Abstract
The high mobility group A (HMGA2) gene encodes a protein that alters chromatin structure and regulates the transcription of many genes; it is implicated in both benign and malignant neoplasias, but its rearrangements are a feature of development of several mesenchymal tumors. Given its implication in these tumors and particularly adipocytic tumors, and the availability of antibodies usable on paraffin-embedded tissues, we evaluated the immunohistochemical expression of this gene in a series of 1052 mesenchymal tumors. The objective was to define the value and limitations of HMGA2 immunohistochemical expression for histotyping, and compare with molecular data reported in the literature. We thus analyzed 880 cases on tissue microarray and 182 cases on whole sections (211 adipocytic tumors, 628 sarcomas, 213 benign mesenchymal tumors, and 10 normal adipose tissues). A nuclear immunostaining was detected in 86% of conventional and intramuscular lipomas, in 86% of well-differentiated liposarcomas and in 67% of dedifferentiated liposarcomas, as opposed to 16% of other benign adipose tumors and to 15% of non-well-differentiated liposarcoma/dedifferentiated liposarcoma sarcomas. Among benign mesenchymal tumors and lesions, it was detected in 90% of nodular fasciitis and in 88% of benign fibrous histiocytomas with respective specificities of 85 and 100%, and in 90% of aggressive angiomyxoma, contrary to other vulvovaginal tumor types, which expressed HMGA2 only rarely. The normal adipose tissue was always negative for HMGA2. Although not specific, immunohistochemical detection of the HMGA2 protein is helpful for the distinction of normal adipose tissue from well-differentiated lesions, particularly on biopsy or on re-excision. It is less sensitive than MDM2/CDK4 for dedifferentiated liposarcomas diagnosis, but it appears more specific to distinguish dedifferentiated liposarcomas from other poorly differentiated sarcomas. Finally, and may be more importantly, HMGA2 is useful for the diagnosis of benign fibrous histiocytoma, nodular fasciitis and vulvovaginal benign mesenchymal tumors.
Similar content being viewed by others
Main
The human high mobility group A (HMGA2) gene encodes a non-histone chromatin protein that belongs to a family of the HMG proteins including also HMGA1a, HMGA1b, HMGA1c, which result from alternative splicing of the HMGA1 mRNA.1, 2 HMGA2 can modify chromatin structure enabling a fine tuning of transcription in a variety of cellular processes such as differentiation and oncogenesis.3, 4, 5, 6 With a length of 160 kb, it is located at the chromosome band 12q14.3 and consists of five exons. Each of the first three exons codes for an AT hook domain, which is separated from the fourth exon by a large intronic sequence (140 kB), whereas the fifth exon codes for the acidic C-terminal domain. The 3′UTR of HMGA2 contains binding sites for the let-7 family of microRNAs that inhibit gene expression at the posttranscriptional level.7, 8 HMGA2 is at high levels during embryonic development,9, 10 but it is not detectable at any significant level in adult tissues.11, 12 Recent studies report increased expression of HMGA2 in several human malignancies, including lung and pancreatic tumors, oral squamous cell carcinomas and a subset of breast cancers.13, 14, 15, 16, 17, 18 This expression represents a poor prognostic index as overexpression often correlates with the presence of metastasis. This supports the idea of the implication of HMGA2 in pathogenesis, although its function is not yet well understood in these neoplasms.4 However, this gene is also a target of chromosomal rearrangements in some benign solid tumors19, 20 such as mesenchymal tumors:21, 22, 23, 24 in particular, HMGA2 is a prominent feature of conventional lipomas as 75% of these harbor translocation t(3;12) involving it,3, 25, 26 but it is also overexpressed or amplified in atypical lipomatous tumor/well-differentiated27, 28, 29 and dedifferentiated liposarcomas. Given the involvement of HMGA2 in mesenchymal tumors and particularly adipocytic tumors and the availability of antibodies usable on paraffin-embedded tissues, we evaluated immunohistochemical expression of this gene in a large series of mesenchymal tumors in order to define its value and limitations for histotyping.
Materials and methods
We selected 1052 mesenchymal tumors from 1014 patients. Cases were issued from the archives of the Department of Pathology of Institut Bergonié and from the database of the French Sarcoma Group (http://www.conticabase.org) with the participation of Paul Papin Center, Angers; University hospital, Caen; Henri Mondor Hospital, Créteil; Georges-Francois Leclerc Center, Dijon; University Institute of Pathology, Lausanne; Oscard Lambret Center, Lille; Leon-Berard Center, Lyon; Val d’Aurelle Center, Montpellier; Alexis Vautrin Center, Nancy; René Huguenin Center, Saint-Cloud; Claudius Regaud Institute, Toulouse and Gustave Roussy Institute, Villejuif. These cases have been collegially reviewed and classified according to the latest World Health Organization classification for bone and soft tissue tumors.
The tumors selected belong to four main categories: (1) adipocytic tumors with 218 cases including 73 benign tumors (31 conventional lipomas, 5 intramuscular lipomas, 34 spindle cell and pleomorphic lipomas, 2 hibernomas and 1 myolipoma) and 145 liposarcomas (43 atypical lipomatous tumor/well-differentiated, 69 dedifferentiated liposarcomas, 19 pleomorphic liposarcomas and 14 myxoid/round cell liposarcomas); (2) non-adipocytic sarcomas with 628 cases including 217 leiomyosarcomas, 179 unclassified sarcomas, 84 myxofibrosarcomas, 35 rhabdomyosarcomas (18 pleomorphic, 11 embryonal and 6 alveolar), 14 synovial sarcomas, 13 gastro-intestinal stromal tumors, 12 malignant peripheral nerve sheath tumors, 10 angiosarcomas, 10 dermatofibrosarcoma protuberans, 9 osteosarcomas, 8 primitive neuroectodermal tumors, 6 epithelioid sarcomas, 6 clear cell sarcomas, 5 soft part alveolar sarcomas, 4 extra-skeletal myxoid chondrosarcomas, 4 Kaposi's sarcomas, 3 epithelioid hemangioendotheliomas, 3 low-grade fibromyxoid sarcomas, 3 desmoplastic small round cell tumors, 2 malignant mesenchymomas, 1 malignant solitary fibrous tumor; (3) benign mesenchymal tumors or lesions with 176 cases including 81 desmoid tumors, 20 nodular fasciitis, 17 benign fibrous histiocytomas, 13 uterine leiomyomas, 10 intramuscular myxomas, 5 angiomas, 5 angiolipomas, 5 schwannomas, 3 neurofibromas, 3 solitary fibrous tumors, 3 myofibromas, 2 angiomyolipomas, 2 perineuriomas, 1 angioleiomyoma, 1 granular cell tumor, 1 giant cell tumor of soft tissue and 4 reactive fibrosis; and (4) benign mesenchymal vulvovaginal tumors with 30 cases including 11 fibroepithelial stromal polyps, 10 aggressive angiomyxomas, 4 angiomyofibroblastomas, 2 cellular angiofibromas and 3 leiomyomas. We also analyzed 10 samples of normal adipose tissue.
Eight hundred and eighty mesenchymal tumors were analyzed on tissue microarray, divided into 18 paraffin blocks. Each case was represented by three spots of 1 mm. The other 182 cases were analyzed on whole sections. In particular, this concerned the 10 samples of normal adipose tissues, the benign adipocytic tumors (excluding spindle cell lipomas) and atypical lipomatous tumor/well-differentiated because of their properties with tissue unsuitable for tissue microarray. To this we added some samples of benign tumors including some cases of fasciitis, or benign fibrous histiocytomas, solitary fibrous tumors, mesenchymal vulvovaginal tumors and leiomyomas to supplement the number of cases treated by this method to improve the representativeness of the results.
Immunohistochemistry
A total of 4 μm-thick paraffin sections were cut and mounted on glass slides (Superfrost+ Menzel Glazer). Subsequently, the preparations were entirely treated on automate VENTANA-Benchmark-XT®. The primary antibody was a polyclonal anti-rabbit antibody from BIOCHECK laboratories (Foster City, Ref. 59170AP) and was used at a 1:500 dilution. Sections were incubated at room temperature in standard CCI buffer (Ventana) and were revealed with ‘Ultra ViewTr’ Universal DAB kit, after that they stained with Hematoxylin ReaDi Ventana solution (Ref. 760-2021).
A tumor was considered as HMGA2 positive when at least one-cell nuclei stained per high-power field. The number of positive cells was counted in medium-power fields for each histology section and quantified as a percentage. For each case derived from tissue microarray, we used the average value of the three spots.
Statistical Analysis
Specificity and sensitivity for HMGA2 immunostainings were evaluated in the well-differentiated liposarcoma subgroup and compared with that of the normal adipose tissue, and evaluated in the well-differentiated/dedifferentiated liposarcoma and compared with that of the so-called ‘others sarcoma’ type subgroup. Sensitivity for HMGA2 was calculated as the ratio of positively stained atypical lipomatous tumor/well-differentiated/dedifferentiated liposarcomas to the number of atypical lipomatous tumor/well-differentiated/dedifferentiated liposarcomas examined. Specificity for HMGA2 was calculated as the ratio of unstained non-atypical lipomatous tumor/well-differentiated/dedifferentiated liposarcomas tumors to the total number of non-atypical lipomatous tumor/well-differentiated/dedifferentiated liposarcomas. Among the benign tumors or the vulvovaginal area, sensitivities and specificities were evaluated on the same principle with the aggressive angiomyxomas as the referent positive tumor. According to cytogenetic data, we hypothesized that conventional lipomas, atypical lipomatous tumor/well-differentiated and dedifferentiated liposarcomas would be positive as opposed to normal adipose tissue and to others sarcomas.
Results
HMGA2 staining according to tumor histotype is summarized in Tables 1, 2 and 3. Among benign adipocytic tumors (Table 1), HMGA2 was expressed in 86% of conventional and intramuscular lipomas, whereas other tumors were rarely positive (six cases of spindle cell and pleomorphic lipomas, no case of hibernoma or myolipoma, ie 16% of cases). Among liposarcomas (Table 1), HMGA2 was expressed almost exclusively by atypical lipomatous tumor/well-differentiated and dedifferentiated liposarcomas (86 and 67% of cases, respectively), with only two positive cases among pleomorphic liposarcomas and no cases for myxoid/round cell liposarcomas (6%).
As opposed to 67% positivity in dedifferentiated liposarcomas, only 16% of potential dedifferentiated liposarcomas simulators were positive, especially in myxofibrosarcomas (26%), and embryonal rhabdomyosarcomas (36%). Only 5 out of 78 (8%) of other sarcomas were positive (Table 2).
Among non-adipocytic benign mesenchymal tumors, HMGA2 was expressed in 88% of benign fibrous histiocytomas, in 90% of nodular fasciitis and in 30% of leiomyomas. Among vulvovaginal mesenchymal tumors, we obtained a positive staining in 90% of aggressive angiomyxoma, contrary to other tumor types, which expressed HMGA2 rarely (Table 3). In our study, none of the 10 cases of normal adipose tissue was positive for HMGA2.
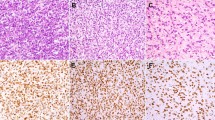
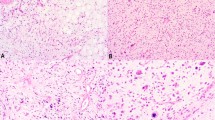
HMGA2 shows a nuclear positivity with a variable number of positive cells: in 128 cases (48% of positive cases), at least 50% of tumor cells were immunostained for HMGA2, especially in lipomas, atypical lipomatous tumor/well-differentiated liposarcoma and dedifferentiated liposarcomas (Figures 1, 2 and 3).
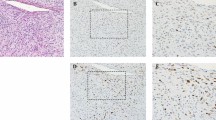
Regarding atypical lipomatous tumor/well-differentiated, we analyzed expression according to histologic subtype: 86% of atypical lipomatous tumor/well-differentiated lipoma-like subtypes expressed HMGA2 with only 4% of cases showing >50% of positive cells. Eighty-three percent of atypical lipomatous tumor/well-differentiated sclerosing subtypes expressed HMGA2 with half of them showing >50% of labeled cells. In other HMGA2-positive sarcomas, between 5 and 30% of the cells were stained. Finally, among nodular fasciitis and benign fibrous histiocytomas, which frequently expressed HMGA2 (Figures 4 and 5), only three cases showed <50% of positive cells. The proportion of positive cells in leiomyomas varies widely between 10 and 100% of cells. Among vulvovaginal tumors, 90% of aggressive angiomyxomas expressed HMGA2 with an intense nuclear positivity involving at least 75% of cells in all cases (Figure 6). Nearly 30% of fibroepithelial stromal polyps expressed this antibody, but the immunostaining was less intense in fewer cells. Table 4 summarizes the statistical analysis regarding sensitivity and specificity of HMGA2.
Discussion
This study of immunohistochemical expression of HMGA2 in a large series of mesenchymal tumors showed a strong positivity in most conventional and intramuscular lipomas and in atypical lipomatous tumor/well-differentiated. These results are in agreement with our cytogenetics knowledge as such tumors show rearrangements in the 12q13–15 region in which the HMGA2 gene is the target. Translocation (3;12) is the most common cytogenetic aberration in conventional lipomas, and leads to a loss of C-terminal region and the formation of a fusion gene HMGA2-LPP. HMGA2 is certainly the important gene of this rearrangement in lipomas3, 4 as there are no fewer than 41 other chromosomal loci as partners of HMGA2.3, 30, 31 However, conventional lipomas with normal karyotypes31 or with aberrations in 6p21 or on chromosome 13q have been reported. These data could explain that in our series, roughly 14% of conventional lipomas do not express HMGA2. Moreover, although the amplification of MDM2 and CDK4 genes is considered to be the major event in atypical lipomatous tumor/well-differentiated/dedifferentiated liposarcomas, overexpression of HMGA2 in these tumors (86 and 67%, respectively) has been suggested a possible function of HMGA2 in atypical lipomatous tumor/well-differentiated.32
HMGA2 does not allow us to distinguish a lipoma from an atypical lipomatous tumor/well-differentiated. Although it is not more sensitive than MDM2/CDK4 for diagnosing dedifferentiated liposarcomas, with a positivity of 67% of cases collected, HMGA2 can be useful for the diagnosis of dedifferentiated liposarcomas, especially on biopsy, or for distinguishing areas of atypical lipomatous tumor/well-differentiated from normal adipose tissue. However, we should keep in mind that the gold standard for the diagnosis of difficult cases of atypical lipomatous tumor/well-differentiated and dedifferentiated liposarcomas is currently the demonstration of MDM2/CDK4 amplification by FISH.33
In our study, the most frequent potential simulators of dedifferentiated liposarcomas, which are HMGA2 positive are myxofibrosarcomas, poorly differentiated sarcomas and leiomyosarcomas with a positivity of 26, 18 and 13%, respectively. Therefore, in an individual case, HMGA2 expression is of rather limited value in the differential diagnosis of dedifferentiated liposarcoma.
Our results are similar to those reported by Bartuma,27 showing that the expression levels of HMGA2 evaluated by RT–PCR were correlated with morphological and cytogenetic subgroups of adipocytic tumors. These data suggest a practical interest of HMGA2 immunohistochemistry for the diagnosis in three situations: for distinguishing normal adipocytic tissue from a mature adipocytic tumor when dealing with a small quantity of tissue; for evaluating the margin status in atypical lipomatous tumor/well-differentiated and for separating dedifferentiated liposarcomas from simulators.
This study also showed that HMGA2 immunohistochemistry can be useful for the diagnosis of some benign mesenchymal lesions. It is positive in almost 90% of benign fibrous histiocytoma and negative in dermatofibrosarcoma protuberans. This result is in agreement with a recent study, which reported that only 1 case out of 14 dermatofibrosarcoma protuberans focally overexpressed HMGA2 and that 96% of benign fibrous histiocytoma expressed HMGA2 intensely and diffusely; the difference was statistically significant.34 In our study, 90% of nodular fasciitis, also overexpressed HMGA2. In contrast, only 16% of desmoid tumors were positive. Therefore, this marker can be useful when dealing with nodular fasciitis. HMGA2 gene has a function in the development and differentiation of soft tissues35, 36 and its dysregulated expression has been reported in benign mesenchymal lesions.37, 38 Given the physiological function of HMGA2, its overexpression can be the cause of cellular proliferation. On the other hand, HMGA2 expression can be induced by growth factors such as platelet-derived growth factor39 and, therefore, could be the consequence rather than the cause of cellular proliferation, particularly in reactive lesions such as nodular fasciitis. As nodular fasciitis and cellular fibrous histiocytoma are easily confused, this overexpression of HMGA2 in both lesions may make the differential diagnosis more difficult.
Among vulvovaginal benign mesenchymal tumors, aggressive angiomyxoma also overexpressed HMGA2 in 90% of cases. However, fibroepithelial stromal polyps and leiomyomas were positive in 27 and 33% of our cases.
This HMGA2 overexpression may certainly be explained by the complex cytogenetic rearrangements of HMGA2 in benign lesions. The most frequent cytogenetic abnormalities in leiomyomas (especially in uterus) are on the one hand a reciprocal translocation t(12;14) (q15; q23–24) with an extragenic breakpoint at the 5′part of HMGA2 and on the other hand, a deletion of the long arm of chromosome 7.23, 40 But FISH showed complex rearrangements on chromosomes 3, 6, 10 and 12.41, 42 Cytogenetic abnormalities of HMGA2 are reported in 40–50% of uterine leiomyomas. These data are significantly correlated with immunohistochemical expression of HMGA2 in our series and in several other studies.42, 43, 44 Among vulvovaginal lesions, aggressive angiomyxomas also have rearrangements of 12q13–15 region with translocations involving chromosomes 1, 7, 8 and 21.45, 46, 47, 48 But HMGA2 was identified as the only regulated gene, and RT–PCR techniques have reported breakpoints in HMGA2 at different levels: preferentially in the 5′ part of gene and also described in intron 3. More widely, a study of the characterization of rearrangements of HMGA2 in 90 genital tumors showed their presence in 33% of aggressive angiomyxomas, but not in other tumors (18 angiomyofibroblastomas, 6 cellular angiofibromas, 5 fibroepithelial stromal polyps and 5 leiomyomas).49 The resulting immunohistochemical application presents a clear interest in practice. Although the correlation between rearrangements of the HMGA2 protein and its expression is not clearly established in these lesions, the literature reports an overexpression in about 50% of cases of aggressive angiomyxoma.50 Our results corroborated these data with an intense and diffuse positivity in 9 out of 10 cases of aggressive angiomyxoma as opposed to other differential diagnoses such as cellular angiofibroma and angiomyofibroblastoma, which are always negative for this marker. However, HMGA2 positivity is not specific as a third of fibroepithelial stromal polyps and leiomyomas cases of this location express this protein; but in these cases, there are other morphologic criteria for diagnosis, except when dealing with an extensively myxoid leiomyoma, which is very similar to aggressive angiomyxoma and, therefore, represents one of the most difficult differential diagnosis for this tumor. This marker is also potentially interesting for assessing surgical margins of atypical lipomatous tumor/well-differentiated as this is challenging at the histological level.
In conclusion, although not specific of a line of differentiation, immunohistochemical detection of HMGA2 is a potential valuable aid in the diagnosis of mesenchymal tumors. In particular, it is useful for distinguishing well-differentiated adipocytic tumors from normal tissue, for margin assessment in atypical lipomatous tumor/well-differentiated, for dedifferentiated liposarcomas and, potentially more importantly, for the diagnosis of benign fibrous histiocytoma, nodular fasciitis and vulvovaginal benign mesenchymal tumors.
References
Giancotti V, Bandiera A, Buratti E, et al. Comparison of multiple forms of the high mobility group I proteins in rodent and human cells. Identification of the human high mobility group I-C protein. Eur J Biochem 1991;198:211–216.
Johnson KR, Cook SA, Davisson MT . Chromosomal localization of the murine gene and two related sequences encoding high-mobility-group I and Y proteins. Genomics 1992;12:503–509.
Fedele M, Battista S, Manfioletti G, et al. Role of the high mobility group A proteins in human lipomas. Carcinogenesis 2001;22:1583–1591.
Fusco A, Fedele M . Roles of HMGA proteins in cancer. Nat Rev Cancer 2007;7:899–910.
Narita M, Krizhanovsky V, Nunez S, et al. A novel role for high-mobility group a proteins in cellular senescence and heterochromatin formation. Cell 2006;126:503–514.
Reeves R . Molecular biology of HMGA proteins: hubs of nuclear function. Gene 2001;277:63–81.
Lee YS, Dutta A . The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev 2007;21:1025–1030.
Mayr C, Hemann MT, Bartel DP . Disrupting the pairing between let-7 and Hmga2 enhances oncogenic transformation. Science 2007;315:1576–1579.
Chiappetta G, Avantaggiato V, Visconti R, et al. High level expression of the HMGI (Y) gene during embryonic development. Oncogene 1996;13:2439–2446.
Hirning-Folz U, Wilda M, Rippe V, et al. The expression pattern of the Hmgic gene during development. Genes Chromosomes Cancer 1998;23:350–357.
Li O, Li J, Droge P . DNA architectural factor and proto-oncogene HMGA2 regulates key developmental genes in pluripotent human embryonic stem cells. FEBS Lett 2007;581:3533–3537.
Rogalla P, Drechsler K, Frey G, et al. HMGI-C expression patterns in human tissues. Implications for the genesis of frequent mesenchymal tumors. Am J Pathol 1996;149:775–779.
Abe N, Watanabe T, Masaki T, et al. Pancreatic duct cell carcinomas express high levels of high mobility group I(Y) proteins. Cancer Res 2000;60:3117–3122.
Fabjani G, Tong D, Wolf A, et al. HMGA2 is associated with invasiveness but not a suitable marker for the detection of circulating tumor cells in breast cancer. Oncol Rep 2005;14:737–741.
Hristov AC, Cope L, Reyes MD, et al. HMGA2 protein expression correlates with lymph node metastasis and increased tumor grade in pancreatic ductal adenocarcinoma. Mod Pathol 2009;22:43–49.
Meyer B, Loeschke S, Schultze A, et al. HMGA2 overexpression in non-small cell lung cancer. Mol Carcinog 2007;46:503–511.
Motoyama K, Inoue H, Nakamura Y, et al. Clinical significance of high mobility group A2 in human gastric cancer and its relationship to let-7 microRNA family. Clin Cancer Res 2008;14:2334–2340.
Miyazawa J, Mitoro A, Kawashiri S, et al. Expression of mesenchyme-specific gene HMGA2 in squamous cell carcinomas of the oral cavity. Cancer Res 2004;64:2024–2029.
Hess JL . Chromosomal translocations in benign tumors: the HMGI proteins. Am J Clin Pathol 1998;109:251–261.
Schoenmakers EF, Geurts JM, Kools PF, et al. A 6-Mb yeast artificial chromosome contig and long-range physical map encompassing the region on chromosome 12q15 frequently rearranged in a variety of benign solid tumors. Genomics 1995;29:665–678.
Dahlen A, Mertens F, Rydholm A, et al. Fusion, disruption, and expression of HMGA2 in bone and soft tissue chondromas. Mod Pathol 2003;16:1132–1140.
Ingraham SE, Lynch RA, Surti U, et al. Identification and characterization of novel human transcripts embedded within HMGA2 in t(12;14)(q15;q24.1) uterine leiomyoma. Mutat Res 2006;602:43–53.
Mine N, Kurose K, Nagai H, et al. Gene fusion involving HMGIC is a frequent aberration in uterine leiomyomas. J Hum Genet 2001;46:408–412.
Ashar HR, Tkachenko A, Shah P, et al. HMGA2 is expressed in an allele-specific manner in human lipomas. Cancer Genet Cytogenet 2003;143:160–168.
Nilsson M, Mertens F, Hoglund M, et al. Truncation and fusion of HMGA2 in lipomas with rearrangements of 5q32-->q33 and 12q14-->q15. Cytogenet Genome Res 2006;112:60–66.
Petit MM, Schoenmakers EF, Huysmans C, et al. LHFP, a novel translocation partner gene of HMGIC in a lipoma, is a member of a new family of LHFP-like genes. Genomics 1999;57:438–441.
Bartuma H, Panagopoulos I, Collin A, et al. Expression levels of HMGA2 in adipocytic tumors correlate with morphologic and cytogenetic subgroups. Mol Cancer 2009;8:36.
Dei Tos AP, Doglioni C, Piccinin S, et al. Coordinated expression and amplification of the MDM2, CDK4, and HMGI-C genes in atypical lipomatous tumours. J Pathol 2000;190:531–536.
Tallini G, Dal Cin P, Rhoden KJ, et al. Expression of HMGI-C and HMGI(Y) in ordinary lipoma and atypical lipomatous tumors: immunohistochemical reactivity correlates with karyotypic alterations. Am J Pathol 1997;151:37–43.
Arlotta P, Tai AK, Manfioletti G, et al. Transgenic mice expressing a truncated form of the high mobility group I-C protein develop adiposity and an abnormally high prevalence of lipomas. J Biol Chem 2000;275:14394–14400.
Bartuma H, Hallor KH, Panagopoulos I, et al. Assessment of the clinical and molecular impact of different cytogenetic subgroups in a series of 272 lipomas with abnormal karyotype. Genes Chromosomes Cancer 2007;46:594–606.
Italiano A, Bianchini L, Keslair F, et al. HMGA2 is the partner of MDM2 in well-differentiated and dedifferentiated liposarcomas whereas CDK4 belongs to a distinct inconsistent amplicon. Int J Cancer 2008;122:2233–2241.
Sirvent N, Coindre JM, Maire G, et al. Detection of MDM2-CDK4 amplification by fluorescence in situ hybridization in 200 paraffin-embedded tumor samples utility in diagnosing adipocytic lesions and comparison with immunohistochemistry and real-time PCR. Am J Surg Pathol 2007;31:1476–1489.
Li N, McNiff J, Hui P, et al. Differential expression of HMGA1 and HMGA2 in dermatofibroma and dermatofibrosarcoma protuberans: potential diagnostic applications, and comparison with histologic findings, CD34, and factor XIIIa immunoreactivity. Am J Dermatopathol 2004;26:267–272.
Zhou X, Benson KF, Ashar HR, et al. Mutation responsible for the mouse pygmy phenotype in the developmentally regulated factor HMGI-C. Nature 1995;376:771–774.
Battista S, Fidanza V, Fedele M, et al. The expression of truncated HMGI-C gene induces gigantism associated with lipomatosis. Cancer Res 1999;59:4793–4797.
Schoenmakers EF, Wanschura S, Mols R, et al. Recurrent rearrangements in the high mobility group protein gene, HMGI-C, in benign mesenchymal tumors. Nat Genet 1995;10:436–444.
Tallini G, Dal Cin P . HMGI(Y) and HMGI-C dysregulation: a common occurrence in human tumors. Adv Anat Pathol 1999;6:237–246.
Ayoubi TA, Jansen E, Meulemans SM, et al. Regulation of HMGIC expression : an architectural transcription factor involved in growth control and development. Oncogene 1999;18:5076–5087.
Kazmierczak B, Pohnke Y, Bullerdiek J . Fusion transcripts between the HMGIC gene and RTVL-H-related sequences in mesenchymal tumors without cytogenetic aberrations. Genomics 1996;38:223–226.
Nilbert M, Heim S, Mandahl N, et al. Characteristic chromosome abnormalities, including rearrangements of 6p, del(7q), +12, and t(12;14), in 44 uterine leiomyomas. Hum Genet 1990;85:605–611.
Rein MS, Friedman AJ, Barbieri RL, et al. Cytogenetic abnormalities in uterine leiomyomata. Obstet Gynecol 1991;77:923–926.
Gross KL, Neskey DM, Manchanda N, et al. HMGA2 expression in uterine leiomyomata and myometrium: quantitative analysis and tissue culture studies. Genes Chromosomes Cancer 2003;38:68–79.
Peng Y, Laser J, Shi G, et al. Antiproliferative effects by Let-7 repression of high-mobility group A2 in uterine leiomyoma. Mol Cancer Res 2008;6:663–673.
Horsman D, Berean K . Aggressive angiomyxoma of the pelvis: cytogenetic findings in a single case. Cancer Genet and Cytogenet. [Abstract] 1991;56:130.
Betz J, Meloni A, Moore G . Cytogenetics findings in a case of angiomyxoma of the vaginal wall. Cancer Genet Cytogenet 1995 Abstract 84:157.
Kazmierczak B, Dal Cin P, Wanschura S . Involvement of the MAR genes in two rares benign mesenchymal tumors: a hamartoma of the breast and a aggressive angiomyxoma. Cancer Genet Cytogenet. [Abstract] 1995;84:148.
Nucci MR, Weremowicz S, Neskey DM, et al. Chromosomal translocation t(8;12) Induces aberrant HMGIC expression in aggressive angiomyxoma of the vulva. Genes Chromosomes Cancer 2001;32:172–176.
Medeiros F, Erickson-Johnson MR, Keeney GL, et al. Frequency and characterization of HMGA2 and HMGA1 rearrangements in mesenchymal tumors of the lower genital tract. Genes Chromosomes Cancer 2007;46:981–990.
Medeiros F, Oliveira A, Llyod R . HMGA2 expression was a biomarker for aggressive angiomyxoma. Mod Pathol. [Abstract]. 2008;21:214A.
Acknowledgements
We are grateful to the following pathologists for contributing cases and paraffin blocks: A Carbonelle, Créteil; MC Chateau, Montpellier; B Chetaille, Marseille; F Collin, Dijon; L Guillou, Lausanne; A Leroux, Nancy; B Marques, Toulouse; JJ Michels, Caen; F Mishellany, Clermont-Ferrand; YM Robin, Lille; M Trassard, Saint-Cloud; I Valo, Angers and to Pippa McKelvie-Sebileau for help with the English manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Dreux, N., Marty, M., Chibon, F. et al. Value and limitation of immunohistochemical expression of HMGA2 in mesenchymal tumors: about a series of 1052 cases. Mod Pathol 23, 1657–1666 (2010). https://doi.org/10.1038/modpathol.2010.174
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2010.174
Keywords
This article is cited by
-
Retrospective analysis of clinicopathological features and prognosis for aggressive angiomyxoma of 27 cases in a tertiary center: a 14-year survey and related literature review
Archives of Gynecology and Obstetrics (2020)
-
HMGA2 immunostaining is a straightforward technique which helps to distinguish pulmonary fat-forming lesions from normal adipose tissue in small biopsies: a retrospective observational study about a series of 13 lung biopsies
Diagnostic Pathology (2017)