Abstract
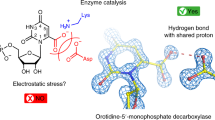
Acetoacetate decarboxylase (AADase) has long been cited as the prototypical example of the marked shifts in the pKa values of ionizable groups that can occur in an enzyme active site. In 1966, it was hypothesized that in AADase the origin of the large pKa perturbation (-4.5 log units) observed in the nucleophilic Lys 115 results from the proximity of Lys 116, marking the first proposal of microenvironment effects in enzymology. The electrostatic perturbation hypothesis has been demonstrated in a number of enzymes, but never for the enzyme that inspired its conception, owing to the lack of a three-dimensional structure. Here we present the X-ray crystal structures of AADase and of the enamine adduct with the substrate analogue 2,4-pentanedione. Surprisingly, the shift of the pKa of Lys 115 is not due to the proximity of Lys 116, the side chain of which is oriented away from the active site. Instead, Lys 116 participates in the structural anchoring of Lys 115 in a long, hydrophobic funnel provided by the novel fold of the enzyme. Thus, AADase perturbs the pKa of the nucleophile by means of a desolvation effect by placement of the side chain into the protein core while enforcing the proximity of polar residues, which facilitate decarboxylation through electrostatic and steric effects.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
The coordinates and structure factors have been deposited in the Protein Data Bank with accession codes 3BH2, 3BGT and 3BH3 corresponding, respectively, to C. acetobutylicum acetoacetate decarboxylase and C. violaceum acetoacetate decarboxylase in the unliganded form and complexed with 2,4-pentanedione.
References
Westheimer, F. H. Coincidences, decarboxylation and electrostatic effects. Tetrahedron 51, 3–20 (1995)
Hamilton, G. A. & Westheimer, F. H. On the mechanism of the enzymatic decarboxylation of acetoacetate. J. Am. Chem. Soc. 81, 6332–6333 (1959)
Fridovich, I. & Westheimer, F. H. On the mechanism of the enzymatic decarboxylation of acetoacetate. II. J. Am. Chem. Soc. 84, 3208–3209 (1962)
Laursen, R. A. & Westheimer, F. H. The active site of acetoacetate decarboxylase. J. Am. Chem. Soc. 88, 3426–3430 (1966)
Warren, S., Zerner, B. & Westheimer, F. H. Acetoacetate decarboxylase. Identification of lysine at the active site. Biochemistry 5, 817–823 (1966)
Kokesh, F. C. & Westheimer, F. H. A reporter group at the active site of acetoacetate decarboxylase. II. Ionization constant of the amino group. J. Am. Chem. Soc. 93, 7270–7274 (1971)
Harris, T. K. & Turner, G. J. Structural basis of perturbed pKa values of catalytic groups in enzyme active sites. IUBMB Life 53, 85–98 (2002)
Gerlt, J. A. Obituary, Frank H. Westheimer. Nature 447, 543 (2007)
Gallivan, J. P. & Dougherty, D. A. Cation–π interactions in structural biology. Proc. Natl Acad. Sci. USA 96, 9459–9464 (1999)
Highbarger, L. A., Gerlt, J. A. & Kenyon, G. L. Mechanism of the reaction catalyzed by acetoacetate decarboxylase. Importance of lysine 116 in determining the pKa of active-site lysine 115. Biochemistry 35, 41–46 (1996)
García-Moreno, E. B. et al. Experimental measurement of the effective dielectric in the hydrophobic core of a protein. Biophys. Chem. 64, 211–224 (1997)
Lee, J. K. & Houk, K. N. A proficient enzyme revisited: the predicted decarboxylase mechanism for orotidine monophosphate. Science 276, 942–945 (1997)
Rashin, V. & Honig, B. Reevaluation of the Born model of ion hydration. J. Phys. Chem. 89, 5588–5593 (1985)
Stites, W. E., Gittis, A. G., Lattman, E. E. & Shortle, D. In a staphylococcal nuclease mutant the side-chain of a lysine replacing valine66 is fully buried in the hydrophobic core. J. Mol. Biol. 221, 7–14 (1991)
Barbas, C. F. et al. Immune versus natural selection: antibody aldolases with enzymic rates but broader scope. Science 278, 2085–2092 (1997)
Crosby, J., Stone, R. & Liehard, G. E. Mechanisms of thiamine-catalyzed reactions. Decarboxylation of 2-(1-carboxy-1-hydroxyethyl)-3,4-dimethylthiazolium chloride. J. Am. Chem. Soc. 92, 2891–2900 (1970)
Fridovich, I. A study of the interaction of acetoacetic decarboxylase with several inhibitors. J. Biol. Chem. 243, 1043–1051 (1968)
O’Leary, M. H. in Mechanisms of Catalysis (ed. Sigman, D. S.) 239 (Academic, 1992)
Tagaki, W. & Westheimer, F. H. Acetoacetate decarboxylase. Reassociation of subunits. Biochemistry 7, 891–894 (1968)
Wiesenborn, D. P., Rudolph, F. B. & Papoutsakis, E. T. Thiolase from Clostridium acetobutylicum ATCC 824 and its role in the synthesis of acids and solvents. Appl. Environ. Microbiol. 54, 2717–2722 (1988)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Pape, T. & Schneider, T. R. HKL2MAP: a graphical user interface for phasing with SHELX programs. J. Appl. Crystallogr. 37, 843–844 (2004)
Usón, I. & Sheldrick, G. M. Advances in direct methods for protein crystallography. Curr. Opin. Struct. Biol. 9, 643–648 (1999)
Terwilliger, T. C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999)
Terwilliger, T. C. Maximum likelihood density modification. Acta Crystallogr. D 56, 965–972 (2000)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphic. Acta Crystallogr. D 60, 2126–2132 (2004)
Collaborative computation project number 4 The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Brünger, A. T. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Storoni, L. C., McCoy, A. J. & Read, R. J. Likelihood-enhanced fast rotation function. Acta Crystallogr. D 60, 432–438 (2004)
Potterton, E., Briggs, P., Turkenburg, M. & Dodson, E. A graphical user interface to the CCP4 program suite. Acta Crystallogr. D 59, 1131–1137 (2003)
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. Procheck: a program to check the sterochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993)
Smolsky, I. L. et al. Biological small-angle x-ray scattering facility at the Stanford Synchrotron Radiation Laboratory. J. Appl. Crystallogr. 40, S453 (2007)
Konarev, P. V. et al. PRIMUS: a Windows PC-based system for small-angle scattering data analysis. J. Appl. Crystallogr. 36, 1277–1282 (2003)
Tjioe, E. & Heller, W. T. ORNL_SAS: software for calculation of small-angle scattering intensities of proteins and protein complexes. J. Appl. Crystallogr. 40, 782–785 (2007)
Svergun, D. I. et al. Large differences are observed between the crystal and solution quaternary structures of allosteric aspartate transcarbamylase in the R state. Proteins 27, 110–117 (1997)
Svergun, D. I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 25, 495–503 (1992)
Ménétret, J. F. et al. The structure of ribosome–channel complexes engaged in protein translocation. Mol. Cell 6, 1219–1232 (2000)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
Ludtke, S. J., Chen, D. H., Song, J. L., Chuang, D. T. & Chiu, W. Seeing GroEL at 6 Å resolution by single particle electron cryomicroscopy. Structure 12, 1129–1136 (2004)
Acknowledgements
We thank A. Murzin for provisional classification of the AADase fold and C. Akey for valuable advice on the execution and interpretation of the electron microscopy work. We also thank H. Robinson and A. Soares for help with data collection. This work was supported by a grant to K.N.A. from the National Science Foundation. Data for this study were measured at Beamlines X12B, X25 and X29A of the National Synchrotron Light Source. Financial support comes principally from the Offices of Biological and Environmental Research (BER) and of Basic Energy Sciences (BES) of the US Department of Energy (DOE), and from the National Center for Research Resources (NCRR) of the National Institutes of Health (NIH). Small-angle X-ray scattering analyses were carried out at the Stanford Synchrotron Radiation Lightsource, funded by DOE, BES. The SSRL Structural Molecular Biology Program is supported by DOE, BER, and by NIH, NCRR. The contents of this work are solely the responsibility of the authors and do not necessarily represent the official view of NCRR or NIH.
Author Contributions M.-C.H. cloned, expressed, purified, crystallized, collected data and performed crystal structure determination, refinement and model analysis. H.T. designed, executed, analysed and wrote the description of the small-angle X-ray scattering analysis. J.F.M. designed, executed, analysed and wrote the description of the electron microscopy experiments. K.N.A. conceived of and designed the project. M.-C.H. and K.N.A. wrote the manuscript; all authors discussed the results and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1-S8 with Legends, Supplementary Tables S1-S4 and a Supplementary Movie Legend. (PDF 1049 kb)
Supplementary Movie
This movie depicts the X-ray crystallographic structure of acetoacetate decarboxylase in the EM density (see file s1 for full Legend). (MP4 3979 kb)
Rights and permissions
About this article
Cite this article
Ho, MC., Ménétret, JF., Tsuruta, H. et al. The origin of the electrostatic perturbation in acetoacetate decarboxylase. Nature 459, 393–397 (2009). https://doi.org/10.1038/nature07938
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature07938
This article is cited by
-
Ground-state destabilization by electrostatic repulsion is not a driving force in orotidine-5′-monophosphate decarboxylase catalysis
Nature Catalysis (2022)
-
Production of the biocommodities butanol and acetone from methanol with fluorescent FAST-tagged proteins using metabolically engineered strains of Eubacterium limosum
Biotechnology for Biofuels (2021)
-
Microbial and enzymatic conversion of levulinic acid, an alternative building block to fermentable sugars from cellulosic biomass
Applied Microbiology and Biotechnology (2020)
-
Lysine relay mechanism coordinates intermediate transfer in vitamin B6 biosynthesis
Nature Chemical Biology (2017)
-
Benchmarking pKa prediction methods for Lys115 in acetoacetate decarboxylase
Journal of Molecular Modeling (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.