Abstract
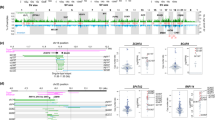
Multiple somatic rearrangements are often found in cancer genomes; however, the underlying processes of rearrangement and their contribution to cancer development are poorly characterized. Here we use a paired-end sequencing strategy to identify somatic rearrangements in breast cancer genomes. There are more rearrangements in some breast cancers than previously appreciated. Rearrangements are more frequent over gene footprints and most are intrachromosomal. Multiple rearrangement architectures are present, but tandem duplications are particularly common in some cancers, perhaps reflecting a specific defect in DNA maintenance. Short overlapping sequences at most rearrangement junctions indicate that these have been mediated by non-homologous end-joining DNA repair, although varying sequence patterns indicate that multiple processes of this type are operative. Several expressed in-frame fusion genes were identified but none was recurrent. The study provides a new perspective on cancer genomes, highlighting the diversity of somatic rearrangements and their potential contribution to cancer development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Kaye, F. J. Mutation-associated fusion cancer genes in solid tumors. Mol. Cancer Ther. 8, 1399–1408 (2009)
Mitelman, F., Johansson, B. & Mertens, F. Fusion genes and rearranged genes as a linear function of chromosome aberrations in cancer. Nature Genet. 36, 331–334 (2004)
Mitelman, F., Johansson, B. & Mertens, F. The impact of translocations and gene fusions on cancer causation. Nature Rev. Cancer 7, 233–245 (2007)
Futreal, P. A. et al. A census of human cancer genes. Nature Rev. Cancer 4, 177–183 (2004)
Stratton, M. R., Campbell, P. J. & Futreal, P. A. The cancer genome. Nature 458, 719–724 (2009)
Sawyers, C. L. Chronic myeloid leukemia. N. Engl. J. Med. 340, 1330–1340 (1999)
Tomlins, S. A. et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310, 644–648 (2005)
Soda, M. et al. Identification of the transforming EML4–ALK fusion gene in non-small-cell lung cancer. Nature 448, 561–566 (2007)
Höglund, M., Gisselsson, D., Sall, T. & Mitelman, F. Coping with complexity. multivariate analysis of tumor karyotypes. Cancer Genet. Cytogenet. 135, 103–109 (2002)
Bignell, G. R. et al. Architectures of somatic genomic rearrangement in human cancer amplicons at sequence-level resolution. Genome Res. 17, 1296–1303 (2007)
Volik, S. et al. End-sequence profiling: sequence-based analysis of aberrant genomes. Proc. Natl Acad. Sci. USA 100, 7696–7701 (2003)
Ruan, Y. et al. Fusion transcripts and transcribed retrotransposed loci discovered through comprehensive transcriptome analysis using Paired-End diTags (PETs). Genome Res. 17, 828–838 (2007)
Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007)
Wood, L. D. et al. The genomic landscapes of human breast and colorectal cancers. Science 318, 1108–1113 (2007)
The Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008)
Weir, B. A. et al. Characterizing the cancer genome in lung adenocarcinoma. Nature 450, 893–898 (2007)
Campbell, P. J. et al. Identification of somatically acquired rearrangements in cancer using genome-wide massively parallel paired-end sequencing. Nature Genet. 40, 722–729 (2008)
Weterings, E. & Chen, D. J. The endless tale of non-homologous end-joining. Cell Res. 18, 114–124 (2008)
van Gent, D. C. & van der Burg, M. Non-homologous end-joining, a sticky affair. Oncogene 26, 7731–7740 (2007)
Hefferin, M. L. & Tomkinson, A. E. Mechanism of DNA double-strand break repair by non-homologous end joining. DNA Repair 4, 639–648 (2005)
Hastings, P. J., Lupski, J. R., Rosenberg, S. M. & Ira, G. Mechanisms of change in gene copy number. Nature Rev. Genet. 10, 551–564 (2009)
Yan, C. T. et al. IgH class switching and translocations use a robust non-classical end-joining pathway. Nature 449, 478–482 (2007)
Bohlander, S. K. ETV6: a versatile player in leukemogenesis. Semin. Cancer Biol. 15, 162–174 (2005)
Knezevich, S. R., McFadden, D. E., Tao, W., Lim, J. F. & Sorensen, P. H. A novel ETV6–NTRK3 gene fusion in congenital fibrosarcoma. Nature Genet. 18, 184–187 (1998)
Lannon, C. L. & Sorensen, P. H. ETV6–NTRK3: a chimeric protein tyrosine kinase with transformation activity in multiple cell lineages. Semin. Cancer Biol. 15, 215–223 (2005)
Jones, D. T. et al. Tandem duplication producing a novel oncogenic BRAF fusion gene defines the majority of pilocytic astrocytomas. Cancer Res. 68, 8673–8677 (2008)
Basecke, J., Whelan, J. T., Griesinger, F. & Bertrand, F. E. The MLL partial tandem duplication in acute myeloid leukaemia. Br. J. Haematol. 135, 438–449 (2006)
Blow, J. J. & Gillespie, P. J. Replication licensing and cancer—a fatal entanglement? Nature Rev. Cancer 8, 799–806 (2008)
Perou, C. M. et al. Molecular portraits of human breast tumours. Nature 406, 747–752 (2000)
Sorlie, T. et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl Acad. Sci. USA 98, 10869–10874 (2001)
Chin, K. et al. Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell 10, 529–541 (2006)
Bergamaschi, A. et al. Distinct patterns of DNA copy number alteration are associated with different clinicopathological features and gene-expression subtypes of breast cancer. Genes Chromosom. Cancer 45, 1033–1040 (2006)
Li, H., Ruan, J. & Durbin, R. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 18, 1851–1858 (2008)
Ning, Z., Cox, A. J. & Mullikin, J. C. SSAHA: a fast search method for large DNA databases. Genome Res. 11, 1725–1729 (2001)
Venkatraman, E. & Olshen, A. A faster circular binary segmentation algorithm for the analysis of array CGH data. Bioinformatics 23, 657–663 (2007)
Lambros, M. et al. Unlocking pathology archives for molecular genetic studies: a reliable method to generate probes for chromogenic and fluorescent in situ hybridization. Lab. Invest. 86, 398–408 (2006)
Abeysinghe, S. S. et al. Translocation and gross deletion breakpoints in human inherited disease and cancer I: Nucleotide composition and recombination-associated motifs. Hum. Mutat. 22, 229–244 (2003)
Scholz, F. W. & Stephens, M. A. K-sample Anderson-Darling tests. Am. J. Stat. Assoc. 82, 918–924 (1987)
Acknowledgements
We are grateful to M. Lambros, F. Geyer and R. Vatcheva for their assistance in the FISH experiments. We would like to acknowledge the support of the Kay Kendall Leukaemia Fund under Grant KKL282, Human Frontiers Award reference LT000561/2009-L, the Dana-Farber/Harvard SPORE in breast cancer under NCI grant reference CA089393, Breakthrough Breast Cancer, the Research Council of Norway Grants no. 155218 and 175240, and the Wellcome Trust under grant reference 077012/Z/05/Z.
Author Contributions M.R.S., P.A.F., P.J.C. and P.J.S. designed the experiment. S.E., D.J.M., P.J.S., M-L.L., I.V., L.J.M., J.B., M.A.Q., H.S., C.C., R.N., A.M.S., A.L., J.W.M.M. and C.Latimer carried out laboratory analyses. J.A.F., J.S.R.-F., L.v.V., A.L.R. D.P.S. and A.-L.B.-D. provided clinical samples. P.J.S., D.J.M., I.V., M.-L.L., E.D.P., J.T.S., L.A.S., C.Leroy, C.D.G., M.J., J.W.T., K.W.L., P.J.C., P.A.F., J.S.R.-F., J.W.M.M., A.M.S., J.A.F., M.R.S., H.E.G.R., A.L.R., A.-L.B.-D., L.v.V., A.L., P.J.C. and P.A.F. performed data analysis, informatics and statistics. M.R.S. wrote the manuscript with comments from P.J.S., P.A.F., P.J.C., A.-L.B.-D., J.S.R.-F., J.A.F., A.L.R., D.P.S. and L.v.V.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-7 with Legends. (PDF 5501 kb)
Supplementary Table 1
Summary of somatic rearrangements found in 24 breast cancers (XLS 845 kb)
Supplementary Table 2
Variation in the prevalence of rearrangement architectures in individual breast cancer genomes. (XLS 28 kb)
Supplementary Table 3
Base pair resolution of rearrangement breakpoints. (XLS 350 kb)
Supplementary Table 4
GC content analysis at rearrangement breakpoints. (XLS 22 kb)
Supplementary Table 5
Evaluation of motif enrichment at rearrangement breakpoints. (XLS 24 kb)
Supplementary Table 6
Gene Fusions. (PDF 135 kb)
Supplementary Table 7
Internally rearranged genes. (XLS 69 kb)
Supplementary Table 8
Known cancer genes that are rearranged. (XLS 36 kb)
Supplementary Table 9
Rearranged genes. (XLS 452 kb)
Rights and permissions
About this article
Cite this article
Stephens, P., McBride, D., Lin, ML. et al. Complex landscapes of somatic rearrangement in human breast cancer genomes. Nature 462, 1005–1010 (2009). https://doi.org/10.1038/nature08645
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08645
This article is cited by
-
Reference Materials for Improving Reliability of Multiomics Profiling
Phenomics (2024)
-
Triple-negatives Mammakarzinom
Die Pathologie (2023)
-
Comparative characterization of 3D chromatin organization in triple-negative breast cancers
Experimental & Molecular Medicine (2022)
-
Role of non-coding RNA in immune microenvironment and anticancer therapy of gastric cancer
Journal of Molecular Medicine (2022)
-
Three-step one-way model in terahertz biomedical detection
PhotoniX (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.