Abstract
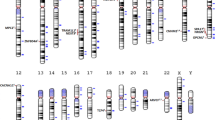
A genomewide linkage scan was carried out in eight clinical samples of informative schizophrenia families. After all quality control checks, the analysis of 707 European-ancestry families included 1615 affected and 1602 unaffected genotyped individuals, and the analysis of all 807 families included 1900 affected and 1839 unaffected individuals. Multipoint linkage analysis with correction for marker–marker linkage disequilibrium was carried out with 5861 single nucleotide polymorphisms (SNPs; Illumina version 4.0 linkage map). Suggestive evidence for linkage (European families) was observed on chromosomes 8p21, 8q24.1, 9q34 and 12q24.1 in nonparametric and/or parametric analyses. In a logistic regression allele-sharing analysis of linkage allowing for intersite heterogeneity, genomewide significant evidence for linkage was observed on chromosome 10p12. Significant heterogeneity was also observed on chromosome 22q11.1. Evidence for linkage across family sets and analyses was most consistent on chromosome 8p21, with a one-LOD support interval that does not include the candidate gene NRG1, suggesting that one or more other susceptibility loci might exist in the region. In this era of genomewide association and deep resequencing studies, consensus linkage regions deserve continued attention, given that linkage signals can be produced by many types of genomic variation, including any combination of multiple common or rare SNPs or copy number variants in a region.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Ng MYM, Levinson DF, Faraone SV, Suarez BK, DeLisi LE, Arinami T et al. Meta-analysis of 32 genomewide linkage studies of schizophrenia. Mol Psychiatry 2008; advance online publication 30 December 2008; doi:10.1038/mp.2008.135.
Manolio TA, Brooks LD, Collins FS . A HapMap harvest of insights into the genetics of common disease. J Clin Invest 2008; 118: 1590–1605.
Levinson DF, Mahtani MM, Nancarrow DJ, Brown DM, Kruglyak L, Kirby A et al. Genome scan of schizophrenia. Am J Psychiatry 1998; 155: 741–750.
Ewen KR, Bahlo M, Treloar SA, Levinson DF, Mowry B, Barlow JW et al. Identification and analysis of error types in high-throughput genotyping. Am J Hum Genet 2000; 67: 727–736.
Schwab SG, Hallmayer J, Albus M, Lerer B, Eckstein GN, Borrmann M et al. A genome-wide autosomal screen for schizophrenia susceptibility loci in 71 families with affected siblings: support for loci on chromosome 10p and 6. Mol Psychiatry 2000; 5: 638–649.
Williams NM, Rees MI, Holmans P, Norton N, Cardno AG, Jones LA et al. A two-stage genome scan for schizophrenia susceptibility genes in 196 affected sibling pairs. Hum Mol Genet 1999; 8: 1729–1739.
Cao Q, Martinez M, Zhang J, Sanders AR, Badner JA, Cravchik A et al. Suggestive evidence for a schizophrenia susceptibility locus on chromosome 6q and a confirmation in an independent series of pedigrees. Genomics 1997; 43: 1–8.
Campion D, d’Amato T, Bastard C, Laurent C, Guedj F, Jay M et al. Genetic study of dopamine D1, D2, and D4 receptors in schizophrenia. Psychiatry Res 1994; 51: 215–230.
Bonnet-Brilhault F, Laurent C, Campion D, Thibaut F, Lafargue C, Charbonnier F et al. No evidence for involvement of KCNN3 (hSKCa3) potassium channel gene in familial and isolated cases of schizophrenia. Eur J Hum Genet 1999; 7: 247–250.
Blouin JL, Dombroski BA, Nath SK, Lasseter VK, Wolyniec PS, Nestadt G et al. Schizophrenia susceptibility loci on chromosomes 13q32 and 8p21. Nat Genet 1998; 20: 70–73.
Kendler KS, FA ON, Burke J, Murphy B, Duke F, Straub RE et al. Irish study on high-density schizophrenia families: field methods and power to detect linkage. Am J Med Genet 1996; 67: 179–190.
Cloninger CR, Kaufmann CA, Faraone SV, Malaspina D, Svrakic DM, Harkavy-Friedman J et al. Genome-wide search for schizophrenia susceptibility loci: the NIMH Genetics Initiative and Millennium Consortium. Am J Med Genet 1998; 81: 275–281.
Schizophrenia Linkage Collaborative Group for Chromosomes 3, 6 and 8. Additional support for schizophrenia linkage on chromosomes six and eight: a multicenter study. Am J Med Genet 1996; 67: 580–594.
Levinson DF, Holmans P, Straub RE, Owen MJ, Wildenauer DB, Gejman PV et al. Multicenter linkage study of schizophrenia candidate regions on chromosomes 5q, 6q, 10p, and 13q: schizophrenia linkage collaborative group III. Am J Hum Genet 2000; 67: 652–663.
Levinson DF, Holmans PA, Laurent C, Riley B, Pulver AE, Gejman PV et al. No major schizophrenia locus detected on chromosome 1q in a large multicenter sample. Science 2002; 296: 739–741.
Mowry BJ, Holmans PA, Pulver AE, Gejman PV, Riley B, Williams NM et al. Multicenter linkage study of schizophrenia loci on chromosome 22q. Mol Psychiatry 2004; 9: 784–795.
Williams NM, Norton N, Williams H, Ekholm B, Hamshere ML, Lindblom Y et al. A systematic genomewide linkage study in 353 sib pairs with schizophrenia. Am J Hum Genet 2003; 73: 1355–1367.
Faraone SV, Matise T, Svrakic D, Pepple J, Malaspina D, Suarez B et al. Genome scan of European–American schizophrenia pedigrees: results of the NIMH Genetics Initiative and Millennium Consortium. Am J Med Genet 1998; 81: 290–295.
Kaufmann CA, Suarez B, Malaspina D, Pepple J, Svrakic D, Markel PD et al. The NIMH Genetics Initiative Millennium Schizophrenia Consortium: linkage analysis of African–American pedigrees. Am J Med Genet 1998; 81: 282–289.
Maier W, Lichtermann D, Minges J, Hallmayer J, Heun R, Benkert O et al. Continuity and discontinuity of affective disorders and schizophrenia: results of a controlled family study. Arch Gen Psychiatry 1993; 50: 871–883.
Faraone SV, Blehar M, Pepple J, Moldin SO, Norton J, Nurnberger JI et al. Diagnostic accuracy and confusability analyses: an application to the Diagnostic Interview for Genetic Studies. Psychol Med 1996; 26: 401–410.
Oliphant A, Barker DL, Stuelpnagel JR, Chee MS . BeadArray technology: enabling an accurate, cost-effective approach to high-throughput genotyping. Biotechniques 2002; 32S: 60–61.
Kong A, Gudbjartsson DF, Sainz J, Jonsdottir GM, Gudjonsson SA, Richardsson B et al. A high-resolution recombination map of the human genome. Nat Genet 2002; 31: 241–247.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007; 81: 559–575.
Sun L, Wilder K, McPeek MS . Enhanced pedigree error detection. Hum Hered 2002; 54: 99–110.
Pritchard JK, Stephens M, Donnelly P . Inference of population structure using multilocus genotype data. Genetics 2000; 155: 945–959.
Freimer NB, Sandkuijl LA, Blower SM . Incorrect specification of marker allele frequencies: effects on linkage analysis. Am J Hum Genet 1993; 52: 1102–1110.
Kong A, Cox NJ . Allele-sharing models: LOD scores and accurate linkage tests. Am J Hum Genet 1997; 61: 1179–1188.
Abecasis GR, Cherny SS, Cookson WO, Cardon LR . Merlin—rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet 2002; 30: 97–101.
Levinson DF, Holmans P . The effect of linkage disequilibrium on linkage analysis of incomplete pedigrees. BMC Genet 2005; 30 (Suppl 1): S6.
Abecasis GR, Wigginton JE . Handling marker–marker linkage disequilibrium: pedigree analysis with clustered markers. Am J Hum Genet 2005; 77: 754–767.
Gudbjartsson DF, Jonasson K, Frigge ML, Kong A . Allegro, a new computer program for multipoint linkage analysis. Nat Genet 2000; 25: 12–13.
Holmans P . Detecting gene–gene interactions using affected sib pair analysis with covariates. Hum Hered 2002; 53: 92–102.
Holmans P, Weissman MM, Zubenko GS, Scheftner WA, Crowe RR, Depaulo Jr JR et al. Genetics of recurrent early-onset major depression (GenRED): final genome scan report. Am J Psychiatry 2007; 164: 248–258.
Suarez BK, Duan J, Sanders AR, Hinrichs AL, Jin CH, Hou C et al. Genomewide linkage scan of 409 European-ancestry and African American families with schizophrenia: suggestive evidence of linkage at 8p23.3-p21.2 and 11p13.1-q14.1 in the combined sample. Am J Hum Genet 2006; 78: 315–333.
Pulver AE, Lasseter VK, Kasch L, Wolyniec P, Nestadt G, Blouin JL et al. Schizophrenia: a genome scan targets chromosomes 3p and 8p as potential sites of susceptibility genes. Am J Med Genet 1995; 60: 252–260.
Stefansson H, Sigurdsson E, Steinthorsdottir V, Bjornsdottir S, Sigmundsson T, Ghosh S et al. Neuregulin 1 and susceptibility to schizophrenia. Am J Hum Genet 2002; 71: 877–892.
Munafo MR, Thiselton DL, Clark TG, Flint J . Association of the NRG1 gene and schizophrenia: a meta-analysis. Mol Psychiatry 2006; 11: 539–546.
Fallin MD, Lasseter VK, Wolyniec PS, McGrath JA, Nestadt G, Valle D et al. Genomewide linkage scan for schizophrenia susceptibility loci among Ashkenazi Jewish families shows evidence of linkage on chromosome 10q22. Am J Hum Genet 2003; 73: 601–611.
Faraone SV, Hwu HG, Liu CM, Chen WJ, Tsuang MM, Liu SK et al. Genome scan of Han Chinese schizophrenia families from Taiwan: confirmation of linkage to 10q22.3. Am J Psychiatry 2006; 163: 1760–1766.
Williams NM, Owen MJ . Genetic abnormalities of chromosome 22 and the development of psychosis. Curr Psychiatry Rep 2004; 6: 176–182.
Stefansson H, Rujescu D, Cichon S, Pietilainen OP, Ingason A, Steinberg S et al. Large recurrent microdeletions associated with schizophrenia. Nature 2008; 455: 232–236.
International Schizophrenia Consortium. Rare chromosomal deletions and duplications increase risk of schizophrenia. Nature 2008; 455: 237–241.
McQueen MB, Devlin B, Faraone SV, Nimgaonkar VL, Sklar P, Smoller JW et al. Combined analysis from eleven linkage studies of bipolar disorder provides strong evidence of susceptibility loci on chromosomes 6q and 8q. Am J Hum Genet 2005; 77: 582–595.
Abkevich V, Camp NJ, Hensel CH, Neff CD, Russell DL, Hughes DC et al. Predisposition locus for major depression at chromosome 12q22–12q23.2. Am J Hum Genet 2003; 73: 1271–1281.
McGuffin P, Knight J, Breen G, Brewster S, Boyd PR, Craddock N et al. Whole genome linkage scan of recurrent depressive disorder from the depression network study. Hum Mol Genet 2005; 14: 3337–3345.
Barden N, Harvey M, Gagne B, Shink E, Tremblay M, Raymond C et al. Analysis of single nucleotide polymorphisms in genes in the chromosome 12q24.31 region points to P2RX7 as a susceptibility gene to bipolar affective disorder. Am J Med Genet B Neuropsychiatr Genet 2006; 141: 374–382.
O’Donovan MC, Craddock N, Norton N, Williams H, Peirce T, Moskvina V et al. Identification of loci associated with schizophrenia by genome-wide association and follow-up. Nat Genet 2008; 40: 1053–1055.
Walsh T, McClellan JM, McCarthy SE, Addington AM, Pierce SB, Cooper GM et al. Rare structural variants disrupt multiple genes in neurodevelopmental pathways in schizophrenia. Science 2008; 320: 539–543.
Xu B, Roos JL, Levy S, van Rensburg EJ, Gogos JA, Karayiorgou M . Strong association of de novo copy number mutations with sporadic schizophrenia. Nat Genet 2008; 40: 880–885.
Pritchard JK, Cox NJ . The allelic architecture of human disease genes: common disease-common variant… or not? Hum Mol Genet 2002; 11: 2417–2423.
Kryukov GV, Pennacchio LA, Sunyaev SR . Most rare missense alleles are deleterious in humans: implications for complex disease and association studies. Am J Hum Genet 2007; 80: 727–739.
Botstein D, Risch N . Discovering genotypes underlying human phenotypes: past successes for mendelian disease, future approaches for complex disease. Nat Genet 2003; 33 (Suppl): 228–237.
Cohen JC, Kiss RS, Pertsemlidis A, Marcel YL, McPherson R, Hobbs HH . Multiple rare alleles contribute to low plasma levels of HDL cholesterol. Science 2004; 305: 869–872.
Ji W, Foo JN, O’Roak BJ, Zhao H, Larson MG, Simon DB et al. Rare independent mutations in renal salt handling genes contribute to blood pressure variation. Nat Genet 2008; 40: 495##.
Roeder K, Bacanu SA, Wasserman L, Devlin B . Using linkage genome scans to improve power of association in genome scans. Am J Hum Genet 2006; 78: 243–252.
Acknowledgements
We thank the many family members who participated in the studies that recruited these samples. This work was supported by National Institute of Mental Health grants 7R01MH062276 (to DFL, CL, MO and DW), 5R01MH068922 (to PG), 5R01MH068921 (to AEP) and 5R01MH068881 (to BR). For the NIMH sample, data and biomaterials were collected in three projects that participated in the National Institute of Mental Health (NIMH) Schizophrenia Genetics Initiative. From 1991 to 97, the principal investigators and co-investigators were: Harvard University, Boston, MA, U01 MH46318, Ming T Tsuang, Stephen Faraone and John Pepple; Washington University, St Louis, MO, U01 MH46276, C Robert Cloninger, Theodore Reich and Dragan Svrakic; Columbia University, New York, NY U01 MH46289, Charles Kaufmann, Dolores Malaspina and Jill Harkavy Friedman. Genotyping services were provided by the Center for Inherited Disease Research (CIDR). CIDR is fully funded through a federal contract from the National Institutes of Health to The Johns Hopkins University, contract number N01-HG-65403. The NIMH Cell Repository at Rutgers University (Dr Douglas Fugman and Dr Jay Tischfield) and the NIMH Center for Collaborative Genetic Studies on Mental Disorders (Dr John Rice) made important contributions to this project.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website (http://www.nature.com/mp)
Supplementary information
Rights and permissions
About this article
Cite this article
Holmans, P., Riley, B., Pulver, A. et al. Genomewide linkage scan of schizophrenia in a large multicenter pedigree sample using single nucleotide polymorphisms. Mol Psychiatry 14, 786–795 (2009). https://doi.org/10.1038/mp.2009.11
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2009.11
Keywords
This article is cited by
-
Evidence for contribution of common genetic variants within chromosome 8p21.2-8p21.1 to restricted and repetitive behaviors in autism spectrum disorders
BMC Genomics (2016)
-
A genome-wide screen for acrophobia susceptibility loci in a Finnish isolate
Scientific Reports (2016)
-
Brain-specific Crmp2 deletion leads to neuronal development deficits and behavioural impairments in mice
Nature Communications (2016)
-
CRMPs: critical molecules for neurite morphogenesis and neuropsychiatric diseases
Molecular Psychiatry (2015)
-
Genome-wide study of association and interaction with maternal cytomegalovirus infection suggests new schizophrenia loci
Molecular Psychiatry (2014)