Abstract
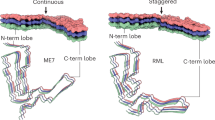
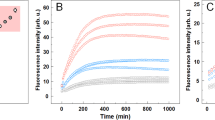
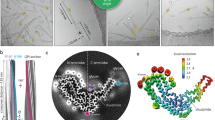
Prions are believed to be infectious, self-propagating polymers of otherwise soluble, host-encoded proteins1,2. This concept is now strongly supported by the recent findings that amyloid fibrils of recombinant prion proteins from yeast3,4,5, Podospora anserina6 and mammals7 can induce prion phenotypes in the corresponding hosts. However, the structural basis of prion infectivity remains largely elusive because acquisition of atomic resolution structural properties of amyloid fibrils represents a largely unsolved technical challenge. HET-s, the prion protein of P. anserina, contains a carboxy-terminal prion domain comprising residues 218–289. Amyloid fibrils of HET-s(218–289) are necessary and sufficient for the induction and propagation of prion infectivity6. Here, we have used fluorescence studies, quenched hydrogen exchange NMR and solid-state NMR to determine the sequence-specific positions of amyloid fibril secondary structure elements of HET-s(218–289). This approach revealed four β-strands constituted by two pseudo-repeat sequences, each forming a β-strand-turn-β-strand motif. By using a structure-based mutagenesis approach, we show that this conformation is the functional and infectious entity of the HET-s prion. These results correlate distinct structural elements with prion infectivity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Alper, T., Cramp, W. A., Haig, D. A. & Clarke, M. C. Does the agent of scrapie replicate without nucleic acid? Nature 214, 764–766 (1967)
Prusiner, S. B. Novel proteinaceous infectious particles cause scrapie. Science 216, 136–144 (1982)
Sparrer, H. E., Santoso, A., Szoka, F. C. Jr & Weissman, J. S. Evidence for the prion hypothesis: induction of the yeast [PSI + ] factor by in vitro-converted Sup35 protein. Science 289, 595–599 (2000)
King, C. Y. & Diaz-Avalos, R. Protein-only transmission of three yeast prion strains. Nature 428, 319–323 (2004)
Tanaka, M., Chien, P., Naber, N., Cooke, R. & Weissman, J. S. Conformational variations in an infectious protein determine prion strain differences. Nature 428, 323–328 (2004)
Maddelein, M. L., Dos Reis, S., Duvezin-Caubet, S., Coulary-Salin, B. & Saupe, S. J. Amyloid aggregates of the HET-s prion protein are infectious. Proc. Natl Acad. Sci. USA 99, 7402–7407 (2002)
Legname, G. et al. Synthetic mammalian prions. Science 305, 673–676 (2004)
Glass, N. L. & Kaneko, I. Fatal attraction: nonself recognition and heterokaryon incompatibility in filamentous fungi. Eukaryot. Cell 2, 1–8 (2003)
Saupe, S. J. Molecular genetics of heterokaryon incompatibility in filamentous ascomycetes. Microbiol. Mol. Biol. Rev. 64, 489–502 (2000)
Turcq, B., Deleu, C., Denayrolles, M. & Begueret, J. Two allelic genes responsible for vegetative incompatibility in the fungus Podospora anserina are not essential for cell viability. Mol. Gen. Genet. 228, 265–269 (1991)
Coustou, V., Deleu, C., Saupe, S. & Begueret, J. The protein product of the het-s heterokaryon incompatibility gene of the fungus Podospora anserina behaves as a prion analog. Proc. Natl Acad. Sci. USA 94, 9773–9778 (1997)
Dos Reis, S. et al. The HET-s prion protein of the filamentous fungus Podospora anserina aggregates in vitro into amyloid-like fibrils. J. Biol. Chem. 277, 5703–5706 (2002)
Balguerie, A. et al. Domain organization and structure-function relationship of the HET-s prion protein of Podospora anserina. EMBO J. 22, 2071–2081 (2003)
Coustou-Linares, V., Maddelein, M. L., Begueret, J. & Saupe, S. J. In vivo aggregation of the HET-s prion protein of the fungus Podospora anserina. Mol. Microbiol. 42, 1325–1335 (2001)
Balguerie, A. et al. The sequences appended to the amyloid core region of the HET-s prion protein determine higher-order aggregate organization in vivo. J. Cell Sci. 117, 2599–2610 (2004)
Hoshino, M. et al. Mapping the core of the β2-microglobulin amyloid fibril by H/D exchange. Nature Struct. Biol. 9, 332–336 (2002)
Lührs, T. et al. The 3D structure of Alzheimer's Aβ(1–42) fibrils. Nature (submitted)
Verel, R., Ernst, M. & Meier, B. H. Adiabatic dipolar recoupling in solid-state NMR: The DREAM scheme. J. Magn. Reson. 150, 81–99 (2001)
Siemer, A. B., Ritter, C., Ernst, M., Riek, R. & Meier, B. H. High-resolution solid-state NMR of the prion protein HET-s in its amyloid conformation. Angew. Chem. Int. Edn Engl. 44, 2441–2444 (2005)
Wishart, D. S. & Sykes, B. D. The 13C chemical-shift index: a simple method for the identification of protein secondary structure using 13C chemical-shift data. J. Biomol. NMR 4, 171–180 (1994)
Javitch, J. A., Shi, L. & Liapakis, G. Use of the substituted cysteine accessibility method to study the structure and function of G protein-coupled receptors. Methods Enzymol. 343, 137–156 (2002)
Tycko, R. Progress towards a molecular-level structural understanding of amyloid fibrils. Curr. Opin. Struct. Biol. 14, 96–103 (2004)
Petkova, A. T. et al. Self-propagating, molecular-level polymorphism in Alzheimer's β-amyloid fibrils. Science 307, 262–265 (2005)
Laws, D. D. et al. Solid-state NMR studies of the secondary structure of a mutant prion protein fragment of 55 residues that induces neurodegeneration. Proc. Natl Acad. Sci. USA 98, 11686–11690 (2001)
Yamaguchi, K. et al. Core and heterogeneity of β2-microglobulin amyloid fibrils as revealed by H/D exchange. J. Mol. Biol. 338, 559–571 (2004)
Harper, J. D. & Lansbury, P. T. Jr Models of amyloid seeding in Alzheimer's disease and scrapie: mechanistic truths and physiological consequences of the time-dependent solubility of amyloid proteins. Annu. Rev. Biochem. 66, 385–407 (1997)
Grzesiek, S. et al. 1H, 13C, and 15N NMR backbone assignments and secondary structure of human interferon-γ. Biochemistry 31, 8180–8190 (1992)
Bracken, C., Palmer, A. G. III & Cavanagh, J. (H)N(COCA)NH and HN(COCA)NH experiments for 1H–15N backbone assignments in 13C/15N-labeled proteins. J. Biomol. NMR 9, 94–100 (1997)
Guntert, P., Dotsch, V., Wider, G. & Wuthrich, K. Processing of multidimensional NMR data with the new software Prosa. J. Biomol. NMR 2, 619–629 (1992)
Samoson, A., Tuherm, T. & Past, J. Rotation sweep NMR. Chem. Phys. Lett. 365, 292–299 (2002)
Acknowledgements
R.R. is a Pew scholar. This research was supported in part by grants from the National Institute of Health, the US Army, the ETH Zurich, the Swiss National Science Foundation, the CNRS and the French Ministry of Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
Contains Supplementary Figures S1-S5 (additional solid and liquid state NMR data, sequence alignments and EM pictures), legends to accompany the figures and Supplementary Tables S1 and S2 giving details of the HET-s infectivity and function assays. (PDF 2848 kb)
Rights and permissions
About this article
Cite this article
Ritter, C., Maddelein, ML., Siemer, A. et al. Correlation of structural elements and infectivity of the HET-s prion. Nature 435, 844–848 (2005). https://doi.org/10.1038/nature03793
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03793
This article is cited by
-
The Possible Role of Prion-Like Viral Protein Domains on the Emergence of Novel Viruses as SARS-CoV-2
Journal of Molecular Evolution (2022)
-
Advances in protein misfolding, amyloidosis and its correlation with human diseases
3 Biotech (2020)
-
A new era for understanding amyloid structures and disease
Nature Reviews Molecular Cell Biology (2018)
-
A 31-residue peptide induces aggregation of tau's microtubule-binding region in cells
Nature Chemistry (2017)
-
Melanosomal formation of PMEL core amyloid is driven by aromatic residues
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.