Abstract
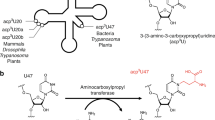
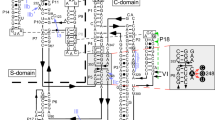
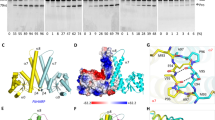
Transfer RNA nucleotidyltransferases (CCA-adding enzymes) are responsible for the maturation or repair of the functional 3′ end of tRNAs by means of the addition of the essential nucleotides CCA. However, it is unclear how tRNA nucleotidyltransferases polymerize CCA onto the 3′ terminus of immature tRNAs without using a nucleic acid template. Here we describe the crystal structure of the Archaeoglobus fulgidus tRNA nucleotidyltransferase in complex with tRNA. We also present ternary complexes of this enzyme with both RNA duplex mimics of the tRNA acceptor stem that terminate with the nucleotides C74 or C75, as well as the appropriate incoming nucleoside 5′-triphosphates. A single nucleotide-binding pocket exists whose specificity for both CTP and ATP is determined by the protein side chain of Arg 224 and backbone phosphates of the tRNA, which are non-complementary to and thus exclude UTP and GTP. Discrimination between CTP or ATP at a given addition step and at termination arises from changes in the size and shape of the nucleotide binding site that is progressively altered by the elongating 3′ end of the tRNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Deutscher, M. P. tRNA nucleotidyltransferase. The Enzymes 15, 183–215 (1982)
Green, R. & Noller, H. F. Ribosomes and translation. Annu. Rev. Biochem. 66, 679–716 (1997)
Nissen, P., Hansen, J., Ban, N., Moore, P. B. & Steitz, T. A. The structural basis of ribosome activity in peptide bond synthesis. Science 289, 920–930 (2000)
Aebi, M. et al. Isolation of a temperature-sensitive mutant with an altered tRNA nucleotidyltransferase and cloning of the gene encoding tRNA nucleotidyltransferase in the yeast Saccharomyces cerevisiae. J. Biol. Chem. 265, 16216–16220 (1990)
Zhu, L. & Deutscher, M. P. tRNA nucleotidyltransferase is not essential for Escherichia coli viability. EMBO J. 6, 2473–2477 (1987)
Holm, L. & Sander, C. DNA polymerase β belongs to an ancient nucleotidyltransferase superfamily. Trends Biochem. Sci. 20, 345–347 (1995)
Yue, D., Maizels, N. & Weiner, A. M. CCA-adding enzymes and poly(A) polymerases are all members of the same nucleotidyltransferase superfamily: characterization of the CCA-adding enzyme from the archaeal hyperthermophile Sulfolobus shibatae. RNA 2, 895–908 (1996)
Li, F. et al. Crystal structures of the Bacillus stearothermophilus CCA-adding enzyme and its complexes with ATP or CTP. Cell 111, 815–824 (2002)
Augustin, M. A. et al. Crystal structure of the human CCA-adding enzyme: insights into template-independent polymerization. J. Mol. Biol. 328, 985–994 (2003)
Xiong, Y., Li, F., Wang, J., Weiner, A. M. & Steitz, T. A. Crystal structures of an archaeal class I CCA-adding enzyme and its nucleotide complexes. Mol. Cell 12, 1165–1172 (2003)
Okabe, M. et al. Divergent evolutions of trinucleotide polymerization revealed by an archaeal CCA-adding enzyme structure. EMBO J. 22, 5918–5927 (2003)
Steitz, T. A. A mechanism for all polymerases. Nature 391, 231–232 (1998)
Deutscher, M. P. Reactions at the 3′ terminus of transfer ribonucleic acid. 3. Catalytic properties of two purified rabbit liver transfer ribonucleic acid nucleotidyl transferases. J. Biol. Chem. 247, 459–468 (1972)
Tomari, Y., Suzuki, T., Watanabe, K. & Ueda, T. The role of tightly bound ATP in Escherichia coli tRNA nucleotidyltransferase. Genes Cells 5, 689–698 (2000)
Seth, M., Thurlow, D. L. & Hou, Y. M. Poly(C) synthesis by class I and class II CCA-adding enzymes. Biochemistry 41, 4521–4532 (2002)
Yue, D., Weiner, A. M. & Maizels, N. The CCA-adding enzyme has a single active site. J. Biol. Chem. 273, 29693–29700 (1998)
Shi, P. Y., Maizels, N. & Weiner, A. M. CCA addition by tRNA nucleotidyltransferase: polymerization without translocation? EMBO J. 17, 3197–3206 (1998)
Shi, P. Y., Weiner, A. M. & Maizels, N. A top-half tDNA minihelix is a good substrate for the eubacterial CCA-adding enzyme. RNA 4, 276–284 (1998)
Pelletier, H., Sawaya, M. R., Kumar, A., Wilson, S. H. & Kraut, J. Structures of ternary complexes of rat DNA polymerase beta, a DNA template-primer, and ddCTP. Science 264, 1891–1903 (1994)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Navaza, J. Implementation of molecular replacement in AMoRe. Acta Crystallogr. D 57, 1367–1372 (2001)
Terwilliger, T. C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Cowtan, K. ’dm’: An automated procedure for phase improvement by density modification. Joint CCP4 ESF-EACBM Newsl. Prot. Crystallogr. 31, 34–38 (1994)
Kleywegt, G. J. & Jones, T. A. Software for handling macromolecular envelopes. Acta Crystallogr. D 55, 941–944 (1999)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Tomita, K. et al. Structural basis for template-independent RNA polymerization. Nature doi:10.1038/nature02712 (this issue)
Acknowledgements
We thank the beamline staff at X25 and X12C of the National Synchrotron Light Source, 8BM at the Advanced Photon Source, A1 at the Cornell High Energy Synchrotron Source, and 8.2.1 and 8.2.2 at the Advanced Light Source for assistance in data collection. We thank A. M. Weiner for suggestions on the manuscript; J. Wang for discussions; members of the Steitz laboratory for assistance at various stages of the project; and R. Evans for work with the ACC75 complex. This work was supported by an NIH grant.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Figure S1
Regions of the 6.5 Å resolution electron density map for the AfCCA-tRNA complex calculated using 2Fo-Fc as coefficients and phases improved by solvent flattening and non-crystallographic symmetry averaging. (JPG 106 kb)
Supplementary Figure S2
Electron density from solvent-flattened and NCS-averaged 2Fo-Fc maps. (JPG 85 kb)
Supplementary Figure S3
The metal ion geometries in the AC74 (a) and ACC75 (b) complexes. (JPG 87 kb)
Supplementary Movie 1
Two tRNA molecules bound to the AfCCA dimer. (MP4 369 kb)
Supplementary Movie 2
The process of the addition of the last two nucleotides by the CCA-adding enzyme. (MP4 1999 kb)
Rights and permissions
About this article
Cite this article
Xiong, Y., Steitz, T. Mechanism of transfer RNA maturation by CCA-adding enzyme without using an oligonucleotide template. Nature 430, 640–645 (2004). https://doi.org/10.1038/nature02711
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02711
This article is cited by
-
poly(UG)-tailed RNAs in genome protection and epigenetic inheritance
Nature (2020)
-
Unbiased screen of RNA tailing activities reveals a poly(UG) polymerase
Nature Methods (2019)
-
Structural mechanism of cytosolic DNA sensing by cGAS
Nature (2013)
-
Replication through an abasic DNA lesion: structural basis for adenine selectivity
The EMBO Journal (2010)
-
tRNA nucleotidyltransferases: ancient catalysts with an unusual mechanism of polymerization
Cellular and Molecular Life Sciences (2010)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.