Abstract
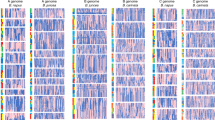
The architecture of higher plants is established through the activity of lateral meristems—small groups of stem cells formed during vegetative and reproductive development. Lateral meristems generate branches and inflorescence structures, which define the overall form of a plant1,2,3, and are largely responsible for the evolution of different plant architectures3. Here, we report the isolation of the barren stalk1 gene, which encodes a non-canonical basic helix–loop–helix protein required for the initiation of all aerial lateral meristems in maize. barren stalk1 represents one of the earliest genes involved in the patterning of maize inflorescences, and, together with the teosinte branched1 gene4, it regulates vegetative lateral meristem development. The architecture of maize has been a major target of selection for early agriculturalists and modern farmers, because it influences harvesting, breeding strategies and mechanization. By sampling nucleotide diversity in the barren stalk1 region, we show that two haplotypes entered the maize gene pool from its wild progenitor, teosinte, and that only one was incorporated throughout modern inbreds, suggesting that barren stalk1 was selected for agronomic purposes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Weigel, D. & Jürgens, G. Stem cells that make stems. Nature 415, 751–754 (2002)
Komatsu, K. et al. LAX and SPA: major regulators of shoot branching in rice. Proc. Natl Acad. Sci. USA 100, 11765–11770 (2003)
Sussex, I. M. & Kerk, N. M. The evolution of plant architecture. Curr. Opin. Plant Biol. 4, 33–37 (2001)
Doebley, J., Stec, A. & Hubbard, L. The evolution of apical dominance in maize. Nature 386, 485–488 (1997)
Hofmeyr, J. D. J. The Inheritance and Linkage Relationships of barren stalk-1 and barren stalk-2, Two Mature-Plant Characters of Maize. Thesis, Cornell Univ., Ithaca, New York (1931)
Ritter, M. K., Padilla, C. M. & Schmidt, R. J. The maize mutant barren stalk1 is defective in axillary meristem development. Am. J. Bot. 89, 203–210 (2002)
Kapitonov, V. V. & Jurka, J. Rolling-circle transposons in eukaryotes. Proc. Natl Acad. Sci. USA 98, 8714–8719 (2001)
Massari, M. E. & Murre, C. Helix-loop-helix proteins: regulators of transcription in eukaryotic organisms. Mol. Cell. Biol. 20, 429–440 (2000)
Toledo-Ortiz, G. E., Huq, E. & Quail, P. H. The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell 15, 1749–1770 (2003)
Liljegren, S. J. et al. Control of fruit patterning in Arabidopsis by INDEHISCENT. Cell 116, 843–853 (2004)
Cheng, P. C., Greyson, R. I. & Walden, D. B. Organ initiation and the development of unisexual flowers in the tassel and ear of Zea mays. Am. J. Bot. 70, 450–462 (1983)
Irish, E. E. Class II tassel seed mutations provide evidence for multiple types of inflorescence meristems in maize (Poaceae). Am. J. Bot. 84, 1502–1515 (1997)
Ambrose, B. A. et al. Molecular and genetic analysis of the silky1 gene reveal conservation in floral organ specification between eudicots and monocots. Mol. Cell 5, 569–579 (2000)
Taguchi-Shiobara, F., Yuan, Z., Hake, S. & Jackson, D. The fasciated ear2 gene encodes a leucine-rich repeat receptor-like protein that regulates shoot meristem proliferation in maize. Genes Dev. 15, 2755–2766 (2001)
McSteen, P. & Hake, S. barren inflorescence2 regulates axillary meristem development in the maize inflorescence. Development 128, 2881–2891 (2001)
Reinhardt, D. et al. Regulation of phyllotaxis by polar auxin transport. Nature 426, 255–260 (2003)
Benkova, E. et al. Local, efflux-dependent auxin gradients as a common module for plant organ formation. Cell 115, 591–602 (2003)
Buckler, E. S. IV, Thornsberry, J. M. & Kresovich, S. Molecular diversity, structure and domestication of grasses. Genet. Res. Camb. 77, 213–218 (2001)
Doebley, J., Stec, A. & Gustus, C. teosinte branched1 and the origin of maize: evidence for epistasis and the evolution of dominance. Genetics 141, 333–346 (1995)
Wang, R. L., Stec, A., Hey, J., Lukens, L. & Doebley, J. The limits of selection during maize domestication. Nature 398, 236–239 (1999)
Hubbard, L., McSteen, P., Doebley, J. & Hake, S. Expression pattern and mutant phenotype of teosinte branched1 correlate with growth suppression in maize and teosinte. Genetics 162, 1927–1935 (2002)
Clark, R. M., Linton, E., Messing, J. & Doebley, J. F. Pattern of diversity in the genomic region near the maize domestication gene tb1. Proc. Natl Acad. Sci. USA 101, 700–707 (2004)
Lukens, L. & Doebley, J. Epistatic and environmental interactions for quantitative trait loci involved in maize evolution. Genet. Res. Camb. 74, 291–302 (1999)
Tenaillon, M. I. et al. Patterns of DNA sequence polymorphism along chromosome 1 of maize (Zea mays ssp. mays L.). Proc. Natl Acad. Sci. USA 98, 9161–9166 (2001)
Tajima, F. Statistical method for testing neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989)
Hudson, R., Kreitman, M. & Aguade, M. A test of neutral molecular evolution based on nucleotide data. Genetics 116, 153–159 (1987)
Whitt, S. R., Wilson, L. M., Tenaillon, M. I., Gaut, B. S. & Buckler, E. S. IV Genetic diversity and selection in the maize starch pathway. Proc. Natl Acad. Sci. USA 99, 12959–12962 (2002)
Bensen, R. J. et al. Cloning and characterization of the maize An1 gene. Plant Cell 7, 75–84 (1995)
Dinneny, J. R., Yadegari, R., Fischer, R. L., Yanofsky, M. F. & Weigel, D. The role of JAGGED in shaping lateral organs. Development 131, 1101–1110 (2004)
Rozas, J. & Rozas, R. DnaSP version 3: an integrated program for molecular population genetics and molecular evolution analysis. Bioinformatics 15, 174–175 (1999)
Acknowledgements
We thank C. J. Whipple for the pictures in Figs 1m and 3b, c, and for discussions; M. Zanis and S. Jeong for critical reading of the manuscript; M. J. Galli for suggestions on quantitative PCR; E. York for assistance with SEMs at the Scripps Institution of Oceanography Analytical Facility; and A. Tsai, E. Durbin and D. Nakamura for technical help. This research was supported by NSF and NIH grants to R.J.S. and J.F.D. A.G. was also supported by MIUR, Ministero dell'Istruzione, dell'Universitá e della Ricerca, Italy.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare they have no competing financial interests.
Supplementary information
Supplementary Data
We provide further details about the CAPS analysis on the 86 inbred lines. (DOC 20 kb)
Supplementary Methods
We describe in detail the isolation of new ba1 alleles, the amplification of regions a, b, c and the CAPS marker analysis on the 86 inbred lines analyzed for haplotype I and II. (DOC 22 kb)
Supplementary Table 1
Supplementary Table 1 is part of the Methods section and lists all the maize and teosinte individuals sequenced at the barren stalk1 gene with the corresponding GenBank Accessions Numbers. (DOC 44 kb)
Rights and permissions
About this article
Cite this article
Gallavotti, A., Zhao, Q., Kyozuka, J. et al. The role of barren stalk1 in the architecture of maize. Nature 432, 630–635 (2004). https://doi.org/10.1038/nature03148
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03148
This article is cited by
-
Hybrid allele-specific ChIP-seq analysis identifies variation in brassinosteroid-responsive transcription factor binding linked to traits in maize
Genome Biology (2023)
-
Accelerating crop domestication through genome editing for sustainable agriculture
Journal of Plant Biochemistry and Biotechnology (2023)
-
Harnessing the role of genes involved in plant architectural changes
Plant Growth Regulation (2023)
-
Deploying QTL-seq rapid identification and separation of the major QTLs of tassel branch number for fine-mapping in advanced maize populations
Molecular Breeding (2023)
-
The genetic basis for panicle trait variation in switchgrass (Panicum virgatum)
Theoretical and Applied Genetics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.