Abstract
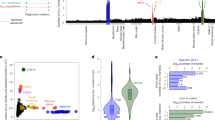
Comprehensive identification of polymorphisms among individuals within a species is essential both for studying the genetic basis of phenotypic differences and for elucidating the evolutionary history of the species. Large-scale polymorphism surveys have recently been reported for human1, mouse2 and Arabidopsis thaliana3. Here we report a nucleotide-level survey of genomic variation in a diverse collection of 63 Saccharomyces cerevisiae strains sampled from different ecological niches (beer, bread, vineyards, immunocompromised individuals, various fermentations and nature) and from locations on different continents. We hybridized genomic DNA from each strain to whole-genome tiling microarrays and detected 1.89 million single nucleotide polymorphisms, which were grouped into 101,343 distinct segregating sites. We also identified 3,985 deletion events of length >200 base pairs among the surveyed strains. We analysed the genome-wide patterns of nucleotide polymorphism and deletion variants, and measured the extent of linkage disequilibrium in S. cerevisiae. These results and the polymorphism resource we have generated lay the foundation for genome-wide association studies in yeast. We also examined the population structure of S. cerevisiae, providing support for multiple domestication events as well as insight into the origins of pathogenic strains.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005)
Frazer, K. A. et al. A sequence-based variation map of 8.27 million SNPs in inbred mouse strains. Nature 448, 1050–1053 (2007)
Clark, R. M. et al. Common sequence polymorphisms shaping genetic diversity in Arabidopsis thaliana. Science 317, 338–342 (2007)
Kellis, M., Patterson, N., Endrizzi, M., Birren, B. & Lander, E. S. Sequencing and comparison of yeast species to identify genes and regulatory elements. Nature 423, 241–254 (2003)
Cliften, P. et al. Finding functional features in Saccharomyces genomes by phylogenetic footprinting. Science 301, 71–76 (2003)
Dujon, B. et al. Genome evolution in yeasts. Nature 430, 35–44 (2004)
Dujon, B. Yeasts illustrate the molecular mechanisms of eukaryotic genome evolution. Trends Genet. 22, 375–387 (2006)
Gresham, D. et al. Genome-wide detection of polymorphisms at nucleotide resolution with a single DNA microarray. Science 311, 1932–1936 (2006)
Carter, D. M. et al. Population genomics of domestic and wild yeasts. Nature (submitted)
Gerton, J. L. et al. Inaugural article: global mapping of meiotic recombination hotspots and coldspots in the yeast Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 97, 11383–11390 (2000)
Pryde, F. E., Gorham, H. C. & Louis, E. J. Chromosome ends: all the same under their caps. Curr. Opin. Genet. Dev. 7, 822–828 (1997)
Schacherer, J. et al. Genome-wide analysis of nucleotide-level variation in commonly used Saccharomyces cerevisiae strains. PLoS ONE 2, e322 (2007)
Winzeler, E. A. et al. Genetic diversity in yeast assessed with whole-genome oligonucleotide arrays. Genetics 163, 79–89 (2003)
Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999)
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000)
Fay, J. C. & Benavides, J. A. Evidence for domesticated and wild populations of Saccharomyces cerevisiae. PLoS Genet. 1, 66–71 (2005)
Mortimer, R. K. & Johnston, J. R. Genealogy of principal strains of the yeast genetic stock center. Genetics 113, 35–43 (1986)
Enache-Angoulvant, A. & Hennequin, C. Invasive Saccharomyces infection: a comprehensive review. Clin. Infect. Dis. 41, 1559–1568 (2005)
de Llanos, R., Querol, A., Peman, J., Gobernado, M. & Fernandez-Espinar, M. T. Food and probiotic strains from the Saccharomyces cerevisiae species as a possible origin of human systemic infections. Int. J. Food Microbiol. 110, 286–290 (2006)
Steinmetz, L. M. et al. Dissecting the architecture of a quantitative trait locus in yeast. Nature 416, 326–330 (2002)
Deutschbauer, A. M. & Davis, R. W. Quantitative trait loci mapped to single-nucleotide resolution in yeast. Nature Genet. 37, 1333–1340 (2005)
Gatbonton, T. et al. Telomere length as a quantitative trait: genome-wide survey and genetic mapping of telomere length-control genes in yeast. PLoS Genet. 2, e35 (2006)
Brem, R. B., Yvert, G., Clinton, R. & Kruglyak, L. Genetic dissection of transcriptional regulation in budding yeast. Science 296, 752–755 (2002)
Perlstein, E. O., Ruderfer, D. M., Roberts, D. C., Schreiber, S. L. & Kruglyak, L. Genetic basis of individual differences in the response to small-molecule drugs in yeast. Nature Genet. 39, 496–502 (2007)
Huson, D. H. & Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 23, 254–267 (2006)
Thornton, K. Libsequence: a C++ class library for evolutionary genetic analysis. Bioinformatics 19, 2325–2327 (2003)
Marjoram, P. & Wall, J. D. Fast “coalescent” simulation. BMC Genet. 7, 16 (2006)
Fu, Y. X. Estimating effective population size or mutation rate using the frequencies of mutations of various classes in a sample of DNA sequences. Genetics 138, 1375–1386 (1994)
Tajima, F. Evolutionary relationship of DNA sequences in finite populations. Genetics 105, 437–460 (1983)
Yang, Z. PAML: a program package for phylogenetic analysis by maximum likelihood. Comput. Appl. Biosci. 13, 555–556 (1997)
Acknowledgements
We are grateful to all the researchers and institutions, and especially J. Fay, for sharing yeast strains. We thank K. Dolinski and J. Matese for technical support and D. Gresham for comments on the manuscript. The authors acknowledge discussions with members of the Kruglyak and Botstein laboratories. This work was supported by NIH grant R37 MH059520 and a James S. McDonnell Foundation Centennial Fellowship to L.K., and NIH grant GM071508 to the Lewis-Sigler Institute.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1- S9 and Supplementary Tables 1 and 4 (PDF 2229 kb)
Table 2
This file contains Supplementary Table 2 (XLS 376 kb)
Table 3
This file contains Supplementary Table 3 (XLS 70 kb)
Rights and permissions
About this article
Cite this article
Schacherer, J., Shapiro, J., Ruderfer, D. et al. Comprehensive polymorphism survey elucidates population structure of Saccharomyces cerevisiae. Nature 458, 342–345 (2009). https://doi.org/10.1038/nature07670
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07670
This article is cited by
-
From beer to breadboards: yeast as a force for biological innovation
Genome Biology (2024)
-
Ancient and recent origins of shared polymorphisms in yeast
Nature Ecology & Evolution (2024)
-
Single nucleotide polymorphisms associated with wine fermentation and adaptation to nitrogen limitation in wild and domesticated yeast strains
Biological Research (2023)
-
Comparative metabolomic and transcriptomic analysis of Saccharomyces cerevisiae W303a and CEN.PK2-1C
World Journal of Microbiology and Biotechnology (2023)
-
Genetic diversity and population structure of Saccharomyces cerevisiae isolated from Turkish sourdough by iPBS-retrotransposons markers
Archives of Microbiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.