Abstract
Aquifex aeolicus was one of the earliest diverging, and is one of the most thermophilic, bacteria known. It can grow on hydrogen, oxygen, carbon dioxide, and mineral salts. The complex metabolic machinery needed for A. aeolicus to function as a chemolithoautotroph (an organism which uses an inorganic carbon source for biosynthesis and an inorganic chemical energy source) is encoded within a genome that is only one-third the size of the E. coli genome. Metabolic flexibility seems to be reduced as a result of the limited genome size. The use of oxygen (albeit at very low concentrations) as an electron acceptor is allowed by the presence of a complex respiratory apparatus. Although this organism grows at 95 °C, the extreme thermal limit of the Bacteria, only a few specific indications of thermophily are apparent from the genome. Here we describe the complete genome sequence of 1,551,335 base pairs of this evolutionarily and physiologically interesting organism.
Similar content being viewed by others
Main
Complete genome sequences have been determined for a number of organisms, including Archaea1, Bacteria2,3,4,5,6,7, and Eukarya8. Here we present and explore the genome sequence of Aquifex aeolicus. With growth-temperature maxima near 95 °C, Aquifex pyrophilus and A. aeolicus are the most thermophilic bacteria known. Although isolated and described only recently9, these species are related to filamentous bacteria first observed at the turn of the century, growing at 89 °C in the outflow of hot springs in Yellowstone National Park10,11. The observation of these macroscopic assemblages would later be instrumental in the drive to culture hyperthermophilic organisms12.
The Aquificaceae represent the most deeply branching family within the bacterial domain on the basis of phylogenetic analysis of 16S ribosomal RNA sequences13,14, although analyses of individual protein sequences vary in their placement of Aquifex relative to other groups15,16,17,18. The genera in this group, Aquifex and Hydrogenobacter, are thermophilic, hydrogen-oxidizing, microaerophilic, obligate chemolithoautotrophs9,19,20,21. A. aeolicus (isolated by R.H. and K. O. Stetter) was cultured at 85 °C under an H2/CO2/O2 (79.5:19.5:1.0) atmosphere in a medium containing only inorganic components. A. aeolicus does not grow on a number of organic substrates, including sugars, amino acids, yeast extract or meat extract. Unlike its close relative A. pyrophilus , A. aeolicus has not been shown to grow anaerobically with nitrate as an electron acceptor in the laboratory.
From study of the physiology of the organism, several predictions can be made. As an autotroph, A. aeolicus must have genes encoding proteins for one or more modes of carbon fixation and a complete set of biosynthetic genes. As autotrophy is a feature that is distributed throughout the Archaea and Bacteria, most of the associated genes are expected to be of ancient origin and clearly related to those characterized elsewhere. The obligate autotrophy suggests a biosynthetic rather than a degradative character. Oxygen respiration implies the presence of corresponding utilization and tolerance genes. The early divergence of the Aquificaceae inferred from ribosomal RNA sequences leads to several questions. Are the machineries for oxygen usage and tolerance homologous to those found in mitochondria and well studied organisms such as Escherichia coli, or were they invented separately? If there was far less oxygen when the lineage originated, is there evidence for use of alternative oxidants?
Genome
General features of the A. aeolicus genome are listed in Box 1. We classified 1,512 open-reading frames (ORFs) into one of three categories, namely, identified Table 1 (Note: Please refer to PDF version — Size: 162K), hypothetical, or unknown. Identified ORFs were further classified into one of 57 cellular role categories adapted from Riley22 (Table 1 — PDF Size: 162k). The relatively high G+ C content of the two 16S-23S-5S rRNA operons (65%) is characteristic of thermophilic bacterial rRNAs23. The genome is densely packed: most genes are apparently expressed in polycistronic operons and many convergently transcribed genes overlap slightly. Nonetheless, many genes that are functionally grouped within operons in other organisms, such as the tryptophan or histidine biosynthesis pathways, are found dispersed throughout the A. aeolicus genome or appear in novel operons. Even when they encode subunits of the same enzyme, the genes are often separated on the chromosome (for example, glt B and glt D, the genes encoding the large and small subunits of glutamate synthase). Operon organization of genes for the biosynthesis of amino acids is found in both Archaea and Bacteria but it is not universal in either group. A. aeolicus is extreme in that no two amino acid biosynthetic genes are found in the same operon. In contrast, genes required for electron transport, hydrogenase subunits, transport systems, ribosomal subunits, and flagella are often in functionally related operons in A. aeolicus (Figure 1) (Note: Please refer to PDF version — Size: 155K). No introns or inteins (protein splicing elements) were detected in the genome.
A single extrachromosomal element (ECE) was identified during sequencing. Sequence redundancy for the total project was calculated to be 4.83. The ECE, however, is significantly over-represented relative to the chromosome; when calculated independently for the final assemblies, redundancies are 4.73 and 8.76 for the chromosome and for the ECE, respectively. The ECE therefore appears to be present at roughly twice the copy number of the chromosome. Although no ORFs on the ECE can be assigned a function with confidence, except for a transposase, two of the predicted proteins show similarity to hypothetical proteins in the Methanococcus jannaschii genome1. One ORF on the ECE is also present in two identical copies on the A. aeolicus chromosome, providing evidence of genetic exchange between the chromosome and the ECE.
Reductive tricarboxylic acid cycle
As an autotroph, A. aeolicus obtains all necessary carbon by fixing CO2 from the environment. An assay for activity of the reductive tricarboxylic acid (TCA) cycle in A. pyrophilus cell extracts showed in vitro activities for each proposed reaction24. The reductive (reverse) TCA cycle fixes two molecules of CO2 to form acetyl-coenzyme A (acetyl-CoA) and other biosynthetic intermediates25. The A. aeolicus genome contains genes encoding malate dehydrogenase, fumarate hydratase, fumarate reductase, succinate-CoA ligase, ferredoxin oxidoreductase, isocitrate dehydrogenase, aconitase and citrate synthase, which together could constitute the TCA pathway. There is no biochemical evidence for alternative carbon-fixation pathways in A. pyrophilus24,25 nor is there sequence evidence for such pathways in A. aeolicus.
The TCA cycle is vital as it provides the substrates of many biosynthetic pathways. (It is beyond the scope of this report to detail these biosynthetic pathways, but they seem to be typically bacterial, and candidate genes for all or most of the enzymes have been identified in A. aeolicus.) The central role of the TCA cycle is emphasized by duplication of many of its constituent genes in A.aeolicus. Two genes encode proteins that are similar to malate dehydrogenase (in addition to a lactate dehydrogenase). The fumarate hydratase is split into amino- and carboxy-terminal subunits, as is the case in M. jannaschii1. Unlinked genes encoding two iron–sulphur proteins of fumarate reductase (alternatively succinate dehydrogenase) accompany a single flavoprotein subunit. Two sets of genes resembling succinate-CoA ligase (both the α- and β-subunits) are present. A. aeolicus has two putative operons encoding four-subunit (α, β, γ, δ) 2-acid ferredoxin oxidoreductases; members of this family catalyze reversible carboxylation/decarboxylation of pyruvate, 2-isoketovalerate, or 2-oxoglutarate with varying specificity26. These duplicated genes may encode paralogous proteins with unique substrate specificity, as opposed to redundant functions. For example, a paralogue of succinate-CoA ligase may activate citrate with coenzyme A to form citryl-CoA, which citrate synthase can cleave to produce oxaloacetate and acetyl-CoA.
Gluconeogenesis through the Embden–Meyerhof–Parnas pathway
Growing autotrophically, A. aeolicus must synthesize pentose and hexose monosaccharides from products of the reductive TCA cycle. Pyruvate produced by pyruvate ferredoxin oxidoreductase or by pyruvate carboxylase (oxaloacetate decarboxylase)24 may enter the Embden–Meyerhof–Parnas pathway of glycolysis and gluconeogenesis. Genes encoding fructose-1,6-bisphosphatase, an essential gluconeogenic enzyme in E. coli, have not been identified in the genomes of the autotrophs A. aeolicus or M. jannaschii1, suggesting that an unidentified pathway may exist. The A. aeolicus genome also encodes enzymes of the pentose-phosphate pathway and enzymes for glycogen synthesis and catabolism. We found neither (phospho) gluconate dehydrase nor 2-keto-3-deoxy-(6-phospho)gluconate aldolase of the Entner–Doudoroff pathway.
Respiration
Aquifex species are able to grow by using oxygen concentrations as low as 7.5 p.p.m. (R.H. and K. O. Stetter, unpublished observations). The enzymes for oxygen respiration are similar to those of other bacteria: ubiquinol cytochrome c oxidoreductase (bc 1 complex), cytochrome c (three different genes) and cytochrome c oxidase (with two different subunit I genes and two different subunit II genes). The alternative system, with cytochrome bd ubiquinol oxidase, is also present. Clearly, the Aquifex lineage did not independently invent oxygen respiration. This leaves at least three possibilities: consistent with the ability of Aquifex to use very low levels of oxygen, the oxygen-respiration system was highly developed when oxygen had only a small fraction of its present concentration before the advent of oxygenic photosynthesis; contrary to what is implied by the 16S phylogeny, the lineage including Aquifex originated after the rise in atmospheric oxygen; or oxygen respiration developed once, and was then laterally transferred among bacterial lineages and acquired by Aquifex.
Many other oxidoreductases are present in addition to those obviously involved in oxygen respiration. The physiological role of most of these oxidoreductases is unknown or ambiguous, but two deserve comment. There is a putative nitrate reductase in the genome, although A. aeolicus has not been observed to perform NO3− respiration, unlike the closely related A. pyrophilus. The nitrate reductase gene is adjacent to a nitrate transporter, and may be involved in nitrogen assimilation rather than respiration. It is also possible that A. aeolicus has a latent ability to respire with nitrate but that the conditions required have not been found. Two gene sequences show strong similarities to Rieske proteins, even though the rest of the ubiquinol cytochrome c oxidoreductase subunits appear only once in the genome. One of these Rieske protein genes is adjacent to a sulphide dehydrogenase subunit, suggesting a role in sulphur respiration.
Oxidative stress
A. aeolicus grows optimally under microaerophilic conditions and consequently possesses various protective enzymes to counter reactive oxygen species, particularly superoxide and peroxide. The genome contains three genes encoding superoxide dismutases, two of the copper/zinc family and one of the iron/manganese family. The latter has also been noted in A. pyrophilus 27. One of the copper/zinc superoxide dismutase genes is located in a large gene cluster encoding formate dehydrogenase.
No catalase genes were identified. There are several genes in the genome that might encode proteins that catalyze the detoxification of H2O2, including cytochrome c peroxidase, thiol peroxidase, and two alkyl hydroperoxide reductase genes. All of these enzymes require an exogenous reductant and therefore do not evolve O2. However, treatment of A. pyrophilus9 or A. aeolicus biomass with H2O2 results in the rapid evolution of gas bubbles. This catalase activity may result from a novel enzyme that cannot yet be identified by sequence similarity.
Motility
Like A. pyrophilus 9, A. aeolicus ismotile and possesses monopolar polytrichous flagella. More than25 genes encoding proteins involved in flagellar structure andbiosynthesis have been identified in A. aeolicus (Box 1). However, no homologues of the bacterial chemotaxis system were identified. In enteric bacteria, membrane-bound receptors bind chemoattractants and repellents and modulate the activity of the histidine kinase CheA28. Phosphoryl groups from CheA are transferred to CheY, which then binds to the flagellar switch, altering the direction of flagellar rotation. Homologous chemotaxis systems are present in the archaea Halobacterium salinarum29 and Pyrococcus sp. OT3 (H. Sizuya, personal communication), although the bacterial and archaeal flagellar apparatuses are not homologous30. The M. jannaschii genome also lacks homologues of known genes required for chemotaxis. Thus, either motility in A. aeolicus and M. jannaschii is undirected or input for controlling taxis is mediated through another, unidentified system. The most studied chemotaxis systems respond to sugars and amino acids, although responses to other inputs (for example, metals, redox potential, and light) may also occur. In contrast to all the organisms known to possess the classical chemotactic signal-transduction pathways, both A. aeolicus and M. jannaschii are obligate chemoautotrophs. Chemoautotrophs may respond to a different set of factors, such as concentrations of dissolved gas (CO2, H2 or O2) or another critical parameter such as temperature.
In E. coli, the flagellar switch is essential for flagellar structure and function and coupling of chemotaxis signals. But the A. aeolicus genome encodes homologues of only two of the three E. coli proteins that make up the switch, FliG and FliN. Biochemical31 and genetic32 studies implicate the missing FliM protein as the receptor for phosphorylated CheY, the switch signal. The absence of both FliM and CheY in A. aeolicus supports the identification of FliM as the receptor for phosphorylated CheY in E. coli. This result also argues against a direct role for FliM in torque generation.
DNA replication and repair
The A. aeolicus primary replicative DNA polymerase, corresponding to the DNA polymerase III holoenzyme in E. coli, probably consists of a core structure containing α- and ε-subunits, a γ-τ-subunit and an additional member of the γ-τ/δ′-family. A gene encoding a protein homologous to the β-sliding clamp was also found. This minimalistic complex lacks homologous θ-, δ-, χ- and ψ-subunits, as does the Mycoplasma genitalium holoenzyme3. Translation of the 54K (relative molecular mass) γ-τ-ATPase subunit may proceed without a programmed frameshift to produce a protein similar to the N-terminal region of the E. coli γ-subunit. DNA polymerase I is present as separate Klenow fragment and 5′ → 3′ exonuclease subunits, encoded by two non-adjacent ORFs. Although the repair polymerase, DNA polymerase II, has not been found in A. aeolicus, one ORF (Aq1422) encodes a protein similar to the eukaryotic DNA repair polymerase-β. A member of the same family has been identified in Thermus aquaticus33 and Bacillus subtilis.
Transcriptional and translational apparatuses
The transcriptional apparatus of A. aeolicus is similar to that of E. coli and lacks any components specific to the Eukarya or Archaea (Fig. 2). In addition to the core RNA polymerase α-, β-, and β′-subunits, four σ-factors which determine promoter specificity are present (Table 1) ( PDF Size: 162k). Several different families of bacterial transcriptional regulators were also identified, including two-component systems. All of the ribosomal proteins and elongation factors common to other bacteria are present, indicating that all bacteria-specific ribosomal proteins were present in the common ancestor of Aquifex and other bacteria. Also present are the four sel genes required for the cotranslational incorporation of selenocysteine. These latter genes are clustered in a 15-kilobase-pair segment that also encodes the biosynthetic and structural proteins for formate dehydrogenase, the only selenocysteine-containing protein identified. The gene that encodes selenocysteine transfer RNA, selC, is apparently cotranscribed with the genes encoding the formate dehydrogenase structural proteins.
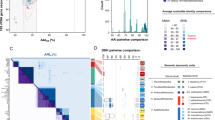
Predicted A. aeolicus proteins representing each category were independently compared to sets of all potential polypeptides (≥100 amino acids) from the two genomes using FASTA44. If the top scoring alignment covered ≥80% of the length of the A. aeolicus protein, the score was plotted. There were more positives found in the E. coli genome in nearly every category. Hypothetical proteins (those identified by database match but of unknown function) are very similarly represented by M. jannaschii and E. coli. There are a small number of very highly conserved hypotheticals that are shared between A. aeolicus and M.jannaschii . Generally, biosynthetic categories show less discrimination than information-processing categories, which are clearly more E. coli -like. The variation in the apparent rates of evolution in different categories suggests that different phylogenies may be inferred depending on the sequence analysed. Within each graph, correspondence to E. coli is shown in white and M. jannaschii is shown in black. Avg id, average identity; count, number of proteins analysed.
A. aeolicus lacks glutaminyl-tRNA and asparaginyl-tRNA synthetases. The genes required for transamidation of glutamyl-tRNAGln are present34. Charging of asparaginyl-tRNA is likely to proceed through the analogous reaction, as shown in halobacteria35, although the genes(s) for that transamidase are unknown. The canonical methionyl- and leucyl-tRNA synthetases have only been seen previously as single polypeptide enzymes; however, in A. aeolicus the homologues appear fragmented into two subunits. In both cases, the genes that encode the N- and C-terminal portions are widely separated on the chromosome. No complete three-dimensional structural data are available for either methionyl- or leucyl-aminoacyl tRNA synthetases, but the subunit organization in the A. aeolicus aminoacyl-tRNA synthetases may reflect domain organization in the homologous proteins.
Thermophily
The A. aeolicus genome is the second completely sequenced genome of a hyperthermophile. By comparing the A. aeolicus and M. jannaschii genomes and contrasting them with the complete genomes of mesophiles, we can discover whether there are aspects of the genome or the encoded information that are diagnostic of hyperthermophiles. The G+ C content of the stable RNAs is clearly indicative of the high growth temperature of the organism. This property can be used to identify stable RNAs against the relatively low G+ C background of the A. aeolicus genome. The gene encoding tmRNA (or 10Sa RNA)36, an RNA involved in tagging polypeptides translated from incomplete messenger RNAs for degradation, was located in this way.
Two genes for reverse gyrase are present in the genome. This is the only protein known to be present only in thermophiles. Other proteins, currently described as hypotheticals, may be diagnostic of hyperthermophiles but the data sets are not yet large enough to decide this with confidence.
Although features of stabilization may not be apparent in any given protein37, a large enough data set may reveal general trends in amino-acid usage that are informative. Particularly important in this regard is inclusion of multiple genomes of hyperthermophiles so as not to allow the idiosyncracies of a single organism to bias the conclusions. As shown in Table 2, comparison of the amino-acid composition encoded by six genomes shows that use of individual amino acids can vary significantly from genome to genome. The data suggest trends that may be correlated with the thermostability of the encoded proteins. One apparent trend is that the hyperthemophile genomes encode higher levels of charged amino acids on average than mesophile genomes38, primarily at the expense of uncharged polar residues. Glutamine in particular seems to be significantly discriminated against in the hyperthermophiles. Although this observation might be rationalized on the basisof an increased rate of deamidation of this residue at highertemperatures, aspargine does not appear subject to similar discrimination.
Phylogeny
The placement of the Aquifex lineage as one of the earliest divergences in the eubacterial tree13,14 is interesting because of the insights it could provide into the ancestral eubacterial phenotype, including the hypothesized thermophilic nature of the first bacteria. Protein-based phylogenies often do not support the original rRNA-based placement15,16,18. Thus, the availability of some 1,500 genes from an Aquifex species would seem to offer a definitive resolution of the phylogeny. However, our analyses of ribosomal proteins, aminoacyl-tRNA synthetases, and other proteins do not do so, showing no consistent picture of the organism's phylogeny. We cannot make a more complete analysis and discussion here, but some observations can be made. These proteins do not yield a statistically significant placement of the Aquifex lineage or of other major eubacterial lineages. This situation partially reflects the inadequacy of some protein sequences as indicators of distant molecular genealogy because of their particular evolutionary dynamic, including the patterns and rates of amino-acid replacements. In some cases (such as the aminoacyl-tRNA synthetases for arginine, cysteine, histidine, proline and tyrosine), the analyses are further complicated by the presence of paralogous genes and/or apparent lateral gene transfers. It seems that a more extensive survey of genes and a better sampling of major eubacterial taxa will be required to confidently confirm or refute an early divergence of the Aquifex lineage.
Conclusions
Advances in sequencing techniques have allowed us to move beyond studies of single genes to studies of complete genomes only recently2. This rapid advance has created the opportunity to begin to characterize an organism with the full knowledge of the genome in hand. The complete genome summarized in this report represents our first view of A. aeolicus. The challenge now is to ask specific questions in ways which take advantage of the whole-genome data.
Beyond studies of any single organism in isolation, complete genomes allow comprehensive comparisons between organisms. For instance, comparisons of the similarity of genes can be made that reveal that genes in different categories vary in their relative conservation (Fig. 2). In addition, genome-wide trends are apparent. For example, why is there not more of a tendency to group functionally related genes (for example, biosynthetic pathways) into operons in A. aeolicus ? This was also seen in the genome sequence of the autotroph M. jannaschii1. Is this because the autotrophic lifestyle decreases the need for selective regulation? There also seem to be a few multifunctional, fused proteins in A.aeolicus and M. jannaschii. Although this seems unlikely to be related to autotrophy, it might be associated with extreme thermophily. The large number of diverse genome sequences that will become available in the coming years will allow more detailed correlation of global genomic properties with particular physiologies.
Methods
Sequencing strategy. The sequencing strategy used to assemble the complete genome was based on the whole genome random (or ‘shotgun’) approach, which has been successfully used for other genomes of similar size1,2,3,4. Shotgun sequencing projects are characterized by two phases: an initial completely random phase in which the bulk of the data is collected, followed by a closure phase where directed techniques are used to close gaps and complete the assembly. By pursuing a strategy where only 97% coverage was initially achieved, we were able to limit the number of sequences needed for the random phase to only 10,500 (ref. 39).
Sequences were generated from a small insert library constructed in λ ZAP II vectors40,41 (average insert length 2.9 kilobase pairs). Two different methods were used for sequencing: first, dye-primer M13-21 and M13 reverse primer ABI Prism CS+ ready reaction kits, analysed on 48-cm 4% polyacrylamide gels; and second, dye-terminator (ABI Prism FS+) reactions using two pBluescript-specific primers. These reactions were analysed on 36-cm 5% Long-Ranger gels.
The sequence fragments were assembled on an Apple Power Macintosh computer using Sequencher (Gene Codes, Ann Arbor, MI), an assembly and editing program. Assembly was typically performed in batches of roughly 200–400 sequences, and was followed by inspection and editing of the assemblies. All sequences in the set were compared with all others through this process. After assembly, the sequences comprised ∼750 contigs at the end of the random phase. Sequences were obtained from both ends of ∼200 randomly chosen clones from a fosmid library42,43. These sequences were then assembled with consensus sequences derived from the contigs of random-phase sequences using Sequencher. Gaps between contigs were closed by direct sequencing on fosmids not wholly contained within a contig. The fosmid library thus served a purpose analogous to that of the λ-scaffold in other projects1,2,3,4. The final eight gaps were closed by direct sequencing of polymerase chain reaction (PCR) products generated with the TaqPlus Long PCR System (Stratagene Cloning Systems, La Jolla, CA).
Consequences of reducing the number of sequences in the random phase are the large number of gaps that remain to be closed in the directed phase, and the reduction in overall coverage. To ensure that reduced coverage did not compromise accuracy, ∼200 oligonucleotide primers were synthesized to resequence regions of ambiguity identified by visual inspection of the entire assembly. 13,785 sequences, with an average edited read length of 557 base pairs, constitute the final assembly. On the basis of a relatively small number of errors identified during the annotation process, we estimate the error frequency to be <0.01%, comparable to other published genomic sequence estimates.
Gene (ORF + RNA) identification and functional assignment approaches. Coding regions of the A. aeolicus genome were analysed and assigned using primarily the programs BLASTP44 and FASTA45 to search against a non-redundant protein database. Many analyses were carried out within the context of MAGPIE46,47, an integrated computing environment for genome analysis. The results of these analyses are available for user interpretation, validation, and categorization. Additional ORFs were identified and start sites refined using the program CRITICA (J. H. Badger and G.J.O., unpublished program). Finally, all presumed ‘intergenic regions’ were examined with BLASTX for similarities to known protein sequences48. Transfer RNA genes were identified with the program tRNAscan-SE49.
References
Bult, C. et al. Complete genome sequence of the methanogenic archaeon, Methanococcus jannaschii . Science 273, 1058– 1073 (1996).
Fleischmann, R. D. et al. Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science 269, 496– 511 (1995).
Fraser, C. M. et al. The minimal gene complement of Mycoplasma genitalium. Science 270, 397–403 (1995).
Tomb, J.-F. et al. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature 388, 539– 547 (1997).
Himmelreich, R. et al . Complete sequence analysis of the genome of the bacterium Mycoplasma pneumoniae. Nucleic Acids Res. 24, 4420–4449 (1996).
Kaneko, T. et al. Sequence analysis of the genome of the unicellular cyanobacterium Synechocystis sp. strain PCC7803. II. Sequence determination of the entire genome and assignment of potential protein-coding regions. DNA Res. 3, 109–136 ( 1996).
Blattner, F. R. et al . The complete genome sequence of Escherichia coli K-12. Science 277, 1453– 1462 (1997).
Goffeau, A. et al. Life with 6000 genes. Science 274, 546 (1996).
Huber, R. et al. Aquifex pyrophilus gen. nov. sp. nov. represents a novel group of marine hyperthermophilic hydrogen oxidizing bacteria. Arch. Micrtobiol. 15, 340–351 (1992).
Reysenbach, L., Wickham, G. S. & Pace, N. R. Phylogenetic analysis of the hyperthermophilic pink filament community in Octopus Spring, Yellowstone National Park. Appl. Environ. Microbiol. 60, 2113– 2119 (1994).
Setchell, W. A. The upper temperature limits of life. Science 17, 934–937 (1903).
Brock, T. D. The road to Yellowstone — and beyond. Annu. Rev. Microbiol. 49, 1–28 (1995 ).
Burggraf, S., Olsen, G. J., Stetter, K. O. & Woese, C. R. Aphylogenetic analysis of Aquifex pyrophilus. Syst. Appl. Microbiol. 15, 353–356 ( 1992).
Pitulle, C. et al. Phylogenetic position of the genus Hydrogenobacter. Int. J. Syst. Bacteriol. 44, 620–626 (1994).
Baldauf, S. L., Palmer, J. D. & Doolittle, W. F. The root of the universal tree and the origin of eukaryotes based on elongation factor phylogeny. Proc. Natl Acad. Sci. USA 93, 7749–7754 ( 1996).
Klenk, H.-P., Palm, P. & Zillig, W. in Molecular Biology of the Archaea(eds Pfeifer, F., Palm, P. & Scleeifer, K. H.) 139–147 (Vch Pub, (1994).
Bocchetta, M. et al . Arrangement and nucleotide sequence of the gene (fus ) encoding elongation factor G (EF-G) from the hyperthermophilic bacterium Aquifex pyrophilus : phylogenetic depth of hyperthermophilic bacteria inferred from analysis of the EF-G/fus sequences. J. Mol. Evol. 41, 803–812 ( 1995).
Wetmur, J. G. et al . Cloning, sequencing, and expression of RecA proteins from three distantly related thermophilic eubacteria. J. Biol. Chem. 269, 25928–25935 ( 1994).
Kawasumi, T., Igarashi, Y., Kodama, T. & Minoda, Y. Hydrogenobacter thermophilus gen. nov., sp. nov. an extremely thermophilic, aerobic, hydrogen-oxidizing bacterium. Int. J. Syst. Bacteriol. 34, 5–10 (1984).
Kristjannson, J., Ingason, A. & Alfredsson, G. A. Isolation of thermophilic obligately autotrophic hydrogen-oxidizing bacteria, similar to Hydrogenobacter thermophilus, from Icelandic hotsprings. Arch. Microbiol. 140, 321–325 (1985).
Kryukov, V. R., Savel'eva, N. D. & Pusheva, M. A. Calderobacterium hydrogenophilum gen. nov., sp. nov. an extreme thermophilic bacterium and its hydrogenase activity. Microbiology (Engl. Trans. Mikrobiologiya) 52, 611–618 (1983).
Riley, M. Functions of the gene products of Escherichia coli. Microbiol. Rev. 57, 862–952 ( 1993).
Weisburg, W. G., Giovannoni, S. J. & Woese, C. R. The Deinococcus-Thermus phylum and the effect of rRNA composition on phylogenetic tree construction. Syst. Appl. Microbiol. 11, 128–134 ( 1989).
Beh, M., Strauss, G., Huber, R., Stetter, K. O. & Fuchs, G. Enzymes of the reductive citric acid cycle in the autotrophic eubacterium Aquifex pyrophilus and in the archaebacterium Thermoproteus neutrophilus. Arch. Microbiol. 160, 306–311 (1993).
Fuchs, G. in Autotrophic Bacteria(eds Schegel, H. G & Bowein, B.) 365– 382 (Springer, New York, (1987).
Mai, X. & Adams, M. W. Characterization of a fourth type of 2-keto acid-oxidizing enzyme from a hyperthermophilic archaeon: 2-ketoglutarate ferredoxin oxidoreductase from Thermococcus litoralis. J. Bacteriol. 178, 5890–5896 (1996).
Lim, J. H. et al. Cloning and expression of superoxide dismutase from Aquifex pyrophilus, a hyperthermophilic bacterium. FEBS Lett. 406, 142–146 (1997).
Bourret, R. B., Borkovich, K. A. & Simon, M. I. Signal transduction pathways involving protein phosphorylation in prokaryotes. Annu. Rev. Biochem. 60, 401–441 (1991).
Rudolph, J., Tolliday, N., Schmitt, C., Schuster, S. C. & Oesterhelt, D. Phosphorylation in halobacterial signal transduction. EMBO J. 14, 4249– 4257 (1995).
Jarrell, K. F., Bayley, D. P. & Kostyukova, A. S. The archaeal flagellum: a unique motility structure. J. Bacteriol. 178, 5057– 5064 (1996).
Welch, M., Oosawa, K., Aizawa, S. I. & Eisenbach, M. Effects of phosphorylation, Mg2+, and conformation of the chemotaxis protein CheY on its binding to the flagellar switch protein FliM. Biochemistry 33, 10470–10467 (1994).
Sockett, H., Yamaguchi, S., Kihara, M., Irikura, V. M. & Macnab, R. M. Molecular analysis of the flagellar switch protein FliM of Salmonella typhimurium. J. Bacteriol. 174, 793–806 ( 1992).
Motoshima, H. et al . Molecular cloning and nucleotide sequence of the aminopeptidase T gene of Thermus aquaticus YT-1 and its high-level expression in Escherichia coli. Agric. Biol. Chem. 54, 2385–2392 (1990).
Curnow, A. W. et al . Glu-tRNAGln amidotransferase: a novel heterotrimeric enzyme required for correct decoding of glutamine codons during translation. Proc. Natl Acad. Sci. USA 94, 11819– 11826 (1997).
Curnow, A. W., Ibba, M. & Söll, D. tRNA-dependent asparagine formation. Nature 382, 589–590 ( 1996).
Tu, G. F., Reid, G. E., Zhang, J. G., Moritz, R. L. & Simpson, R. J. C-terminal extension of truncated proteins in Escherichia coli with a 10Sa decapeptide. J. Biol. Chem. 270, 9322–9326 (1995).
Böhm, G. & Jaenicke, R. Relevance of sequence statistics for the properties of extremophilic proteins. Int. J. Pept. Protein Res. 43, 97–106 ( 1994).
Choi, I.-G. et al. Random sequence analysis of genomic DNA of a hyperthermophile: Aquifex pyrophilus. Extremophiles 1, 125–134 (1997).
Lander, E. S. & Waterman, M. S. Genomic mapping by fingerprinting random clones: a mathematical analysis. Genomics 2, 231–239 (1988).
Short, J. M., Fernandez, J. M., Sorge, J. A. & Huse, W. D. Lambda ZAP: a bacteriophage lambda expression vector with in vivo excision properties. Nucleic Acids Res. 16, 7583– 7600 (1988).
Alting-Mees, M. A. & Short, J. M. pBluescript II: gene mapping vectors. Nucleic Acids Res. 17, 9494 (1989).
Shizuya, H. et al. Cloning and stable maintenance of 300-kilobase-pair fragments of human DNA in Escherichia coli using an F-factor-based vector. Proc. Natl Acad. Sci. USA 89, 8794– 8797 (1992).
Kim, U.-J., Shizuya, H., de Jong, P. J., Birren, B. & Simon, M. I. Stable propagation of cosmid sized human DNA inserts in an F factor based vector. Nucleic Acids Res. 20, 1083–1085 ( 1992).
Altschul, S. F., Fish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Pearson, W. R. & Lipman, D. J. Improved tools for biological sequence comparison. Proc. Natl Acad. Sci. USA 85, 2444–2448 (1988).
Gaasterland, T. & Sensen, C. W. MAGPIE: automated genome interpretation. Trends Genet. 12, 76–78 (1996).
Gaasterland, T. & Sensen, C. W. Fully automated genome analysis that reflects user needs and preferences. A detailed introduction to the MAGPIE system architecture. Biochimie 78, 302–310 (1996).
Gish, W. & States, D. J. Identification of protein coding regions by database similarity search. Nature Genet. 3, 266–272 (1993).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 ( 1997).
Acknowledgements
This work was supported in part by Department of Energy Microbial Genome Program grants (to R.V.S., C. R. Woese and G.J.O.). We thank C. Woese for his cooperation in the analysis of the genome and interest in the project; K. Stetter for continuing interest; G. Frey, J. Holaska, S. Peralta, D. Hafenbrandl, S. Delk, T. Robinson, and J. Arnett for technical assistance; and D. Robertson, J. Stein, I.Sanyal, T. Richardson, G. Hauska, and K. Williams for discussions.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Deckert, G., Warren, P., Gaasterland, T. et al. The complete genome of the hyperthermophilic bacterium Aquifex aeolicus . Nature 392, 353–358 (1998). https://doi.org/10.1038/32831
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/32831
This article is cited by
-
Tectonically-driven oxidant production in the hot biosphere
Nature Communications (2022)
-
Genomic attributes of thermophilic and hyperthermophilic bacteria and archaea
World Journal of Microbiology and Biotechnology (2022)
-
Occurrence of randomly recombined functional 16S rRNA genes in Thermus thermophilus suggests genetic interoperability and promiscuity of bacterial 16S rRNAs
Scientific Reports (2019)
-
Structural characterization of HypX responsible for CO biosynthesis in the maturation of NiFe-hydrogenase
Communications Biology (2019)
-
Characterization of a mildly alkalophilic and thermostable recombinant Thermus thermophilus laccase with applications in decolourization of dyes
Biotechnology Letters (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.