Abstract
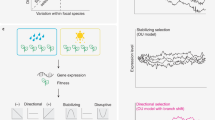
Mutation is the ultimate source of biological diversity because it generates the variation that fuels evolution1. Gene expression is the first step by which an organism translates genetic information into developmental change. Here we estimate the rate at which mutation produces new variation in gene expression by measuring transcript abundances across the genome during the onset of metamorphosis in 12 initially identical Drosophila melanogaster lines that independently accumulated mutations for 200 generations2. We find statistically significant mutational variation for 39% of the genome and a wide range of variability across corresponding genes. As genes are upregulated in development their variability decreases, and as they are downregulated it increases, indicating that developmental context affects the evolution of gene expression. A strong correlation between mutational variance and environmental variance shows that there is the potential for widespread canalization3. By comparing the evolutionary rates that we report here with differences between species4,5, we conclude that gene expression does not evolve according to strictly neutral models. Although spontaneous mutations have the potential to generate abundant variation in gene expression, natural variation is relatively constrained.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lynch, M. & Walsh, B. Genetics and Analysis of Quantitative Traits (Sinauer Associates, Sunderland, Massachusetts, 1998)
Houle, D. & Nuzhdin, S. V. Mutation accumulation and the effect of copia insertions in Drosophila melanogaster. Genet. Res. 83, 7–18 (2004)
Wagner, G. P., Booth, G. & Bagheri-Chaichian, H. A population genetic theory of canalization. Evolution 51, 329–347 (1997)
Rifkin, S. A., Kim, J. & White, K. P. Evolution of gene expression in the Drosophila melanogaster subgroup. Nature Genet. 33, 138–144 (2003)
Gu, Z. L., Rifkin, S. A., White, K. P. & Li, W. H. Duplicate genes increase gene expression diversity within and between species. Nature Genet. 36, 577–579 (2004)
Oleksiak, M. F., Roach, J. L. & Crawford, D. L. Natural variation in cardiac metabolism and gene expression in Fundulus heteroclitus. Nature Genet. 37, 67–72 (2004)
Morley, M. et al. Genetic analysis of genome-wide variation in human gene expression. Nature 430, 743–747 (2004)
Yang, H.-P., Tanikawa, A. Y. & Kondrashov, A. S. Molecular nature of 11 spontaneous de novo mutations in Drosophila melanogaster. Genetics 157, 1285–1292 (2001)
White, K. P., Rifkin, S. A., Hurban, P. & Hogness, D. S. Microarray analysis of Drosophila development during metamorphosis. Science 286, 2179–2184 (1999)
Li, T.-R. & White, K. P. Tissue-specific gene expression and ecdysone-regulated genomic networks in Drosophila. Dev. Cell 5, 59–72 (2003)
Lande, R. The maintenance of genetic variability by mutation in a polygenic character with linked loci. Genet. Res. 26, 221–235 (1976)
Houle, D., Morikawa, B. & Lynch, M. Comparing mutational variabilities. Genetics 143, 1467–1483 (1996)
Khaitovich, P. et al. A neutral model of transcriptome evolution. PLoS Biol. 2, e132 (2004)
Yanai, I., Graur, D. & Ophir, R. Incongruent expression profiles between human and mouse orthologous genes suggest widespread neutral evolution of transcription control. OMICS 8, 15–24 (2004)
Hsieh, W.-P., Chub, T.-M., Wolfinger, R. D. & Gibson, G. Mixed-model reanalysis of primate data suggests tissue and species biases in oligonucleotide-based gene expression profiles. Genetics 165, 747–757 (2003)
Lemos, B., Meiklejohn, C. D., Caceres, M. & Hartl, D. L. Rates of divergence in gene expression profiles of primates, mice, and flies: stabilizing selection and variability among functional categories. Evolution 59, 126–137 (2005)
Denver, D. R. et al. The transcriptional consequences of mutation and natural selection in Caenorhabditis elegans. Nature Genet. 37, 544–548 (2005)
Lynch, M. & Hill, W. G. Phenotypic evolution by neutral mutation. Evolution 40, 915–935 (1986)
Waddington, C. H. Genetic assimilation of an acquired character. Evolution 7, 118–126 (1953)
Meiklejohn, C. D. & Hartl, D. L. A single mode of canalization. Trends Ecol. Evol. 17, 468–473 (2002)
Houle, D. How should we explain variation in the genetic variance of traits? Genetica 103, 241–253 (1998)
Lynch, M. et al. Perspective: spontaneous deleterious mutation. Evolution 53, 645–663 (1999)
Kacser, H. & Burns, J. The control of flux. Symp. Soc. Exp. Biol. 27, 65–104 (1973)
Hartl, D., Dykhuizen, D. & Dean, A. Limits of adaptation: the evolution of selective neutrality. Genetics 111, 655–674 (1985)
Benton, M. J. & Ayala, F. J. Dating the tree of life. Science 300, 1698–1700 (2003)
Wolfinger, R. et al. Assessing gene significance from cDNA microarray expression data via mixed models. J. Comput. Biol. 8, 625–637 (2001)
Littell, R. C., Milliken, G. A., Stroup, W. W. & Wolfinger, R. D. SAS System for Mixed Models (SAS Institute, Cary, North Carolina, 1996)
Fry, J. D. in Genetic Analysis of Complex Traits Using SAS (ed. Saxton, A.) 11–34 (SAS Institute, Cary, North Carolina, 2004)
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995)
Sokal, R. R. & Rohlf, F. J. Biometry (W. H. Freeman and Company, New York, 1995)
Acknowledgements
We thank G. Wagner, S. Rice, J. Powell, M. Lynch, J. Fry, G. Gibson, P. Gibbs, T.-R. Li, J. D. Lambert and members of the Kim and White laboratories for suggestions, help and comments. This work was supported by a National Library of Medicine fellowship (to S.A.R.), an NIH grant (to J.K.), and grants from the W.M. Keck Foundation, the Arnold and Mabel Beckman Foundation, and the NIH and National Human Genome Research Institute (to K.P.W.). Author Contributions S.A.R. and J.K. planned and designed the project in consultation with D.H. and K.P.W. Expression data was collected by S.A.R. using spotted microarrays developed by K.P.W. D.H. generated and maintained the mutation accumulation lines. S.A.R. and J.K. developed the computational analyses and carried out the quantitative genetics modelling. S.A.R. wrote the paper with contributions from all authors.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
Microarray data have been deposited in the Gene Expression Omnibus under accession numbers GSE2126 (mutation accumulation lines), GSE2641 (technical error) and GSE2642 (comparative data update and extension). Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Data 1
Estimates from the genome-wide mixed model analyses of the full and jackknifed datasets. (TXT 4289 kb)
Supplementary Data 2
SAS PROC MIXED calls for the model hierarchy. (DOC 35 kb)
Supplementary Methods
Additional details of the methods used in this study. (PDF 57 kb)
Supplementary Figure 1
Schematic of sample collection. (DOC 393 kb)
Supplementary Figure 2
Mutational heritability versus expression level. (DOC 72 kb)
Supplementary Figure 3
Experimental design. (DOC 192 kb)
Supplementary Figure 4
Likelihood ratio tests order along the model hierarchy. (DOC 220 kb)
Rights and permissions
About this article
Cite this article
Rifkin, S., Houle, D., Kim, J. et al. A mutation accumulation assay reveals a broad capacity for rapid evolution of gene expression. Nature 438, 220–223 (2005). https://doi.org/10.1038/nature04114
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04114
This article is cited by
-
The impact of species-wide gene expression variation on Caenorhabditis elegans complex traits
Nature Communications (2022)
-
Molecular and evolutionary processes generating variation in gene expression
Nature Reviews Genetics (2021)
-
Long-term experimental evolution reveals purifying selection on piRNA-mediated control of transposable element expression
BMC Biology (2020)
-
Immigration shapes evolutionary tolerance to toxic cyanobacteria in two cladoceran grazers
Hydrobiologia (2019)
-
The genesis of an exceptionally lethal venom in the timber rattlesnake (Crotalus horridus) revealed through comparative venom-gland transcriptomics
BMC Genomics (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.