Abstract
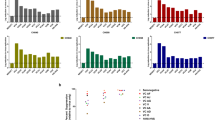
The extreme polymorphism in the human leukocyte antigen (HLA) class I region of the human genome is suggested to provide an advantage in pathogen defence mediated by CD8+ T cells1,2,3. HLA class I molecules present pathogen-derived peptides on the surface of infected cells for recognition by CD8+ T cells. However, the relative contributions of HLA-A and -B alleles have not been evaluated. We performed a comprehensive analysis of the class I restricted CD8+ T-cell responses against human immunodeficiency virus (HIV-1), immune control of which is dependent upon virus-specific CD8+ T-cell activity4,5. In 375 HIV-1-infected study subjects from southern Africa, a significantly greater number of CD8+ T-cell responses are HLA-B-restricted, compared to HLA-A (2.5-fold; P = 0.0033). Here we show that variation in viral set-point, in absolute CD4 count and, by inference, in rate of disease progression in the cohort, is strongly associated with particular HLA-B but not HLA-A allele expression (P < 0.0001 and P = 0.91, respectively). Moreover, substantially greater selection pressure is imposed on HIV-1 by HLA-B alleles than by HLA-A (4.4-fold, P = 0.0003). These data indicate that the principal focus of HIV-specific activity is at the HLA-B locus. Furthermore, HLA-B gene frequencies in the population are those likely to be most influenced by HIV disease, consistent with the observation that B alleles evolve more rapidly than A alleles6,7,8. The dominant involvement of HLA-B in influencing HIV disease outcome is of specific relevance to the direction of HIV research and to vaccine design.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Jeffery, K. J. & Bangham, C. R. Do infectious diseases drive MHC diversity? Microbes Infect. 2, 1335–1341 (2000)
Parham, P. & Ohta, T. Population biology of antigen presentation by MHC class I molecules. Science 272, 67–74 (1996)
Marsh, S. G. E., Parham, P. & Barber, L. D. The HLA Facts Book (Academic, London, 2000)
Carrington, M. et al. HLA and HIV-1: heterozygote advantage and B*35-Cw*04 disadvantage. Science 283, 1748–1752 (1999)
McMichael, A. J. & Rowland-Jones, S. L. Cellular immune responses to HIV. Nature 410, 980–987 (2001)
Belich, M. P. et al. Unusual HLA-B alleles in two tribes of Brazilian Indians. Nature 357, 326–329 (1992)
Watkins, D. I. et al. New recombinant HLA-B alleles in a tribe of South American Amerindians indicate rapid evolution of MHC class I loci. Nature 357, 329–333 (1992)
McAdam, S. N. et al. A uniquely high level of recombination at the HLA-B locus. Proc. Natl Acad. Sci. USA 91, 5893–5897 (1994)
Snary, D., Barnstable, C. J., Bodmer, W. F. & Crumpton, M. J. Molecular structure of human histocompatibility antigens: the HLA-C series. Eur. J. Immunol. 7, 580–585 (1977)
Addo, M. M. et al. Comprehensive epitope analysis of human immunodeficiency virus type 1 (HIV-1)-specific T-cell responses directed against the entire expressed HIV-1 genome demonstrate broadly directed responses, but no correlation to viral load. J. Virol. 77, 2081–2092 (2003)
Mellors, J. W. et al. Prognosis in HIV-1 infection predicted by the quantity of virus in plasma. Science 272, 1167–1170 (1996)
Badri, M. et al. Initiating highly active antiretroviral therapy in sub-Saharan Africa: an assessment of the revised World Health Organization scaling-up guidelines. AIDS 18, 1159–1168 (2004)
Goulder, P. J. & Watkins, D. I. HIV and SIV CTL escape: implications for vaccine design. Nature Rev. Immunol. 4, 630–640 (2004)
Moore, C. B. et al. Evidence of HIV-1 adaptation to HLA-restricted immune responses at a population level. Science 296, 1439–1443 (2002)
Cooper, E. R. et al. Combination antiretroviral strategies for the treatment of pregnant HIV-1-infected women and prevention of perinatal HIV-1 transmission. J. Acquir. Immune Defic. Syndr. 29, 484–494 (2002)
Garcia, P. M. et al. Maternal levels of plasma human immunodeficiency virus type 1 RNA and the risk of perinatal transmission. Women and Infants Transmission Study Group. N. Engl. J. Med. 341, 394–402 (1999)
Kaslow, R. A. et al. Influence of combinations of human major histocompatibility complex genes on the course of HIV-1 infection. Nature Med. 2, 405–411 (1996)
O'Brien, S. J., Gao, X. & Carrington, M. HLA and AIDS: a cautionary tale. Trends Mol. Med. 7, 379–381 (2001)
Migueles, S. A. et al. HLA B*5701 is highly associated with restriction of virus replication in a subgroup of HIV-infected long term nonprogressors. Proc. Natl Acad. Sci. USA 97, 2709–2714 (2000)
Gao, X. et al. Effect of a single amino acid change in MHC class I molecules on the rate of progression to AIDS. N. Engl. J. Med. 344, 1668–1675 (2001)
Trachtenberg, E. et al. Advantage of rare HLA supertype in HIV disease progression. Nature Med. 9, 928–935 (2003)
Flores-Villanueva, P. O. et al. Control of HIV-1 viremia and protection from AIDS are associated with HLA-Bw4 homozygosity. Proc. Natl Acad. Sci. USA 98, 5140–5145 (2001)
Martin, M. P. et al. Epistatic interaction between KIR3DS1 and HLA-B delays the progression to AIDS. Nature Genet. 31, 429–434 (2002)
Cooke, G. S. & Hill, A. V. Genetics of susceptibility to human infectious disease. Nature Rev. Genet. 2, 967–977 (2001)
Ramos, M. & Lopez de Castro, J. A. HLA-B27 and the pathogenesis of spondyloarthritis. Tissue Antigens 60, 191–205 (2002)
Thio, C. L. et al. Comprehensive analysis of class I and class II HLA antigens and chronic hepatitis B virus infection. J. Virol. 77, 12083–12087 (2003)
Bunce, M. PCR-sequence-specific primer typing of HLA class I and class II alleles. Methods Mol. Biol. 210, 143–171 (2003)
Leslie, A. J. et al. HIV evolution: CTL escape mutation and reversion after transmission. Nature Med. 10, 282–289 (2004)
Acknowledgements
We thank those who made possible the collection of blood samples for the study, in particular the patients and staff at Cato Manor Clinic and Sinikithemba Clinic, Durban, South Africa. We also thank S. McAdam, J. Webster and R. Phillips for comments and suggestions. This work was supported by a contract from the National Institutes of Health (B.D.W.), with additional support from the Wellcome Trust (P.J.R.G., A.J.L., P.Z., P.K.), the Elizabeth Glaser Pediatric AIDS Foundation (P.J.R.G., B.T.K.), the National Institutes of Health and the Doris Duke Charitable Foundation. P.J.R.G. and B.T.K. are Elizabeth Glaser Pediatric AIDS Foundation Scientists, B.D.W. is a Doris Duke Distinguished Science Professor.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Information Table 1
Peptides recognized in association with individual HLA class I allele expression. (PDF 68 kb)
Supplementary Information Table 2
List of the 69 associations identified between HLA class I expression and overlapping peptide targeted, for which 0.05<p<0.001. (PDF 62 kb)
Supplementary Information Table 3
Linkage disequilibrium involving the alleles that were identified in association with high or low viral loads. (PDF 41 kb)
Supplementary Information Figure 1
HLA restriction and epitope optimisation for 10 additional epitopes where the HLA-peptide association was identified by the statistical approach described in the text. (PDF 75 kb)
Supplementary Information Figure 2
HLA class I molecule expression and absolute CD4 count in chronically infected Zulu/Xhosa (n=706). (PDF 50 kb)
Supplementary Information Methods
Detailed description of the statistical methods used in this study. (DOC 25 kb)
Supplementary Data
GenBank accession number information. (DOC 19 kb)
Rights and permissions
About this article
Cite this article
Kiepiela, P., Leslie, A., Honeyborne, I. et al. Dominant influence of HLA-B in mediating the potential co-evolution of HIV and HLA. Nature 432, 769–775 (2004). https://doi.org/10.1038/nature03113
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03113
This article is cited by
-
Prognostic impact of HLA supertype mismatch in single-unit cord blood transplantation
Bone Marrow Transplantation (2024)
-
Innate receptors modulating adaptive T cell responses: KIR-HLA interactions and T cell-mediated control of chronic viral infections
Immunogenetics (2023)
-
Africa-specific human genetic variation near CHD1L associates with HIV-1 load
Nature (2023)
-
Immunoescape of HIV-1 in Env-EL9 CD8 + T cell response restricted by HLA-B*14:02 in a Non progressor who lost twenty-seven years of HIV-1 control
Retrovirology (2022)
-
Allelic imbalance of HLA-B expression in human lung cells infected with coronavirus and other respiratory viruses
European Journal of Human Genetics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.