Abstract
Immunohistochemistry plays a key role in the diagnosis of soft tissue tumors. Until recently, however, the primary purpose of immunohistochemistry in this context was simply to attempt to demonstrate a line of differentiation. Unfortunately, most traditional markers (predominantly directed against cytoplasmic determinants) show relatively limited specificity. Over the last decade or so, much more specific immunohistochemical markers for soft tissue tumors have been developed. This review will provide an update of some of the most useful new diagnostic markers, which are significantly changing clinical practice for surgical pathologists, separated into three general categories: (1) lineage-restricted transcription factors, (2) protein correlates of molecular alterations, and (3) diagnostic markers identified by gene expression profiling.
Similar content being viewed by others
Main
Immunohistochemistry plays a key role in the diagnosis of soft tissue tumors. Until recently, however, the primary purpose of immunohistochemistry in this context was simply to attempt to demonstrate a line of differentiation. Unfortunately, most traditional markers (predominantly directed against cytoplasmic determinants) show relatively limited specificity. For example, smooth muscle (α) actin (SMA) is not only a marker of smooth muscle neoplasms but is also typically expressed by myofibroblastic lesions; SMA may sometimes also be detected in essentially any neoplasm showing spindle cell morphology (eg, sarcomatoid carcinoma). S100 protein (S100B), the most widely used Schwann cell marker, is consistently expressed in benign nerve sheath tumors and more variably in a subset of malignant peripheral nerve sheath tumors, but is also positive in melanocytic, cartilaginous, and myoepithelial tumors, as well as chordoma, interdigitating dendritic cell sarcoma, Langerhans cell histiocytosis, and occasional carcinomas (most notably, breast carcinoma). Over the last decade or so, much more specific immunohistochemical markers for soft tissue tumors have been developed. This review will provide an update of some of the most useful new diagnostic markers, which are significantly changing clinical practice for surgical pathologists, separated into three general categories: (1) lineage-restricted transcription factors, (2) protein correlates of molecular alterations, and (3) diagnostic markers identified by gene expression profiling (Table 1).
Lineage-restricted transcription factors
Antibodies that recognize nuclear transcription factors are increasingly finding applications in surgical pathology, such as the determination of possible anatomic sites for a carcinoma of unknown primary. Some of these markers (eg, TTF1 and CDX2) have been available in most clinical immunohistochemistry laboratories for over a decade. Other markers (eg, PAX8) have more recently been introduced into clinical practice. Until recently, few antibodies directed against transcription factors have been used for the diagnosis of soft tissue tumors.
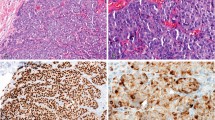
The most widely used lineage-specific transcription factor for soft tissue tumors is myogenin (MYF4), whose expression is limited to skeletal muscle cells.1 Myogenin is indispensable in the diagnosis of rhabdomyosarcoma, and can also be applied to confirm the presence of heterologous rhabdomyoblastic differentiation in other tumor types (eg, dedifferentiated liposarcoma and malignant peripheral nerve sheath tumor).2 The extent of staining correlates with histologic type: whereas diffuse nuclear reactivity for myogenin is typical of alveolar rhabdomyosarcoma, embryonal rhabdomyosarcoma usually shows more heterogeneous staining (in 30–70% of tumor cells; Figure 1).3 Pleomorphic rhabdomyosarcoma and spindle cell/sclerosing rhabdomyosarcoma may show only limited myogenin expression in a small subset of cells.4, 5 MYOD1 (MYF3) is another skeletal muscle-specific transcription factor. Available antibodies may be more difficult to optimize than those directed against myogenin and sometimes show cytoplasmic background staining.1
Myogenin expression in rhabdomyosarcomas. (a) Alveolar rhabdomyosarcoma showing a solid growth pattern. (b) Myogenin shows strong nuclear staining in nearly all tumor cells. (s) Embryonal rhabdomyosarcoma composed of round-to-short spindled cells in a myxoid stroma. (d) Only a subset of tumor cells is positive for myogenin.
The expression of two related ETS family transcription factors is relatively restricted to endothelial cells and derived neoplasms. Although FLI1 (most famous for its role in the Ewing sarcoma translocation; see below) is expressed in nearly all endothelial lesions,6 FLI1 is also positive in some normal lymphocytes, lymphoblastic lymphomas, and a subset of a wide range of mesenchymal neoplasms.7, 8 Consequently, FLI1 has limited utility in differential diagnosis. In contrast, recent studies have shown that ERG is a highly sensitive and specific marker for endothelial differentiation.7, 9, 10 ERG shows nuclear staining in benign, rarely metastasizing, and malignant vascular lesions, including nearly all epithelioid hemangioendotheliomas, pseudomyogenic hemangioendotheliomas, and even poorly differentiated (epithelioid and spindle cell) angiosarcomas, which are particularly challenging to diagnose (Figure 2), but in almost no other mesenchymal neoplasms.9, 10 Of note, some anti-ERG antibodies show cross-reactivity with FLI1, reducing the specificity of such reagents. Immunoreactivity for ERG may also be observed in epithelioid sarcomas, particularly using antibodies that recognize the N-terminus of ERG;11, 12 an antibody directed against the C-terminus of ERG is rarely positive in epithelioid sarcoma.12 ERG is also positive in a selected group of tumors that harbor translocations involving ERG: ∼50% of prostatic adenocarcinomas, 5–10% of Ewing sarcomas, and a small subset of acute myeloid leukemias.13
Brachyury (T) is a transcriptional activator involved in the development of the notochord.14 Nuclear expression of brachyury is a consistent feature of chordoma (Figure 3), which is a helpful diagnostic immunohistochemical finding to distinguish this tumor type from potential histologic mimics, including chondrosarcoma, metastatic carcinoma, and myoepithelial tumors.14, 15, 16, 17 No other consistently brachyury-positive tumor types have been identified. Of note, a subset of chordomas show copy number gains at the T locus.18 T gene duplications have been identified as a susceptibility factor for familial chordoma,19 and single-nucleotide variants in T are also associated with a risk of developing chordoma.20
SOX10 is a member of the SOX (SRY-related HMG-box) family of transcription factors that is important for embryonic development and cell fate determination of the neural crest and peripheral nervous system.21 As such, SOX10 is a relatively specific marker for neuroectodermal neoplasms, consistently expressed in benign nerve sheath tumors, clear cell sarcoma, and melanoma (including desmoplastic and spindle cell variants; Figure 4).21, 22, 23 Unfortunately, SOX10 shows lower sensitivity for malignant peripheral nerve sheath tumors (30–50%) and, when positive, is usually detected in only a small subset of tumor cells (Figure 4).22, 24 Reactivity for SOX10 is also observed in astrocytomas, myoepithelial tumors, selected other types of salivary gland neoplasms, and a subset of breast carcinomas, particularly basal-like, ‘triple-negative,’ and metaplastic carcinomas.21, 25, 26
SOX10 (SRY-related HMG-box 10) expression in nerve sheath tumors. (a) Neurofibroma with buckled nuclei and prominent collagenous stroma. (b) SOX10 is positive in the Schwann cell component. (c) Malignant peripheral nerve sheath tumor with fascicular architecture, tapering nuclei, and a high mitotic rate. (d) Nuclear reactivity for SOX10 is observed in a small subset of tumor cells.
The most recent addition to this group of markers is SATB2 (special AT-rich sequence-binding protein 2). SATB2 is a nuclear matrix protein that plays a critical role in osteoblast lineage commitment.27 SatB2 knockout mice show impaired osteoblast differentiation and craniofacial defects.27 Deletion of SATB2 also partially underlies the craniofacial malformations observed in the rare human chromosome 2q32–q33 deletion syndrome.28 SATB2 is a sensitive and specific immunohistochemical marker of osteoblastic differentiation in bone and soft tissue tumors.29 Although nuclear staining for SATB2 is seen in nearly all cases of osteosarcoma (Figure 5), SATB2 expression is also a feature of benign osteoblastic neoplasms, as well as other sarcoma types with heterologous osteosarcomatous differentiation (eg, dedifferentiated chondrosarcoma and dedifferentiated liposarcoma).29 As such, SATB2 expression must be interpreted carefully in the context of histologic features and radiologic findings. SATB2 may be a particularly helpful diagnostic adjunct for osteosarcoma in two specific settings: (1) when the histologic features of the matrix are equivocal (ie, osteoid vs hyalinized collagen) and (2) when the biopsy only samples tumor with an undifferentiated appearance. Of note, soft tissue tumors with abundant hyalinized collagen (eg, sclerosing epithelioid fibrosarcoma (SEF) and sclerosing rhabdomyosarcoma) are consistently negative for SATB2.29 SATB2 has previously also been shown to be a sensitive and specific marker for colorectal origin among carcinoma.30
Protein correlates of molecular alterations in soft tissue tumors
β-Catenin is the product of the CTNNB1 gene involved in the Wnt signaling pathway (along with APC). Mutations in CTNNB1 are found in 85–90% of sporadic desmoid tumors;31 patients with familial adenomatous polyposis and a germline mutation in APC also develop desmoid fibromatosis (Gardner syndrome), especially in the mesentery of the small intestine. Immunohistochemistry for β-catenin shows aberrant nuclear staining (as opposed to the normal cell membranous pattern of staining) in the majority of cases of desmoid fibromatosis (Figure 6) and is therefore useful to confirm the diagnosis, particularly in core needle biopsies or other limited samples in which the distinction between scar tissue and desmoid tumor (or other bland spindle cell neoplasms) may be challenging.32, 33, 34 Of note, nuclear immunoreactivity may sometimes be limited in extent; examination under high magnification is often required to identify nuclear staining. Although β-catenin immunohistochemistry has a reasonable sensitivity for desmoid fibromatosis (70–90%) and is consistently negative in other intra-abdominal mesenchymal neoplasms (including gastrointestinal stromal tumors (GISTs) and smooth muscle tumors),32, 34 it is not entirely specific; a subset of solitary fibrous tumors (20–40%) and low-grade myofibroblastic sarcomas (30%) also show nuclear staining.35 β-Catenin must therefore be interpreted carefully in the context of clinical findings and morphology.
Atypical lipomatous tumor/well-differentiated liposarcoma and dedifferentiated liposarcoma are characterized by supernumerary ring and giant marker chromosomes, composed of amplified material from the long arm of chromosome 12 (q13∼15), within which several oncogenes are located, leading to overexpression of MDM2 and CDK4.36 Nuclear reactivity for MDM2 and CDK4 is therefore highly sensitive for well-differentiated and dedifferentiated liposarcomas, but is not entirely specific, as some other spindle cell and pleomorphic sarcomas (such as malignant peripheral nerve sheath tumor, myxofibrosarcoma, and rhabdomyosarcoma) show nuclear staining in a subset of cases.37, 38, 39, 40 Combined positive staining for both MDM2 and CDK4 is much more specific.37 Identification of MDM2 gene amplification by fluorescence in situ hybridization (FISH) is highly specific for well-differentiated and dedifferentiated liposarcoma (more specific than immunohistochemistry).41 Nonetheless, in the proper context, immunohistochemistry for MDM2 and CDK4 can be very helpful in differential diagnosis, particularly in the distinction between well-differentiated liposarcoma and spindle cell/pleomorphic lipoma or conventional lipoma, and between dedifferentiated liposarcoma and other high-grade sarcomas in a core needle biopsy of a large retroperitoneal or intra-abdominal mass (Figure 7). Our understanding of the molecular genetics of these tumor types has also helped clarify the nosology of so-called ‘inflammatory malignant fibrous histiocytoma,’ nearly all cases of which in fact represent not a distinct tumor type but a morphologic pattern in dedifferentiated liposarcoma (Figure 7).42
MDM2 and CDK4 expression in liposarcomas. (a) Well-differentiated liposarcoma showing variation in adipocyte size and scattered hyperchromatic nuclei. (b) Occasional nuclei are positive for MDM2. (c) Dedifferentiated liposarcoma showing non-distinctive histology with mildly atypical spindle cells arranged in a storiform architecture. (d) CDK4 is positive in the majority of tumor cells. (e) Dedifferentiated liposarcoma showing an ‘inflammatory malignant fibrous histiocytoma (MFH)’ histologic pattern. (f) Nuclear reactivity for MDM2 is observed. Nearly all cases of so-called ‘inflammatory MFH’ represent dedifferentiated liposarcomas.
SMARCB1 (also known as INI1 and SNF5) is a member of the SWI/SNF multisubunit chromatin remodeling complex that plays a fundamental role in regulating transcription.43 SMARCB1 is ubiquitously expressed in normal cells. Biallelic inactivation of SMARCB1 is essentially a defining feature of malignant rhabdoid tumors of infancy.44 Loss of normal nuclear expression of SMARCB1 is useful to confirm the diagnosis of this rare, aggressive tumor type (Figure 8).45, 46 More recent studies have also demonstrated a tumor suppressor role of SMARCB1 in epithelioid sarcoma (both conventional and proximal type).47, 48, 49 Homozygous deletion of the SMARCB1 locus is also common in epithelioid sarcoma,50 and ∼95% of epithelioid sarcomas show loss of SMARCB1 protein expression by immunohistochemistry (Figure 8).48 Immunohistochemistry is therefore helpful in differential diagnosis, as metastatic carcinomas and epithelioid vascular neoplasms (angiosarcoma and epithelioid hemangioendothelioma) consistently retain SMARCB1 protein expression (Figure 8).48 Several other tumor types also show loss of SMARCB1 expression, including renal medullary carcinomas, ∼50% of epithelioid malignant peripheral nerve sheath tumors, and a subset of myoepithelial carcinomas of soft tissue, extraskeletal myxoid chondrosarcomas, and poorly differentiated chordomas.43, 48, 51, 52
SMARCB1 (INI1)-deficient tumors. (a) Malignant rhabdoid tumor composed of epithelioid cells with vesicular nuclei, prominent nucleoli, and eosinophilic hyaline cytoplasmic inclusions. (b) Loss of SMARCB1 protein expression is a characteristic finding. (c) Conventional epithelioid sarcoma composed of bland epithelioid cells infiltrating dermal collagen. (d) Expression of SMARCB1 is lost in tumor cells. Note the normal (retained) nuclear staining in stromal and endothelial cells. (e) Proximal-type epithelioid sarcoma showing large-cell morphology and marked nuclear atypia. (f) Loss of SMARCB1 staining is observed in nearly all cases. (g) Adrenal cortical carcinoma mimicking malignant rhabdoid tumor. (h) Expression of SMARCB1 protein is retained in tumor cells.
Familial paraganglioma syndrome, the most common inherited paraganglioma predisposition syndrome, is caused by germline mutations in the succinate dehydrogenase (SDH) subunit genes.53, 54 The SDH complex resides in the inner mitochondrial membrane, where it participates in both the citric acid cycle (catalyzing the oxidation of succinate to fumarate) and the electron transport chain (complex II). Mutations in SDH subunit genes lead to loss of protein expression and enzyme function. Immunohistochemistry for SDHB shows loss of the normal granular cytoplasmic (mitochondrial) staining pattern in such tumors, irrespective of which gene is mutated.54 Similarly, immunohistochemistry for SDHB can help identify a clinically and histologically distinctive class of GISTs of the stomach, referred to as SDH-deficient GISTs (Figure 9).55, 56, 57, 58, 59 GISTs with loss of SDH expression are a feature of Carney–Stratakis syndrome, a variant of familial paraganglioma syndrome in which patients also develop such GISTs, as well as Carney triad (paraganglioma, gastric GIST, and pulmonary chondroma), although the genetic basis of the latter non-familial syndrome has not been determined.58 The rare GISTs that present in childhood are nearly all SDH-deficient, and recent studies have shown that 8–10% of gastric GISTs overall are also SDH-deficient.56, 57 The molecular mechanisms underlying SDH dysfunction in the majority of SDH-deficient GISTs (outside of the Carney–Stratakis syndrome) are unknown; SDH subunit mutations are found in only a subset of patients.53, 56
Succinate dehydrogenase (SDH)-deficient gastrointestinal stromal tumor (GIST). (a) SDH-deficient GISTs arise in the stomach and show a multinodular growth pattern. (b) Similar to conventional GISTs, KIT is strongly positive. (c) Most SDH-deficient GISTs are dominated by epithelioid cytomorphology. (d) Loss of SDHB protein expression confirms the diagnosis. Note the granular cytoplasmic staining in endothelial cells, which serve as an internal control.
SDH-deficient GISTs (previously briefly referred to as ‘pediatric-type’ or ‘type 2’ GISTs) show a distinctive multinodular or plexiform growth pattern and epithelioid or mixed morphology, and can therefore be recognized histologically (Figure 9).55, 59, 60 Similar to conventional KIT-mutant GISTs, SDH-deficient GISTs are usually positive for KIT, DOG1 (discovered on GIST1), and CD34 by immunohistochemistry. Risk assessment criteria currently widely used for GIST do not reliably predict behavior for this class of tumors, although they have a significant potential for metastasis to lymph nodes (unlike conventional GIST), peritoneum, and liver.53, 60 Metastases are clinically indolent (some patients with metastatic disease survive without therapy for decades) and show limited (if any) response to imatinib. Most recently, SDHA mutations have been identified as a common cause of SDH-deficient GIST (around one-third of all such tumors); of note, loss of SDHA protein expression reliably predicts the presence of SDHA mutations (unlike loss of SDHB, which does not indicate the gene involved, if any).61, 62, 63, 64 SDH-deficient GISTs with mutations in other SDH subunit genes (and those without identifiable SDH mutations) retain normal cytoplasmic staining for SDHA.62, 64 A small subset of renal cell carcinomas also shows loss of SDHB protein expression; some (but not all) such tumors are characterized by a distinctive oncocytoid cytomorphology.65, 66
Several antibodies that recognize protein products of gene rearrangements in sarcomas can be applied clinically for diagnostic surgical pathology. Alveolar soft part sarcoma harbors the translocation t(X;17), which leads to the ASPSCR1-TFE3 fusion and overexpression of the TFE3 protein.67 A similar translocation is found in Xp11 translocation renal cell carcinomas,68 and TFE3 rearrangements have recently been identified in a small subset of PEComas69 and a distinctive subset of epithelioid hemangioendotheliomas.70 By immunohistochemistry, these tumors show nuclear staining for TFE3, which can be used to confirm the diagnosis (Figure 10).69, 70, 71
The most common translocation in Ewing sarcoma is t(11;22), found in ∼90% of cases. This translocation leads to EWSR1-FLI1 fusion and overexpression of the FLI1 protein. By immunohistochemistry, Ewing sarcomas usually show strong nuclear staining for FLI1.8, 72 However, as mentioned previously, FLI1 is also expressed in endothelial cells and derived neoplasms, some non-neoplastic lymphocytes, lymphoblastic lymphomas, and occasionally in a variety of other tumor types.6, 7, 8, 9 Given its lack of specificity, FLI1 immunohistochemistry is not particularly helpful in differential diagnosis. As mentioned above, ERG is an excellent marker for endothelial neoplasms.9 In addition, a small subset of Ewing sarcomas (ie, those with EWSR1-ERG rearrangement) is strongly positive for ERG as well.73
Approximately 50% of inflammatory myofibroblastic tumors (IMTs) harbor rearrangements of the ALK locus, which usually results in ALK protein expression detectable by immunohistochemistry.74, 75, 76, 77, 78 Interestingly, the pattern of ALK staining appears to correlate with the fusion partner. Most translocations (involving the fusion partners TPM3, TPM4, CARS, ATIC, SEC31A, and CLTC) result in cytoplasmic staining, whereas a nuclear membrane pattern of staining is seen with the RANBP2 fusion partner (a nuclear pore protein; Figure 11).79 IMTs with RANBP2-ALK fusion have a predilection for the omentum and mesentery; show distinctive (epithelioid) morphology, myxoid stroma, and often a neutrophilic inflammatory infiltrate; and pursue an aggressive clinical course (leading to the designation ‘epithelioid inflammatory myofibroblastic sarcoma’).79 As this distinctive IMT variant can be positive for CD30, it may be confused with anaplastic large-cell lymphoma; desmin expression is helpful to distinguish among these possibilities, as desmin is consistently negative in this lymphoma type. Among mesenchymal neoplasms, ALK expression is not entirely specific for IMT; eg, it is also positive in some rhabdomyosarcomas and malignant peripheral nerve sheath tumors.74, 80 A subset of neuroblastomas and pulmonary adenocarcinomas also overexpress ALK (the latter usually secondary to EML4-ALK fusion).81, 82 Most other fibroblastic/myofibroblastic tumors (including desmoid fibromatosis, nodular fasciitis, myofibroma, and infantile fibrosarcoma) are negative for ALK.74
ALK expression in inflammatory myofibroblastic tumor. (a) Inflammatory myofibroblastic tumor composed of plump spindle cells arranged in loose fascicles with prominent stromal lymphocytes. (b) Cytoplasmic staining for ALK is observed in ∼50% of cases. (c) Epithelioid inflammatory myofibroblastic sarcoma with amphophilic cytoplasm, prominent nucleoli, myxoid stroma, and stromal neutrophils. (d) There is a characteristic nuclear membrane pattern of ALK staining.
Recently, a NAB2-STAT6 fusion has been identified by several independent groups as a consistent feature of solitary fibrous tumors (including cellular examples formerly known as ‘hemangiopericytoma,’ as well as histologically atypical and malignant variants).83, 84, 85 The so-called ‘meningeal hemangiopericytoma’ also harbors this fusion gene, supporting the classification of this tumor type with solitary fibrous tumor (similar to examples arising in soft tissue and at visceral sites).86 As these two fusion partners reside in close proximity on chromosome 12q13, this gene fusion cannot be identified by conventional FISH analysis.84 NAB2-STAT6 fusion leads to high-level nuclear expression of the STAT6 protein.86 By immunohistochemistry, nearly all cases of solitary fibrous tumor (including cellular and malignant variants) show strong nuclear staining for STAT6 (Figure 12), whereas other spindle cell neoplasms that might be considered in the differential diagnosis are consistently negative.86, 87
Diagnostic markers identified by gene expression profiling
DOG1 was shown to be highly expressed in GIST by gene expression profiling.88 Also known as ANO1 (anoctamin 1) and TMEM16A, this protein is a calcium-activated chloride channel expressed in normal interstitial cells of Cajal that plays a fundamental role in generation of slow waves during peristalsis.89 DOG1 is a highly sensitive and specific marker for GIST, and it is also positive in a subset of KIT-negative tumors (Figure 13).90, 91, 92, 93 DOG1 is most helpful as a diagnostic adjunct for epithelioid gastric and omental GISTs with PDGFRA mutations (which are often KIT negative or show limited KIT immunoreactivity). Notable other tumor types that sometimes express DOG1 include a small subset of uterine-type retroperitoneal leiomyomas and synovial sarcomas.93
DOG1 (discovered on GIST1) expression in gastrointestinal stromal tumor. (a) Epithelioid gastrointestinal stromal tumor of the stomach with a PDGFRA mutation. (b) KIT is negative in tumor cells. Note the strong membranous staining in occasional mast cells, which serve as an internal control. (c) DOG1 is positive with a membranous staining pattern.
TLE1 (transducin-like enhancer of split 1) is a transcriptional corepressor that inhibits Wnt signaling. Using gene expression profiling, TLE1 was found to be an excellent discriminator of synovial sarcoma from other sarcoma types.94 By immunohistochemistry, TLE1 shows moderate-to-strong, diffuse nuclear staining in the majority of synovial sarcomas (80–90%) (Figure 14).95, 96, 97, 98 TLE1 is a relatively specific marker for synovial sarcoma that can be helpful in differential diagnosis, as it is only positive in a small subset of malignant peripheral nerve sheath tumors and solitary fibrous tumors (usually with only weak staining), and it is consistently negative in Ewing sarcomas (which can be mistaken for poorly differentiated synovial sarcomas).95, 97
MUC4 is a high-molecular-weight transmembrane glycoprotein expressed on the surface of some glandular epithelial cell types. Gene expression profiling has recently indicated that this epithelial mucin is highly expressed by low-grade fibromyxoid sarcoma (LGFMS),99 which is typically characterized by the t(7;16) translocation, resulting in FUS-CREB3L2 rearrangement, or more rarely t(11;16), resulting in FUS-CREB3L1 fusion.100, 101 MUC4 is a highly sensitive and specific immunohistochemical marker for LGFMS among spindle cell neoplasms (MUC4 is negative in soft tissue perineurioma, malignant peripheral nerve sheath tumor, solitary fibrous tumor, desmoid fibromatosis, and intramuscular/cellular myxoma) and can therefore be helpful in confirming the diagnosis (Figure 15).102 SEF is a rare aggressive sarcoma type composed of cords of epithelioid cells in a densely hyalinized stroma. Until recently, no helpful diagnostic markers were available. Some cases of SEF harbor the same genetic alterations as, and also may show hybrid histologic features with LGFMS.100, 103 Hybrid tumors with both LGFMS and SEF components, as well as ∼70% of ‘pure’ SEF cases, are strongly positive for MUC4 (Figure 15).103 Immunohistochemistry for MUC4 can be helpful in differential diagnosis, as nearly all other epithelioid bone and soft tissue tumors are negative for this marker, with the exception of the glandular component of biphasic synovial sarcoma, although this tumor type is unlikely to pose diagnostic problems.103, 104
MUC4 expression in fibroblastic sarcomas. (a) Low-grade fibromyxoid sarcoma showing a storiform-to-whorled growth pattern and bland spindle cell morphology. These tumors can easily be mistaken for benign neoplasms. (b) Strong cytoplasmic staining for MUC4 is highly specific for this tumor type among spindle cell neoplasms. (c) Sclerosing epithelioid fibrosarcoma composed of cords of epithelioid cells embedded in a densely hyalinized stroma. (d) MUC4 is positive in most cases.
GRIA2 is a subunit of a class of glutamate receptors involved in the transmission of excitatory synaptic signals in the central nervous system. In a recent study, global gene expression profiling identified GRIA2 as the top upregulated gene in solitary fibrous tumors (100-fold higher than control tissues).84 By immunohistochemistry using a tissue microarray, GRIA2 was positive in the majority of solitary fibrous tumors and was negative in other spindle cell neoplasms.84 These preliminary observations require confirmation in additional studies.
Summary
We have entered a new era of diagnostic immunohistochemistry for soft tissue tumor pathology. Antibodies directed against lineage-restricted transcription factors are helpful and relatively specific diagnostic markers. Molecular genetic findings can lead to new inexpensive and rapid diagnostic tests in the form of a single immunohistochemical stain. Finally, gene expression profiling can provide novel markers to discriminate among classes of histologically similar tumors, in some cases potentially obviating confirmation by molecular genetic techniques.
References
Wang NP, Marx J, McNutt MA et al. Expression of myogenic regulatory proteins (myogenin and MyoD1) in small blue round cell tumors of childhood. Am J Pathol 1995;147:1799–1810.
Folpe AL . MyoD1 and myogenin expression in human neoplasia: a review and update. Adv Anat Pathol 2002;9:198–203.
Cessna MH, Zhou H, Perkins SL et al. Are myogenin and myoD1 expression specific for rhabdomyosarcoma? A study of 150 cases, with emphasis on spindle cell mimics. Am J Surg Pathol 2001;25:1150–1157.
Folpe AL, McKenney JK, Bridge JA et al. Sclerosing rhabdomyosarcoma in adults: report of four cases of a hyalinizing, matrix-rich variant of rhabdomyosarcoma that may be confused with osteosarcoma, chondrosarcoma, or angiosarcoma. Am J Surg Pathol 2002;26:1175–1183.
Furlong MA, Mentzel T, Fanburg-Smith JC . Pleomorphic rhabdomyosarcoma in adults: a clinicopathologic study of 38 cases with emphasis on morphologic variants and recent skeletal muscle-specific markers. Mod Pathol 2001;14:595–603.
Folpe AL, Chand EM, Goldblum JR et al. Expression of Fli-1, a nuclear transcription factor, distinguishes vascular neoplasms from potential mimics. Am J Surg Pathol 2001;25:1061–1066.
McKay KM, Doyle LA, Lazar AJ et al. Expression of ERG, an Ets family transcription factor, distinguishes cutaneous angiosarcoma from histological mimics. Histopathology 2012;61:989–991.
Rossi S, Orvieto E, Furlanetto A et al. Utility of the immunohistochemical detection of FLI-1 expression in round cell and vascular neoplasm using a monoclonal antibody. Mod Pathol 2004;17:547–552.
Miettinen M, Wang ZF, Paetau A et al. ERG transcription factor as an immunohistochemical marker for vascular endothelial tumors and prostatic carcinoma. Am J Surg Pathol 2011;35:432–441.
Yaskiv O, Rubin BP, He H et al. ERG protein expression in human tumors detected with a rabbit monoclonal antibody. Am J Clin Pathol 2012;138:803–810.
Miettinen M, Wang Z, Sarlomo-Rikala M et al. ERG expression in epithelioid sarcoma: a diagnostic pitfall. Am J Surg Pathol 2013;37:1580–1585.
Stockman DL, Hornick JL, Deavers MT et al. ERG and FLI1 protein expression in epithelioid sarcoma. Mod Pathol advance online publication, 27 September 2013; doi:10.1038/modpathol.2013.161 (e-pub ahead of print).
Shah RB . Clinical applications of novel ERG immunohistochemistry in prostate cancer diagnosis and management. Adv Anat Pathol 2013;20:117–124.
Vujovic S, Henderson S, Presneau N et al. Brachyury, a crucial regulator of notochordal development, is a novel biomarker for chordomas. J Pathol 2006;209:157–165.
Oakley GJ, Fuhrer K, Seethala RR . Brachyury, SOX-9, and podoplanin, new markers in the skull base chordoma vs chondrosarcoma differential: a tissue microarray-based comparative analysis. Mod Pathol 2008;21:1461–1469.
Sangoi AR, Karamchandani J, Lane B et al. Specificity of brachyury in the distinction of chordoma from clear cell renal cell carcinoma and germ cell tumors: a study of 305 cases. Mod Pathol 2011;24:425–429.
Tirabosco R, Mangham DC, Rosenberg AE et al. Brachyury expression in extra-axial skeletal and soft tissue chordomas: a marker that distinguishes chordoma from mixed tumor/myoepithelioma/parachordoma in soft tissue. Am J Surg Pathol 2008;32:572–580.
Presneau N, Shalaby A, Ye H et al. Role of the transcription factor T (brachyury) in the pathogenesis of sporadic chordoma: a genetic and functional-based study. J Pathol 2011;223:327–335.
Yang XR, Ng D, Alcorta DA et al. T (brachyury) gene duplication confers major susceptibility to familial chordoma. Nat Genet 2009;41:1176–1178.
Pillay N, Plagnol V, Tarpey PS et al. A common single-nucleotide variant in T is strongly associated with chordoma. Nat Genet 2012;44:1185–1187.
Ordonez NG . Value of SOX10 immunostaining in tumor diagnosis. Adv Anat Pathol 2013;20:275–283.
Nonaka D, Chiriboga L, Rubin BP . Sox10: a pan-schwannian and melanocytic marker. Am J Surg Pathol 2008;32:1291–1298.
Palla B, Su A, Binder S et al. SOX10 expression distinguishes desmoplastic melanoma from its histologic mimics. Am J Dermatopathol 2013;35:576–581.
Karamchandani JR, Nielsen TO, van de Rijn M et al. Sox10 and S100 in the diagnosis of soft-tissue neoplasms. Appl Immunohistochem Mol Morphol 2012;20:445–450.
Cimino-Mathews A, Subhawong AP, Elwood H et al. Neural crest transcription factor Sox10 is preferentially expressed in triple-negative and metaplastic breast carcinomas. Hum Pathol 2013;44:959–965.
Ohtomo R, Mori T, Shibata S et al. SOX10 is a novel marker of acinus and intercalated duct differentiation in salivary gland tumors: a clue to the histogenesis for tumor diagnosis. Mod Pathol 2013;26:1041–1050.
Dobreva G, Chahrour M, Dautzenberg M et al. SATB2 is a multifunctional determinant of craniofacial patterning and osteoblast differentiation. Cell 2006;125:971–986.
FitzPatrick DR, Carr IM, McLaren L et al. Identification of SATB2 as the cleft palate gene on 2q32-q33. Hum Mol Genet 2003;12:2491–2501.
Conner JR, Hornick JL . SATB2 is a novel marker of osteoblastic differentiation in bone and soft tissue tumours. Histopathology 2013;63:36–49.
Magnusson K, de Wit M, Brennan DJ et al. SATB2 in combination with cytokeratin 20 identifies over 95% of all colorectal carcinomas. Am J Surg Pathol 2011;35:937–948.
Lazar AJ, Tuvin D, Hajibashi S et al. Specific mutations in the beta-catenin gene (CTNNB1) correlate with local recurrence in sporadic desmoid tumors. Am J Pathol 2008;173:1518–1527.
Bhattacharya B, Dilworth HP, Iacobuzio-Donahue C et al. Nuclear beta-catenin expression distinguishes deep fibromatosis from other benign and malignant fibroblastic and myofibroblastic lesions. Am J Surg Pathol 2005;29:653–659.
Montgomery E, Folpe AL . The diagnostic value of beta-catenin immunohistochemistry. Adv Anat Pathol 2005;12:350–356.
Montgomery E, Torbenson MS, Kaushal M et al. Beta-catenin immunohistochemistry separates mesenteric fibromatosis from gastrointestinal stromal tumor and sclerosing mesenteritis. Am J Surg Pathol 2002;26:1296–1301.
Carlson JW, Fletcher CD . Immunohistochemistry for beta-catenin in the differential diagnosis of spindle cell lesions: analysis of a series and review of the literature. Histopathology 2007;51:509–514.
Coindre JM, Pedeutour F, Aurias A . Well-differentiated and dedifferentiated liposarcomas. Virchows Arch 2010;456:167–179.
Binh MB, Sastre-Garau X, Guillou L et al. MDM2 and CDK4 immunostainings are useful adjuncts in diagnosing well-differentiated and dedifferentiated liposarcoma subtypes: a comparative analysis of 559 soft tissue neoplasms with genetic data. Am J Surg Pathol 2005;29:1340–1347.
Dei Tos AP, Doglioni C, Piccinin S et al. Molecular abnormalities of the p53 pathway in dedifferentiated liposarcoma. J Pathol 1997;181:8–13.
Dei Tos AP, Doglioni C, Piccinin S et al. Coordinated expression and amplification of the MDM2, CDK4, and HMGI-C genes in atypical lipomatous tumours. J Pathol 2000;190:531–536.
Thway K, Flora R, Shah C et al. Diagnostic utility of p16, CDK4, and MDM2 as an immunohistochemical panel in distinguishing well-differentiated and dedifferentiated liposarcomas from other adipocytic tumors. Am J Surg Pathol 2012;36:462–469.
Sirvent N, Coindre JM, Maire G et al. Detection of MDM2-CDK4 amplification by fluorescence in situ hybridization in 200 paraffin-embedded tumor samples: utility in diagnosing adipocytic lesions and comparison with immunohistochemistry and real-time PCR. Am J Surg Pathol 2007;31:1476–1489.
Coindre JM, Hostein I, Maire G et al. Inflammatory malignant fibrous histiocytomas and dedifferentiated liposarcomas: histological review, genomic profile, and MDM2 and CDK4 status favour a single entity. J Pathol 2004;203:822–830.
Hollmann TJ, Hornick JL . INI1-deficient tumors: diagnostic features and molecular genetics. Am J Surg Pathol 2011;35:e47–e63.
Biegel JA, Zhou JY, Rorke LB et al. Germ-line and acquired mutations of INI1 in atypical teratoid and rhabdoid tumors. Cancer Res 1999;59:74–79.
Hoot AC, Russo P, Judkins AR et al. Immunohistochemical analysis of hSNF5/INI1 distinguishes renal and extra-renal malignant rhabdoid tumors from other pediatric soft tissue tumors. Am J Surg Pathol 2004;28:1485–1491.
Judkins AR, Mauger J, Ht A et al. Immunohistochemical analysis of hSNF5/INI1 in pediatric CNS neoplasms. Am J Surg Pathol 2004;28:644–650.
Chbani L, Guillou L, Terrier P et al. Epithelioid sarcoma: a clinicopathologic and immunohistochemical analysis of 106 cases from the French sarcoma group. Am J Clin Pathol 2009;131:222–227.
Hornick JL, Dal Cin P, Fletcher CD . Loss of INI1 expression is characteristic of both conventional and proximal-type epithelioid sarcoma. Am J Surg Pathol 2009;33:542–550.
Orrock JM, Abbott JJ, Gibson LE et al. INI1 and GLUT-1 expression in epithelioid sarcoma and its cutaneous neoplastic and nonneoplastic mimics. Am J Dermatopathol 2009;31:152–156.
Sullivan LM, Folpe AL, Pawel BR et al. Epithelioid sarcoma is associated with a high percentage of SMARCB1 deletions. Mod Pathol 2013;26:385–392.
Calderaro J, Moroch J, Pierron G et al. SMARCB1/INI1 inactivation in renal medullary carcinoma. Histopathology 2012;61:428–435.
Cheng JX, Tretiakova M, Gong C et al. Renal medullary carcinoma: rhabdoid features and the absence of INI1 expression as markers of aggressive behavior. Mod Pathol 2008;21:647–652.
Barletta JA, Hornick JL . Succinate dehydrogenase-deficient tumors: diagnostic advances and clinical implications. Adv Anat Pathol 2012;19:193–203.
van Nederveen FH, Gaal J, Favier J et al. An immunohistochemical procedure to detect patients with paraganglioma and phaeochromocytoma with germline SDHB, SDHC, or SDHD gene mutations: a retrospective and prospective analysis. Lancet Oncol 2009;10:764–771.
Doyle LA, Nelson D, Heinrich MC et al. Loss of succinate dehydrogenase subunit B (SDHB) expression is limited to a distinctive subset of gastric wild-type gastrointestinal stromal tumours: a comprehensive genotype-phenotype correlation study. Histopathology 2012;61:801–809.
Janeway KA, Kim SY, Lodish M et al. Defects in succinate dehydrogenase in gastrointestinal stromal tumors lacking KIT and PDGFRA mutations. Proc Natl Acad Sci USA 2011;108:314–318.
Miettinen M, Wang ZF, Sarlomo-Rikala M et al. Succinate dehydrogenase-deficient GISTs: a clinicopathologic, immunohistochemical, and molecular genetic study of 66 gastric GISTs with predilection to young age. Am J Surg Pathol 2011;35:1712–1721.
Gaal J, Stratakis CA, Carney JA et al. SDHB immunohistochemistry: a useful tool in the diagnosis of Carney-Stratakis and Carney triad gastrointestinal stromal tumors. Mod Pathol 2011;24:147–151.
Gill AJ, Chou A, Vilain R et al. Immunohistochemistry for SDHB divides gastrointestinal stromal tumors (GISTs) into 2 distinct types. Am J Surg Pathol 2010;34:636–644.
Rege TA, Wagner AJ, Corless CL et al. ‘Pediatric-type’ gastrointestinal stromal tumors in adults: distinctive histology predicts genotype and clinical behavior. Am J Surg Pathol 2011;35:495–504.
Dwight T, Benn DE, Clarkson A et al. Loss of SDHA expression identifies SDHA mutations in succinate dehydrogenase-deficient gastrointestinal stromal tumors. Am J Surg Pathol 2013;37:226–233.
Miettinen M, Killian JK, Wang ZF et al. Immunohistochemical loss of succinate dehydrogenase subunit A (SDHA) in gastrointestinal stromal tumors (GISTs) signals SDHA germline mutation. Am J Surg Pathol 2013;37:234–240.
Oudijk L, Gaal J, Korpershoek E et al. SDHA mutations in adult and pediatric wild-type gastrointestinal stromal tumors. Mod Pathol 2013;26:456–463.
Wagner AJ, Remillard SP, Zhang YX et al. Loss of expression of SDHA predicts SDHA mutations in gastrointestinal stromal tumors. Mod Pathol 2013;26:289–294.
Gill AJ, Pachter NS, Chou A et al. Renal tumors associated with germline SDHB mutation show distinctive morphology. Am J Surg Pathol 2011;35:1578–1585.
Miettinen M, Sarlomo-Rikala M, Cue PM et al. Mapping of succinate dehydrogenase losses in 2258 epithelial neoplasms. Appl Immunohistochem Mol Morphol advance online publication, 25 March 2013 (e-pub ahead of print).
Ladanyi M, Lui MY, Antonescu CR et al. The der(17)t(X;17)(p11;q25) of human alveolar soft part sarcoma fuses the TFE3 transcription factor gene to ASPL, a novel gene at 17q25. Oncogene 2001;20:48–57.
Argani P, Antonescu CR, Illei PB et al. Primary renal neoplasms with the ASPL-TFE3 gene fusion of alveolar soft part sarcoma: a distinctive tumor entity previously included among renal cell carcinomas of children and adolescents. Am J Pathol 2001;159:179–192.
Argani P, Aulmann S, Illei PB et al. A distinctive subset of PEComas harbors TFE3 gene fusions. Am J Surg Pathol 2010;34:1395–1406.
Antonescu CR, Le Loarer F, Mosquera JM et al. Novel YAP1-TFE3 fusion defines a distinct subset of epithelioid hemangioendothelioma. Genes Chromosomes Cancer 2013;52:775–784.
Argani P, Lal P, Hutchinson B et al. Aberrant nuclear immunoreactivity for TFE3 in neoplasms with TFE3 gene fusions: a sensitive and specific immunohistochemical assay. Am J Surg Pathol 2003;27:750–761.
Folpe AL, Hill CE, Parham DM et al. Immunohistochemical detection of FLI-1 protein expression: a study of 132 round cell tumors with emphasis on CD99-positive mimics of Ewing's sarcoma/primitive neuroectodermal tumor. Am J Surg Pathol 2000;24:1657–1662.
Wang WL, Patel NR, Caragea M et al. Expression of ERG, an Ets family transcription factor, identifies ERG-rearranged Ewing sarcoma. Mod Pathol 2012;25:1378–1383.
Cessna MH, Zhou H, Sanger WG et al. Expression of ALK1 and p80 in inflammatory myofibroblastic tumor and its mesenchymal mimics: a study of 135 cases. Mod Pathol 2002;15:931–938.
Chan JK, Cheuk W, Shimizu M . Anaplastic lymphoma kinase expression in inflammatory pseudotumors. Am J Surg Pathol 2001;25:761–768.
Coffin CM, Hornick JL, Fletcher CD . Inflammatory myofibroblastic tumor: comparison of clinicopathologic, histologic, and immunohistochemical features including ALK expression in atypical and aggressive cases. Am J Surg Pathol 2007;31:509–520.
Coffin CM, Patel A, Perkins S et al. ALK1 and p80 expression and chromosomal rearrangements involving 2p23 in inflammatory myofibroblastic tumor. Mod Pathol 2001;14:569–576.
Cook JR, Dehner LP, Collins MH et al. Anaplastic lymphoma kinase (ALK) expression in the inflammatory myofibroblastic tumor: a comparative immunohistochemical study. Am J Surg Pathol 2001;25:1364–1371.
Marino-Enriquez A, Wang WL, Roy A et al. Epithelioid inflammatory myofibroblastic sarcoma: An aggressive intra-abdominal variant of inflammatory myofibroblastic tumor with nuclear membrane or perinuclear ALK. Am J Surg Pathol 2011;35:135–144.
Yoshida A, Shibata T, Wakai S et al. Anaplastic lymphoma kinase status in rhabdomyosarcomas. Mod Pathol 2013;26:772–781.
Sholl LM, Weremowicz S, Gray SW et al. Combined use of ALK immunohistochemistry and FISH for optimal detection of ALK-rearranged lung adenocarcinomas. J Thorac Oncol 2013;8:322–328.
Conklin CM, Craddock KJ, Have C et al. Immunohistochemistry is a reliable screening tool for identification of ALK rearrangement in non-small-cell lung carcinoma and is antibody dependent. J Thorac Oncol 2013;8:45–51.
Chmielecki J, Crago AM, Rosenberg M et al. Whole-exome sequencing identifies a recurrent NAB2-STAT6 fusion in solitary fibrous tumors. Nat Genet 2013;45:131–132.
Mohajeri A, Tayebwa J, Collin A et al. Comprehensive genetic analysis identifies a pathognomonic NAB2/STAT6 fusion gene, nonrandom secondary genomic imbalances, and a characteristic gene expression profile in solitary fibrous tumor. Genes Chromosomes Cancer 2013;52:873–886.
Robinson DR, Wu YM, Kalyana-Sundaram S et al. Identification of recurrent NAB2-STAT6 gene fusions in solitary fibrous tumor by integrative sequencing. Nat Genet 2013;45:180–185.
Schweizer L, Koelsche C, Sahm F et al. Meningeal hemangiopericytoma and solitary fibrous tumors carry the NAB2-STAT6 fusion and can be diagnosed by nuclear expression of STAT6 protein. Acta Neuropathol 2013;125:651–658.
Doyle LA, Vivero M, Fletcher CD et al. Nuclear expression of STAT6 distinguishes solitary fibrous tumor from histologic mimics. Mod Pathol advance online publication, 13 September 2013; doi:10.1038/modpathol.2013.164 (e-pub ahead of print).
West RB, Corless CL, Chen X et al. The novel marker, DOG1, is expressed ubiquitously in gastrointestinal stromal tumors irrespective of KIT or PDGFRA mutation status. Am J Pathol 2004;165:107–113.
Gomez-Pinilla PJ, Gibbons SJ, Bardsley MR et al. Ano1 is a selective marker of interstitial cells of Cajal in the human and mouse gastrointestinal tract. Am J Physiol Gastrointest Liver Physiol 2009;296:G1370–G1381.
Espinosa I, Lee CH, Kim MK et al. A novel monoclonal antibody against DOG1 is a sensitive and specific marker for gastrointestinal stromal tumors. Am J Surg Pathol 2008;32:210–218.
Lee CH, Liang CW, Espinosa I . The utility of discovered on gastrointestinal stromal tumor 1 (DOG1) antibody in surgical pathology-the GIST of it. Adv Anat Pathol 2010;17:222–232.
Liegl B, Hornick JL, Corless CL et al. Monoclonal antibody DOG1.1 shows higher sensitivity than KIT in the diagnosis of gastrointestinal stromal tumors, including unusual subtypes. Am J Surg Pathol 2009;33:437–446.
Miettinen M, Wang ZF, Lasota J . DOG1 antibody in the differential diagnosis of gastrointestinal stromal tumors: a study of 1840 cases. Am J Surg Pathol 2009;33:1401–1408.
Terry J, Saito T, Subramanian S et al. TLE1 as a diagnostic immunohistochemical marker for synovial sarcoma emerging from gene expression profiling studies. Am J Surg Pathol 2007;31:240–246.
Foo WC, Cruise MW, Wick MR et al. Immunohistochemical staining for TLE1 distinguishes synovial sarcoma from histologic mimics. Am J Clin Pathol 2011;135:839–844.
Jagdis A, Rubin BP, Tubbs RR et al. Prospective evaluation of TLE1 as a diagnostic immunohistochemical marker in synovial sarcoma. Am J Surg Pathol 2009;33:1743–1751.
Knosel T, Heretsch S, Altendorf-Hofmann A et al. TLE1 is a robust diagnostic biomarker for synovial sarcomas and correlates with t(X;18): analysis of 319 cases. Eur J Cancer 2010;46:1170–1176.
Kosemehmetoglu K, Vrana JA, Folpe AL . TLE1 expression is not specific for synovial sarcoma: a whole section study of 163 soft tissue and bone neoplasms. Mod Pathol 2009;22:872–878.
Moller E, Hornick JL, Magnusson L et al. FUS-CREB3L2/L1-positive sarcomas show a specific gene expression profile with upregulation of CD24 and FOXL1. Clin Cancer Res 2011;17:2646–2656.
Guillou L, Benhattar J, Gengler C et al. Translocation-positive low-grade fibromyxoid sarcoma: clinicopathologic and molecular analysis of a series expanding the morphologic spectrum and suggesting potential relationship to sclerosing epithelioid fibrosarcoma: a study from the French Sarcoma Group. Am J Surg Pathol 2007;31:1387–1402.
Mertens F, Fletcher CD, Antonescu CR et al. Clinicopathologic and molecular genetic characterization of low-grade fibromyxoid sarcoma, and cloning of a novel FUS/CREB3L1 fusion gene. Lab Invest 2005;85:408–415.
Doyle LA, Moller E, Dal Cin P et al. MUC4 is a highly sensitive and specific marker for low-grade fibromyxoid sarcoma. Am J Surg Pathol 2011;35:733–741.
Doyle LA, Wang WL, Dal Cin P et al. MUC4 is a sensitive and extremely useful marker for sclerosing epithelioid fibrosarcoma: association with FUS gene rearrangement. Am J Surg Pathol 2012;36:1444–1451.
Tirabosco R, Berisha F, Ye H et al. Assessment of MUC4 expression in primary bone tumours. Histopathology 2013;63:142–143.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no conflict of interest.
Rights and permissions
About this article
Cite this article
Hornick, J. Novel uses of immunohistochemistry in the diagnosis and classification of soft tissue tumors. Mod Pathol 27 (Suppl 1), S47–S63 (2014). https://doi.org/10.1038/modpathol.2013.177
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2013.177
Keywords
This article is cited by
-
FET(EWSR1)-TFCP2 Rhabdomyosarcoma: An Additional Example of this Aggressive Variant with Predilection for the Gnathic Bones
Head and Neck Pathology (2021)
-
Aggressive Intraosseous Myofibroma of the Maxilla: Report of a Rare Case and Literature Review
Head and Neck Pathology (2021)
-
The pathology of soft tissue sarcomas
La radiologia medica (2019)
-
Ewing sarcoma
Nature Reviews Disease Primers (2018)