Abstract
Arising from: C. E. Crespo-Hernández, B. Cohen & B. Kohler Nature 436, 1141–1144 (2005); Crespo-Hernández et al. reply
Absorption of ultraviolet light by DNA is known to lead to carcinogenic mutations, but the processes between photon absorption and the photochemical reactions are poorly understood. In their study of the excited-stated dynamics of model DNA helices using femtosecond transient absorption spectroscopy1, Crespo-Hernández et al. observe that the picosecond component of the transient signals recorded for the adenine–thymine oligonucleotide (dA)18·(dT)18 is close to that for (dA)18, but quite different from that for (dAdT)9·(dAdT)9; from this observation, they conclude that excimer formation limits excitation energy to one strand at a time. Here we use time-resolved fluorescence spectroscopy to probe the excited-state dynamics, which reveals the complexity of these systems and indicates that the interpretation of Crespo-Hernández et al. is an oversimplification. We also comment on the pertinence of separating base stacking and base pairing in excited-state dynamics of double helices and question the authors' assignment of the long-lived signal component found for (dA)18·(dT)18 to adenine excimers.
Similar content being viewed by others
Main
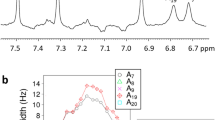
Figure 1a shows the fluorescence decays of (dA)20 at three different wavelengths. Combining these results with our previous measurements obtained for (dA)20 by femtosecond fluorescence upconversion2, at least five exponentials are needed to fit the decays over the 100 femtoseconds to 20 nanoseconds time range. A crucial point is that all time constants vary strongly with the emission wavelength. The same effect is encountered for (dAdT)10·(dAdT)10 and has been reported previously3 for poly(dA)·poly(dT).
a, (dA)20 at 330 nm (magenta), 360 nm (red) and 420 nm (yellow). Black lines correspond to fits with multi-exponential functions, yielding the following sets of time constants (3.2, 37.5, 186 and 748 ps), (11.6, 101, 253 and 1,830 ps) and (39, 198, 551 and 5,050 ps), respectively. The instrumental response function (grey) is represented by the sub-picosecond fluorescence decay at 330 nm of thymidine-5′-monophosphate. b, (dA)20 at 360 nm (red), (dA)20·(dT)20 at 330 nm (blue), (dAdT)10·(dAdT)10 at 420 nm (green). Methods. DNA oligomers (from Eurogentec) dissolved in phosphate buffer (pH 6.8) were excited by femtosecond pulses (100 fs, 267 nm). All decay curves correspond to the total fluorescence F(t) constructed from the parallel (Ipar) and perpendicular (Iperp) components according to: F(t)=Ipar(t)+2GIperp(t), where G accounts for the polarization-dependent sensitivity of the detection system.For further experimental details, see ref. 3.
We interpret this complex behaviour in terms of a model in which a large number of excited states are formed that are delocalized over several bases, which may be located both on the same strand and on opposite strands, with subsequent energy transfer3. This model is based on calculations made in the frame of exciton theory and combines quantum chemistry data and molecular dynamics simulations4,5,6,7, and accounts not only for the fluorescence decays but also the steady-state absorption and fluorescence spectra. Delocalization of the excitation energy is governed by the electronic coupling, which depends on the oligomer's conformation. Conformational changes occurring on pico- and nanosecond timescales are controlled by an ensemble of interactions involving not only the bases but also the backbone, counterions and water molecules5,8. In this sense, both base stacking and base pairing determine excited-state dynamics.
For these reasons, the time constants provide a phenomenological description of the decays and do not correspond to specific excited states. However, it is possible to make a rough comparison of the overall excited-state dynamics by considering the decays recorded at the maxima of the fluorescence spectra of the three oligomers (Fig. 1b). In the case of (dAdT)10·(dAdT)10 and (dA)20, the fluorescence maxima have been assigned to excimer emission9,10. We observe that, in contrast to the transient absorption signals, the fluorescence decay obtained for (dA)20·(dT)20 on the sub-nanosecond timescale is much shorter than that for (dA)20. The same observation is valid when comparing the decays of these single and double strands at identical wavelengths.
Neither in fluorescence nor in transient absorption experiments is the amplitude of the detected signals proportional to the excited-state population. The transient absorption signals depend on the difference between the molar extinction coefficients of the S0 → S1 and S1 → Sn transitions (where S0 and S1 denote the ground and first excited electronic states and Sn denotes different, higher excited electronic states) at the probed wavelengths. As the steady-state absorption spectra of these oligomers correspond to a large number of transitions6 and nothing is known about the S1 → Sn spectra of their various excited states, the percentage of the ‘excimer’ population reported by Crespo-Hernández et al. is not necessarily correct.
The important difference between the transient absorption and fluorescence decays of (dA)n·(dT)n indicates the formation of dark transient species, as also discussed by Crespo-Hernández and colleagues1,10. If these dark species are adenine ‘excimers’, they must have a different electronic structure from the fluorescent ‘excimers’ of (dA)n and, therefore, different lifetimes. Consequently, the similar time constants observed by transient absorption for (dA)18·(dT)18 and (dA)18 may be fortuitous. The species observed by transient absorption in (dA)18·(dT)18 could as well be interstrand A–T charge-transfer states, as suggested by theoretical calculations11. The behaviour of such states in deuterated water, as examined by Crespo-Hernández et al., is not readily predictable because water molecules form a variety of interstrand and intrastrand bridges between bases12.
References
Crespo-Hernández, C. E., Cohen, B. & Kohler, B. Nature 436, 1141–1144 (2005).
Markovitsi, D., Sharonov, A., Onidas, D. & Gustavsson, T. Chem. Phys. Chem. 3, 303–305 (2003).
Markovitsi, D., Onidas, D., Gustavsson, T., Talbot, F. & Lazzarotto, E. J. Am. Chem. Soc. 127, 17130–17131 (2005).
Bouvier, B., Gustavsson, T., Markovitsi, D. & Millié, P. Chem. Phys. 275, 75–92 (2002).
Bouvier, B. et al. J. Phys. Chem. B 107, 13512–13522 (2003).
Emanuele, E., Markovitsi, D., Millié, P. & Zakrzewska, K. Chem. Phys. Chem. 6, 1387–1392 (2005).
Emanuele, E., Zakrzewska, K., Markovitsi, D., Lavery, R. & Millié, P. J. Phys. Chem. B 109, 16109–16118 (2005).
Hartmann, B. & Lavery, R. Q. Rev. Biophys. 29, 309–368 (1996).
Ge, G. & Georghiou, S. Photochem. Photobiol. 54, 301–305 (1991).
Crespo-Hernández, C. E. & Kohler, B. J. Phys. Chem. B 108, 11182–11188 (2004).
Starikov, E. B., Lewis, J. P. & Sankey, O. F. Int. J. Mod. Phys. B 19, 4331–4357 (2005).
Kopka, M. L., Fratini, A. V., Drew, H. R. & Dickerson, R. E. J. Mol. Biol. 163, 129–146 (1983).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Markovitsi, D., Talbot, F., Gustavsson, T. et al. Complexity of excited-state dynamics in DNA. Nature 441, E7 (2006). https://doi.org/10.1038/nature04903
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04903
This article is cited by
-
Exciton Absorption and Luminescence in i-Motif DNA
Scientific Reports (2019)
-
Theoretical study on the photodegradation reaction of deca-BDE in THF in the presence of furan
Theoretical Chemistry Accounts (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.