Abstract
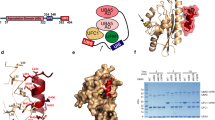
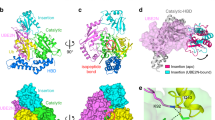
The activation of ubiquitin and related protein modifiers1,2 is catalysed by members of the E1 enzyme family that use ATP for the covalent self-attachment of the modifiers to a conserved cysteine. The Escherichia coli proteins MoeB and MoaD are involved in molybdenum cofactor (Moco) biosynthesis, an evolutionarily conserved pathway3,4. The MoeB- and E1-catalysed reactions are mechanistically similar, and despite a lack of sequence similarity, MoaD and ubiquitin display the same fold including a conserved carboxy-terminal Gly-Gly motif5. Similar to the E1 enzymes, MoeB activates the C terminus of MoaD to form an acyl-adenylate. Subsequently, a sulphurtransferase converts the MoaD acyl-adenylate to a thiocarboxylate that acts as the sulphur donor during Moco biosynthesis6,7. These findings suggest that ubiquitin and E1 are derived from two ancestral genes closely related to moaD and moeB3,5. Here we present the crystal structures of the MoeB–MoaD complex in its apo, ATP-bound, and MoaD-adenylate forms, and highlight the functional similarities between the MoeB– and E1–substrate complexes. These structures provide a molecular framework for understanding the activation of ubiquitin, Rub, SUMO and the sulphur incorporation step during Moco and thiamine biosynthesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hochstrasser, M. All in the ubiquitin family. Science 289, 563–564 (2000).
Hochstrasser, M. Evolution and function of ubiquitin-like protein-conjugation systems. Nature Cell Biol. 2, E153–E157 (2000).
Rajagopalan, K. V. in Escherichia coli and Salmonella typhimurium: Cellular and Molecular Biology (ed. Neidhardt, F. C.) 674–679 (ASM Press, Washington DC, 1996).
Rajagopalan, K. V. Biosynthesis and processing of the molybdenum cofactors. Biochem. Soc. Trans. 25, 757–761 (1997).
Rudolph, M. J., Wuebbens, M. M., Rajagopalan, K. V. & Schindelin, H. Crystal structure of molybdopterin synthase and its evolutionary relationship to ubiquitin activation. Nature Struct. Biol. 8, 42–46 (2001).
Leimkühler, S., Wuebbens, M. M. & Rajagopalan, K. V. Characterization of Escherichia coli MoeB and its involvement in the activation of MPT synthase for the biosynthesis of the molybdenum cofactor. J. Biol. Chem. 276, 34695–34701 (2001).
Pitterle, D. M., Johnson, J. L. & Rajagopalan, K. V. In vitro synthesis of molybdopterin from precursor Z using purified converting factor. J. Biol. Chem. 268, 13506–13509 (1993).
Walker, J. E., Saraste, M., Runswick, M. J. & Gay, N. J. Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J. 1, 945–951 (1982).
Burch, T. J. & Haas, A. L. Site-directed mutagenesis of ubiquitin. Differential roles for arginine in the interaction with ubiquitin-activating enzyme. Biochemistry 33, 7300–7308 (1994).
Arnez, J. G., Dock-Bregeon, A. C. & Moras, D. Glycyl-tRNA synthetase uses a negatively charged pit for specific recognition and activation of glycine. J. Mol. Biol. 286, 1449–1459 (1999).
Hatfield, P. M. & Vierstra, R. D. Multiple forms of ubiquitin-activating enzyme E1 from wheat. J. Biol. Chem. 267, 14799–14803 (1992).
Lauhon, C. T. & Kambampati, R. The iscS gene in Escherichia coli is required for the biosynthesis of 4-thiouridine, thiamin, and NAD. J. Biol. Chem. 275, 20096–20103 (2000).
Palenchar, P. M., Buck, C. J., Cheng, H., Larson, T. J. & Mueller, E. G. Evidence that ThiI, an enzyme shared between thiamin and 4-thiouridine biosynthesis, may be a sulfurtransferase that proceeds through a persulfide intermediate. J. Biol. Chem. 275, 8283–8286 (2000).
Xi, J., Ge, Y., Kinsland, C., McLafferty, F. W. & Begley, T. P. Biosynthesis of the thiazole moiety of thiamin in Escherichia coli: identification of an acyldisulfide-linked protein-protein conjugate that is functionally analogous to the ubiquitin/E1 complex. Proc. Natl Acad. Sci. USA 98, 8513–8518 (2001).
Leimkühler, S. & Rajagopalan, K. V. An Escherichia coli NifS-like sulfurtransferase is required for the transfer of cysteine sulfur in the in vitro synthesis of molybdopterin from precursor Z. J. Biol. Chem. 276, 22024–22031 (2001).
Otwinowski, Z. & Minor, W. in Methods in Enzymology: Macromolecular Crystallography (eds Carter, C. W. & Sweet, R. M. ) 307–326 (Academic, San Diego, 1997).
Sheldrick, G. M. & Schneider, T. R. in Methods in Enzymology: Macromolecular Crystallography (eds Carter, C. W. & Sweet, R. M.) 319–343 (Academic, San Diego, 1997).
DeLaFortelle, E. & Bricogne, G. in Methods in Enzymology: Macromolecular Crystallography (eds Carter, C. W. & Sweet, R. M.) 472–494 (Academic, San Diego, 1997).
Abrahams, J. P. & Leslie, A. G. W. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D 52, 30–42 (1996).
Perrakis, A., Morris, R. & Lamzin, V. S. Automated protein model building combined with iterative structure refinement. Nature Struct. Biol. 6, 458–463 (1999).
Bailey, S. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Murshudov, G., Vagin, A. & Dodson, E. Refinement of macromolecular structures by the maximum likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Johnson, M. E. & Rajagopalan, K. V. In vitro system for molybdopterin biosynthesis. J. Bacteriol. 169, 110–116 (1987).
Laskowski, R. A., Moss, D. S. & Thornton, J. M. Main-chain bond lengths and bond angles in protein structures. J. Mol. Biol. 231, 1049–1067 (1993).
Kraulis, P. J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Barton, G. J. ALSCRIPT: a tool to format multiple sequence alignments. Protein Eng. 6, 37–40 (1993).
Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Acknowledgements
We thank M. J. Rudolph for initial help with crystallization and data collection, J. Daniels for technical assistance, and D. Schneider for support at beamline X26C. This work was supported by National Institutes of Health (NIH) grants to H.S. and K.V.R.. The National Synchrotron Light Source in Brookhaven is supported by DOE and NIH, and beamline X26C is supported in part by the State University of New York at Stony Brook and its Research Foundation.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Lake, M., Wuebbens, M., Rajagopalan, K. et al. Mechanism of ubiquitin activation revealed by the structure of a bacterial MoeB–MoaD complex. Nature 414, 325–329 (2001). https://doi.org/10.1038/35104586
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35104586
This article is cited by
-
Novel causative variants of VEXAS in UBA1 detected through whole genome transcriptome sequencing in a large cohort of hematological malignancies
Leukemia (2023)
-
Structures of UBA6 explain its dual specificity for ubiquitin and FAT10
Nature Communications (2022)
-
Bioinformatics analysis of enzymes involved in cysteine biosynthesis: first evidence for the formation of cysteine synthase complex in cyanobacteria
3 Biotech (2021)
-
Structural and Functional Characterisation of the Domains of Ubiquitin-Activating Enzyme (E1) of Saccharomyces cerevisiae
Cell Biochemistry and Biophysics (2020)
-
Structural insights into E1 recognition and the ubiquitin-conjugating activity of the E2 enzyme Cdc34
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.