Abstract
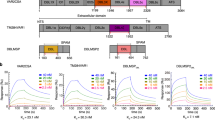
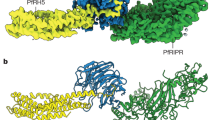
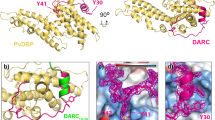
Molecular processes that govern pathogenic features of erythrocyte invasion and cytoadherence in malaria are reliant on Plasmodium-specific Duffy-binding-like domains (DBLs)1. These cysteine-rich modules recognize diverse host cell-surface receptors during pathogenesis. DBLs of parasite erythrocyte-binding proteins mediate invasion, and those from the antigenically variant P. falciparum erythrocyte membrane protein 1 (PfEMP1) have been implicated in cytoadherence. The simian and human malarial parasites, P. knowlesi and P. vivax, invade human erythrocytes exclusively through the host DARC receptor (Duffy antigen receptor for chemokines)2,3. Here we present the crystal structure of the P. knowlesi DBL domain (Pkα-DBL), which binds to DARC during invasion of human erythrocytes. Pkα-DBL retains the overall fold observed in DBLs from P. falciparum erythrocyte-binding antigen (EBA)-175 (ref. 4). Mapping the residues that have previously been implicated in binding5,6,7 highlights a fairly flat but exposed site for DARC recognition in subdomain 2 of Pkα-DBL; this is in sharp contrast to receptor recognition by EBA-175 (ref. 4). In Pkα-DBL, the residues that contact DARC and the clusters of residues under immune pressure map to opposite surfaces of the DBL, and suggest a possible mechanism for immune evasion by P. vivax. Our comparative structural analysis of Pkα-DBL and P. falciparum EBA-175 provides a framework for the understanding of malaria parasite DBLs, and may affect the development of new prophylactic and therapeutic strategies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Miller, L. H., Baruch, D. I., Marsh, K. & Doumbo, O. K. The pathogenic basis of malaria. Nature 415, 673–679 (2002)
Miller, L. H., Mason, S. J., Dvorak, J. A., McGinniss, M. H. & Rothman, I. K. Erythrocyte receptors for (Plasmodium knowlesi) malaria: Duffy blood group determinants. Science 189, 561–563 (1975)
Miller, L. H., Mason, S. J., Clyde, D. F. & McGinniss, M. H. The resistance factor to Plasmodium vivax in blacks. The Duffy-blood-group genotype, FyFy. N. Engl. J. Med. 295, 302–304 (1976)
Tolia, N. H., Enemark, E. J., Sim, B. K. & Joshua-Tor, L. Structural basis for the EBA-175 erythrocyte invasion pathway of the malaria parasite Plasmodium falciparum. Cell 122, 183–193 (2005)
Singh, S. K. et al. Definition of structural elements in Plasmodium vivax and P. knowlesi Duffy-binding domains necessary for erythrocyte invasion. Biochem. J. 374, 193–198 (2003)
Hans, D. et al. Mapping binding residues in the Plasmodium vivax domain that binds Duffy antigen during red cell invasion. Mol. Microbiol. 55, 1423–1434 (2005)
VanBuskirk, K. M., Sevova, E. & Adams, J. H. Conserved residues in the Plasmodium vivax Duffy-binding protein ligand domain are critical for erythrocyte receptor recognition. Proc. Natl Acad. Sci. USA 101, 15754–15759 (2004)
Snow, R. W., Guerra, C. A., Noor, A. M., Myint, H. Y. & Hay, S. I. The global distribution of clinical episodes of Plasmodium falciparum malaria. Nature 434, 214–217 (2005)
Choe, H. et al. Sulphated tyrosines mediate association of chemokines and Plasmodium vivax Duffy binding protein with the Duffy antigen/receptor for chemokines (DARC). Mol. Microbiol. 55, 1413–1422 (2005)
Farzan, M. et al. Tyrosine sulfation of the amino terminus of CCR5 facilitates HIV-1 entry. Cell 96, 667–676 (1999)
Tsuboi, T. et al. Natural variation within the principal adhesion domain of the Plasmodium vivax Duffy binding protein. Infect. Immun. 62, 5581–5586 (1994)
Xainli, J., Adams, J. H. & King, C. L. The erythrocyte binding motif of Plasmodium vivax Duffy binding protein is highly polymorphic and functionally conserved in isolates from Papua New Guinea. Mol. Biochem. Parasitol. 111, 253–260 (2000)
Ampudia, E., Patarroyo, M. A., Patarroyo, M. E. & Murillo, L. A. Genetic polymorphism of the Duffy receptor binding domain of Plasmodium vivax in Colombian wild isolates. Mol. Biochem. Parasitol. 78, 269–272 (1996)
Adams, J. H. et al. The Duffy receptor family of Plasmodium knowlesi is located within the micronemes of invasive malaria merozoites. Cell 63, 141–153 (1990)
Wyatt, R. et al. The antigenic structure of the HIV gp120 envelope glycoprotein. Nature 393, 705–711 (1998)
Kwong, P. D. et al. HIV-1 evades antibody-mediated neutralization through conformational masking of receptor-binding sites. Nature 420, 678–682 (2002)
Wilson, I. A. & Cox, N. J. Structural basis of immune recognition of influenza virus hemagglutinin. Annu. Rev. Immunol. 8, 737–771 (1990)
Mayor, A. et al. Receptor-binding residues lie in central regions of Duffy-binding-like domains involved in red cell invasion and cytoadherence by malaria parasites. Blood 105, 2557–2563 (2005)
Reed, M. B. et al. Targeted disruption of an erythrocyte binding antigen in Plasmodium falciparum is associated with a switch toward a sialic acid-independent pathway of invasion. Proc. Natl Acad. Sci. USA 97, 7509–7514 (2000)
Otwinowski, Z. & Minor, W. Processing of X-ray data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
de La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement in the MIR and MAD methods. Methods Enzymol. 276, 472–494 (1997)
Terwilliger, T. C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999)
Jones, T. A., Zou, J. Y. & Cowan, S. W. Kjeldgaard. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Brunger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check stereochemical quality of protein structure. J. Appl. Crystallogr. 26, 283–291 (1993)
Kleywegt, G. J. Experimental assessment of differences between related protein crystal structures. Acta Crystallogr. D 55, 1878–1884 (1999)
Esnouf, R. M. An extensively modified version of MolScript that includes greatly enhanced coloring capabilities. J. Mol. Graph. Model. 15, 132–134, 112–113 (1997)
Merritt, E. A. & Bacon, D. J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997)
Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991)
Brunger, A. T. The free R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472–474 (1992)
Acknowledgements
We thank D. Stuart and M. Walsh for facilitating data collection. We also thank past and present members of ICGEB, A. Rushdi Shakri and S. Krishnaswamy for help. C.E.C. is a Howard Hughes International Research Scholar. The Sharma laboratory is supported by a grant from the European Commission FP6. C.E.C. and A.S. are International Wellcome Trust Senior Research Fellows.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Atomic coordinates for Pkα-DBL have been deposited in the Protein Data Bank under accession number 2c6j. Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1 and 2. Supplementary Figure 1 details the disulphide structure of DBLs. Supplementary figure 2 details the specific binding of recombinant PkαDBL to human DARC (PDF 122 kb)
Rights and permissions
About this article
Cite this article
Singh, S., Hora, R., Belrhali, H. et al. Structural basis for Duffy recognition by the malaria parasite Duffy-binding-like domain. Nature 439, 741–744 (2006). https://doi.org/10.1038/nature04443
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04443
This article is cited by
-
PvDBPII elicits multiple antibody-mediated mechanisms that reduce growth in a Plasmodium vivax challenge trial
npj Vaccines (2024)
-
Genetic diversity and natural selection of Plasmodium vivax reticulocyte invasion genes in Ecuador
Malaria Journal (2023)
-
Structural basis for DARC binding in reticulocyte invasion by Plasmodium vivax
Nature Communications (2023)
-
Plasmodium knowlesi Duffy binding protein alpha region II (PkDBPαII) in clinical isolates from Peninsular Malaysia and Malaysian Borneo exhibit different immune responses in animal models
Parasitology Research (2022)
-
Population genetic analysis of the Plasmodium falciparum erythrocyte binding antigen-175 (EBA-175) gene in Equatorial Guinea
Malaria Journal (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.