 1135–1137 (2002)
Lakshmipathy, S. K. et al. Identification of nascent chain interaction sites on trigger factor. J. Biol. Chem. 282, 12186–12193 (2007)
Weiner, M. P. et al. Site-directed mutagenesis of double-stranded DNA by the polymerase chain reaction. Gene 151, 119–123 (1994)
Wiseman, T., Williston, S., Brandts, J. F. & Lin, L. N. Rapid measurement of binding constants and heats of binding using a new titration calorimeter. Anal. Biochem. 179, 131–137 (1989)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: Semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
Pettersen, E. F. et al. UCSF chimera - A visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Chin, J. W., Martin, A. B., King, D. S., Wang, L. & Schultz, P. G. Addition of a photocrosslinking amino acid to the genetic code of Escherichia coli . Proc. Natl Acad. Sci. USA 99, 11020–11024 (2002)
Ryu, Y. & Schultz, P. G. Efficient incorporation of unnatural amino acids into proteins in Escherichia coli . Nature Methods 3, 263–265 (2006)
Leslie, A. G. W. Recent changes to the MOSFLM package for processing film and image plate data. Joint CCP4 and ESF-EACBM Newsletter on Protein Crystallography 26, (1992)
Evans, P. R. Scala. Joint CCP4 and ESF-EACBM Newsletter on Protein Crystallography 33, 22–24 (1997)
Vagin, A. A. & Isupov, M. N. Spherically averaged phased translation function and its application to the search for molecules and fragments in electron-density maps. Acta Crystallogr. Sect D: Biol. Crystallogr. 57, 1451–1456 (2001)
Perrakis, A., Morris, R. & Lamzin, V. S. Automated protein model building combined with iterative structure refinement. Nature Struct. Biol. 6, 458–463 (1999)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. Sect D: Biol. Crystallogr. 60, 2126–2132 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. Sect D: Biol. Crystallogr. 53, 240–255 (1997)
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Cryst. 26, 283–291 (1993)
Acknowledgements
Assistance by K. Valer and J. Basquin in the MPIB crystallization facility, by C. Ungewickell in the Gene Center cryo-EM facility as well as by the staff at SLS-beamline X10SA-PX-II is acknowledged. We thank the Deutsche Forschungsgemeinschaft (SFB 594), the Ernst-Jung Foundation, the Körber Foundation, the European Union and the Center for Integrated Protein Science Munich (CIPSM) for financial support.
Author Contributions C.L. designed and executed most of the Rubisco reconstitution experiments and purified the Syn6301-RbcL8 core complexes with contributions from S.S., B.V.R. and K.V.R. The cryo-EM and three-dimensional image analysis were carried out by A.L.Y., as well as the interpretation and fitting of available crystal structures with contributions by O.B. and T.M. to data collection. A.S.-W. performed the crosslinking experiments, peptide binding assays and mutational analysis. M.H.-H. performed the FFF measurements. A.B. purified the Syn6301-RbcL8–AnaCA-(RbcX2)8 complexes and determined the crystal structure of AnaCA-RbcX2. R.B. supervised the design and interpretation of the cryo-EM analysis. M.H.-H. and F.U.H. supervised the design and interpretation of the biochemical experiments and wrote the manuscript with contributions from A.L.Y. and R.B.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Table 1, Supplementary Figures 1-8 with Legends and Supplementary References. (PDF 2532 kb)
Rights and permissions
About this article
Cite this article
Liu, C., Young, A., Starling-Windhof, A. et al. Coupled chaperone action in folding and assembly of hexadecameric Rubisco. Nature 463, 197–202 (2010). https://doi.org/10.1038/nature08651
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08651
This article is cited by
-
Scaffolding protein CcmM directs multiprotein phase separation in β-carboxysome biogenesis
Nature Structural & Molecular Biology (2021)
-
Engineering the Calvin–Benson–Bassham cycle and hydrogen utilization pathway of Ralstonia eutropha for improved autotrophic growth and polyhydroxybutyrate production
Microbial Cell Factories (2020)
-
Novel bacterial clade reveals origin of form I Rubisco
Nature Plants (2020)
-
Small subunits can determine enzyme kinetics of tobacco Rubisco expressed in Escherichia coli
Nature Plants (2020)
-
Possible solutions to several enigmas of Cretaceous climate
International Journal of Earth Sciences (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.