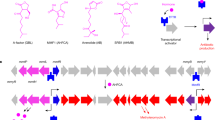
Abstract
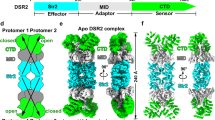
Many proteobacteria are able to monitor their population densities through the release of pheromones known as N-acylhomoserine lactones. At high population densities, these pheromones elicit diverse responses that include bioluminescence, biofilm formation, production of antimicrobials, DNA exchange, pathogenesis and symbiosis1. Many of these regulatory systems require a pheromone-dependent transcription factor similar to the LuxR protein of Vibrio fischeri. Here we present the structure of a LuxR-type protein. TraR of Agrobacterium tumefaciens was solved at 1.66 Å as a complex with the pheromone N-3-oxooctanoyl-l-homoserine lactone (OOHL) and its TraR DNA-binding site. The amino-terminal domain of TraR is an α/β/α sandwich that binds OOHL, whereas the carboxy-terminal domain contains a helix–turn–helix DNA-binding motif. The TraR dimer displays a two-fold symmetry axis in each domain; however, these two axes of symmetry are at an approximately 90° angle, resulting in a pronounced overall asymmetry of the complex. The pheromone lies fully embedded within the protein with virtually no solvent contact, and makes numerous hydrophobic contacts with the protein as well as four hydrogen bonds: three direct and one water-mediated.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
20 July 2011
The name of author Katherine M. Pappas has been corrected in the HTML as described in the accompanying Corrigendum.
References
Whitehead, N. A., Barnard, A. M. L., Slater, H., Simpson, N. J. L. & Salmond, G. P. C. Quorum-sensing in Gram-negative bacteria. FEMS Microbiol. Rev. 25, 365–404 (2001)
Fuqua, C., Parsek, M. R. & Greenberg, E. P. Regulation of gene expression by cell-to-cell communication: acyl-homoserine lactone quorum sensing. Annu. Rev. Genet. 35, 439–468 (2001)
Hanzelka, B. L. & Greenberg, E. P. Evidence that the N-terminal region of the Vibrio fischeri LuxR protein constitutes an autoinducer-binding domain. J. Bacteriol. 177, 815–817 (1995)
Choi, S. H. & Greenberg, E. P. Genetic evidence for multimerization of LuxR, the transcriptional regulator of Vibrio fischeri luminescence. Mol. Mar. Biol. Biotechnol. 1, 408–413 (1992)
Choi, S. H. & Greenberg, E. P. The C-terminal region of the Vibrio fischeri LuxR protein contains an inducer-independent lux gene-activating domain. Proc. Natl Acad. Sci. USA 88, 1115–1119 (1991)
Stevens, A. M., Dolan, K. M. & Greenberg, E. P. Synergistic binding of the Vibrio fischeri LuxR transcriptional activator domain and RNA polymerase to the lux promoter region. Proc. Natl Acad. Sci. USA 91, 12619–12623 (1994)
Fuqua, W. C. & Winans, S. C. A LuxR-LuxI type regulatory system activates Agrobacterium Ti plasmid conjugal transfer in the presence of a plant tumour metabolite. J. Bacteriol. 176, 2796–2806 (1994)
Egland, K. A. & Greenberg, E. P. Conversion of the Vibrio fischeri transcriptional activator, LuxR, to a repressor. J. Bacteriol. 182, 805–811 (2000)
Luo, Z. Q. & Farrand, S. K. Signal-dependent DNA binding and functional domains of the quorum-sensing activator TraR as identified by repressor activity. Proc. Natl Acad. Sci. USA 96, 9009–9014 (1999)
Chai, Y., Zhu, J. & Winans, S. C. A defective TraR-like protein of Agrobacterium tumefaciens forms heterodimers with TraR in vitro, thereby blocking TraR-mediated quorum sensing. Mol. Microbiol. 40, 414–421 (2001)
Zhu, J. & Winans, S. C. Autoinducer binding by the quorum-sensing regulator TraR increases affinity for target promoters in vitro and decreases traR turnover rates in whole cells. Proc. Natl Acad. Sci USA 96, 4832–4837 (1999)
Zhu, J. & Winans, S. C. The quorum-sensing regulator TraR of Agrobacterium tumefaciens requires autoinducer for protein folding, dimerization, and protease resistance. Proc. Natl Acad. Sci. USA 98, 1507–1512 (2001)
Qin, Y. et al. Quorum-sensing signal binding results in dimerization of TraR and its release from membranes into the cytoplasm. EMBO J. 19, 5212–5221 (2000)
Sandler, B. H., Nikonova, L., Leal, W. S. & Clardy, J. Sexual attraction in the silkworm moth: structure of the pheromone-binding-protein-bombykol complex. Chem. Biol. 7, 143–151 (2000)
Welch, M. et al. N-acyl homoserine lactone binding to the CarR receptor determines quorum-sensing specificity in Erwinia. EMBO J. 19, 631–641 (2000)
von Bodman, S. B., Majerczak, D. R. & Coplin, D. L. A negative regulator mediates quorum-sensing control of exopolysaccharide production in Pantoea stewartii subsp. Proc. Natl Acad. Sci. USA 95, 7687–7692 (1998)
Nelson, H. C. Structure and function of DNA-binding proteins. Curr. Opin. Genet. Dev. 5, 180–189 (1995)
Baikalov, I. et al. NarL dimerization? Suggestive evidence from a new crystal form. Biochemistry. 37, 3665–3676 (1998)
Luo, Z.-Q., Qin, Y. & Farrand, S. K. The antiactivator TraM interferes with the autoinducer-dependent binding of TraR to DNA by interacting with the C-terminal region of the quorum-sensing activator. J. Biol. Chem. 275, 7713–7722 (2000)
Egland, K. A. & Greenberg, E. P. Quorum sensing in Vibrio fischeri: analysis of the LuxR DNA binding region by alanine-scanning mutagenesis. J. Bacteriol. 183, 382–386 (2001)
Tempe, J., Petit, A., Holsters, M., Van Montagu, M. & Schell, J. Theromsensitive step associated with transfer of the Ti plasmid during conjugation: possible relation to transformation in crown gall. Proc. Natl Acad. Sci. USA 74, 2848–2849 (1977)
Joachimiak, A. & Sigler, P. B. Crystallization of protein-DNA complexes. Methods Enzymol. 208, 82–99 (1991)
Walsh, M. A., Dementieva, I., Evans, G., Sanishvili, R. & Joachimiak, A. Taking MAD to the extreme: ultrafast protein structure determination. Acta Crystallogr. D 55, 1168–1173 (1999)
Walsh, M. A., Evans, G., Sanishvili, R., Dementieva, I. & Joachimiak, A. MAD data collection—current trends. Acta Crystallogr. D 55, 1726–1732 (1999)
Otwinowski, Z. & Minor, W. Processing of x-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Brünger, A. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Perrakis, A., Morris, R. & Lamzin, V. S. Automated protein model building combined with iterative structure refinement. Nature Struct. Biol. 6, 458–463 (1999)
QUANTA, Molecular Simulations Inc, San Diego. (2000).
Acknowledgements
This work was supported by Monsanto Company, the US Department of Energy, Office of Biological and Environmental Research, and a National Research Service Award to S.C.W.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Zhang, Rg., Pappas, K., Brace, J. et al. Structure of a bacterial quorum-sensing transcription factor complexed with pheromone and DNA. Nature 417, 971–974 (2002). https://doi.org/10.1038/nature00833
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature00833
This article is cited by
-
Quorum sensing-based interactions among drugs, microbes, and diseases
Science China Life Sciences (2023)
-
Rapid evolution of bacterial mutualism in the plant rhizosphere
Nature Communications (2021)
-
Assessment of shikonin and acetyl-shikonin for mitigating quorum sensing potential of C. violaceum
Plant Growth Regulation (2021)
-
Evolution of LuxR solos in bacterial communication: receptors and signals
Biotechnology Letters (2020)
-
Biofilm Formation in Different Salmonella Serotypes Isolated from Poultry
Current Microbiology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.