Abstract
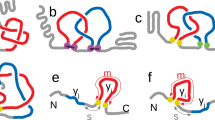
A β-strand is a particular type of extended sequence of amino acid residues, an element of secondary structure of proteins. β-sheets are an assembly of strands, often bringing together parts of the protein which are separated along the backbone. As such, β-sheets are an element of tertiary structure. Parallel βP) and antiparallel (βA) arrangements of strands in a sheet differ in the hydrogen bond pattern between strands, as shown schematically in Fig. 1, and in the type of chain connectivity they allow: short reverse turn connections for βA and longer crossover connections for βP (refs 1–3). Most present secondary structure prediction methods (for reviews refs 4–6) use a four-state distinction of secondary structure: α-helix, β-strand or extended, reverse turn, and ‘random coil’ (everything else). With a data base of 30–40 different protein structures, the conformational preferences for all amino acid residues in these four states seem to have converged7. However, the steadily increasing data base of structurally known proteins makes a refinement of the four-state description feasible. Although more refined classifications of conformational states based on finer subdivisions of (φ,Ψ)-space have been made8,9, we prefer making distinctions based on structural environment. Using a novel definition of β-sheet structure in terms of the tertiary structure juxtaposition of strands, we have analysed residue contacts in known β-sheets and report here secondary structure preferences for the 20 amino acids, separately for antiparallel and parallel arrangements of strands. The distinction between the two arrangements results in strikingly different and sharpened sets of preference parameters, including some of the largest values reported so far for any substructure. These results point the way towards a basic improvement of secondary structure predictions by further distinction of secondary structure elements according to tertiary structure environment. Beyond secondary structure prediction, the different preferences for βA and βP may aid in predicting the tertiary interaction between strands.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Levitt, M. & Chothia, C. Nature 261, 552–558 (1976).
Richardson, J. S. Nature 268, 495–500 (1977).
Sternberg, M. J. E. & Thornton, J. M. J. molec. Biol. 115, 1–17 (1977).
Chou, P. Y. & Fasman, F. D. A. Rev. Biochem. 47, 251–276 (1978).
Sternberg, M. J. E. & Thornton, J. M. Nature 271, 15–20 (1978).
Schulz, G. E. & Schirmer, R. H. Principles of Protein Structure (Springer, New York, 1979).
Robson, B. & Suzuki, E. J. molec. Biol. 107, 327–356 (1976).
Robson, B. & Pain, R. H. Biochem. J. 141, 869–882 (1974).
Burgess, A. W., Ponnuswamy, P. K. & Scheraga, H. A. Israel J. Chem. 12, 239–286 (1975).
Lifson, S. & Sander, C. J. molec. Biol. (submitted).
Feldman, R. J. AMSOM Atlas of Molecular Structures on Microfiche (U.S. NIH, 1976).
Levitt, M. Biochemistry 17, 4277–4285 (1978).
Chou, P. Y. & Fasman, G. D. Biochemistry 13, 222–245 (1974).
Gamier, J., Osguthorpe, D. J. & Robson, B. J. molec. Biol. 120, 97–120 (1978).
Levitt, M. & Greer, J. J. molec. Biol. 114, 181–239 (1977).
Chou, P. Y. & Fasman, G. D. J. molec. Biol. 115, 135–175 (1977).
Sander, C. & Schulz, G. E. J. molec. Evolut. (in the press).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Lifson, S., Sander, C. Antiparallel and parallel β-strands differ in amino acid residue preferences. Nature 282, 109–111 (1979). https://doi.org/10.1038/282109a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/282109a0
This article is cited by
-
Spider-silk inspired polymeric networks by harnessing the mechanical potential of β-sheets through network guided assembly
Nature Communications (2020)
-
Network analysis of dynamically important residues in protein structures mediating ligand-binding conformational changes
European Biophysics Journal (2019)
-
Exploring the potential of 3D Zernike descriptors and SVM for protein–protein interface prediction
BMC Bioinformatics (2018)
-
SVM-SulfoSite: A support vector machine based predictor for sulfenylation sites
Scientific Reports (2018)
-
Brain-specific HIV Nef identified in multiple patients with neurological disease
Journal of NeuroVirology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.