Abstract
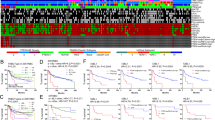
Microarray profiling of invasive breast carcinomas has identified five distinct subtypes of tumors (luminal A, luminal B, normal breast-like, HER2 overexpressing, and basal-like) that are associated with different clinical outcomes. The basal-like subtype is associated with poor clinical outcomes and is the subtype observed in BRCA1-related breast cancers. The aim of this study was to characterize the histologic and immunophenotypic properties of breast basal-like carcinomas that were first positively identified using DNA microarray analysis. Detailed histologic review was performed on 56 tumors with known microarray profiles (23 basal-like, 23 luminal, and 12 HER2+). Immunohistochemistry for estrogen receptor (ER), HER2, EGFR, smooth muscle actin (SMA), p63, CD10, cytokeratin 5/6, cytokeratin 8/18, and vimentin was performed on 18 basal-like, 16 luminal, and 12 HER2+ tumors. The basal-like tumors were grade 3 ductal/NOS (21/23) or metaplastic (2/23) carcinomas that frequently showed geographic necrosis (17/23), a pushing border of invasion (14/23), and a stromal lymphocytic response (13/23). Most basal-like tumors showed immunoreactivity for vimentin (17/18), luminal cytokeratin 8/18 (15/18), EGFR (13/18), and cytokeratin 5/6 (11/18), while positivity for the myoepithelial markers SMA (4/18), p63 (4/18) and CD10 (2/18) was infrequent. All basal-like tumors tested were ER− and HER2−. Morphologic features significantly associated with the basal-like subtype included markedly elevated mitotic count (P<0.0001), geographic tumor necrosis (P=0.0003), pushing margin of invasion (P=0.0001), and stromal lymphocytic response (P=0.01). The most consistent immunophenotype seen in the basal-like tumors was negativity for ER and HER2, and positivity for vimentin, EGFR, cytokeratin 8/18, and cytokeratin 5/6. The infrequent expression of myoepithelial markers in basal-like carcinomas does not support a direct myoepithelial cell derivation of these tumors. These findings should further assist in the identification of basal-like carcinomas in clinical specimens, facilitating treatment and epidemiologic studies of this tumor subtype.
Similar content being viewed by others
Main
Variations in transcriptional programs account for much of the biological diversity of human cells and tumors. Despite this molecular diversity, analyses of invasive breast carcinomas using microarrays have identified gene expression signatures that characterize many of the essential qualities important for biological and clinical classification.1, 2, 3, 4, 5, 6, 7, 8, 9, 10 DNA microarray profiling studies on breast tumors show distinct and reproducible subtypes of breast carcinoma associated with different outcomes. Expression profiles have categorized invasive breast carcinomas into five groups: luminal A, luminal B, HER2+/estrogen receptor (ER)−, basal-like, and normal breast-like. The basal-like subtype is typically ER− and HER2− (ie not amplified) and shows some characteristics of breast myoepithelial cells. The basal-like subtype has been shown to have the highest proliferation rates and poor clinical outcomes,2, 3 and has also been described in association with BRCA1-associated carcinomas.11 Overall, little is known with regard to the development and prevention of these aggressive tumors.
Normal breast ducts contain at least three types of epithelial cells: luminal (glandular) cells, basal/myoepithelial cells, and stem cells. Myoepithelial and luminal epithelia can be distinguished by their different cytokeratin expression patterns. Myoepithelial cells typically express cytokeratin 5/6 and cytokeratin 17, while luminal cells typically express cytokeratins 8 and 18. Immunohistochemical studies for basal/myoepithelial and luminal cytokeratins appear to be helpful in subtyping invasive breast carcinomas into distinct biological subtypes, although the staining patterns for cytokeratins 5/6 and 17 can be highly variable.12
Studies have employed basal/myoepithelial cytokeratins and other markers to identify a subset of ER- and HER2-negative breast carcinomas that are associated with a poor prognosis, further supporting the idea that a basal-like phenotype exists.13, 14, 15, 16, 17, 18 The prevalence and poor prognosis of basal-like breast carcinomas have been validated immunohistochemically; in a 564-case tissue microarray, van de Rijn et al12 demonstrated that 16% of tumors stained positive for cytokeratin 5/6 or cytokeratin 17, and that basal cytokeratin expression was associated with a poor prognosis. Abd El-Rehim evaluated 1944 cases of invasive breast carcinoma and found that approximately 18% of tumors show basal cytokeratin immunoreactivity, and again, these tumors showed a poor prognosis.14 Tsuda et al17, 18 described myoepithelial differentiation in a group of high-grade invasive ductal carcinomas showing a large, central acellular zone. These tumors showed aggressive behavior with increased risk of brain and lung metastasis. Basal cytokeratins are not expressed in all tumors classified as basal-like by gene microarray analysis, which demonstrates a need to identify other markers of basal-like subtype.
An association between EGFR expression and the basal-like phenotype has been demonstrated.16 Breast cancers showing immunoreactivity for cytokeratin 5/6 were also frequently found to coexpress EGFR. Nielsen et al13 proposed a panel of four antibodies (ER, EGFR, HER2, and cytokeratin 5/6) to identify basal-like tumors in which the basal-like tumors are identified as those tumors that were ER−, HER2 not amplified, and showed expression of cytokeratin 5/6 and/or EGFR. It is unknown whether other basal/myoepithelial markers such as smooth muscle actin (SMA), p63, or CD10 are expressed in the basal-like subtype of breast carcinoma. One aim of this study is to evaluate immunohistochemical markers that may further facilitate identification of basal-like tumors in formalin-fixed, paraffin-embedded tissue.
The morphologic features of breast tumors identified as basal-like by gene expression profile have not been well described. It is unclear whether these tumors are similar or heterogeneous in their histology and if there are any supportive morphologic clues to identify this subtype by light microscopy. Therefore, the second aim of this study was to evaluate the grade and morphologic features of a group of known basal-like invasive breast carcinomas identified by gene microarray analysis.
Materials and methods
After IRB approval at our institution, 23 basal-like, 23 luminal/ER+, and 12 HER2+/ER− subtype invasive breast carcinomas were identified by DNA microarray analysis as described previously.2, 3, 19 In this paper, the tumor subtypes were identified from three different microarray data sets. In each data set separately, tumors were subtyped using microarray expression profiles and hierarchical clustering analysis of the samples using a breast tumor intrinsic gene set (see Sorlie et al3 for a detailed description). Each sample was then classified as being basal-like, luminal, normal-like, or HER2+/ER−. In all, 18 basal-like, 16 luminal, and 12 HER2+/ER− tumors had formalin-fixed, paraffin-embedded tissue available for immunohistochemical studies.
Immunohistochemistry
Sections (5 μm) from formalin-fixed, paraffin-embedded tumors were cut and mounted onto Probe On Plus slides (Fisher Scientific). Following deparaffinization in xylene, slides were rehydrated through a graded series of alcohol and placed in running water. Endogenous peroxidase activity was blocked with 3% hydrogen peroxidase and methanol. Samples were steamed for antigen retrieval with 10 mM citrate buffer (pH 6.0) for 30 min. Following protein block, slides were incubated with antibody for ER, HER2, EGFR, SMA, p63, CD10, cytokeratin 8/18, and vimentin (Table 1), and incubated with streptavidin-conjugated HRP using Vectastain ABC kit protocol (Vector Laboratories). 3,3′-Diaminobenzidine tetrahydrochloride (DAB) was used for the visualization of the antibody/enzyme complex. Slides were counterstained with hematoxylin (Biomeda-M10) and examined by light microscopy. Tumor immunoreactivity was scored 0=negative, 1=weak positive, and 2=moderate/strong positive in combination with the percent of cells showing positive staining.
Histopathology
Hematoxylin-and-eosin-stained slides from 23 basal-like, 23 luminal/ER+, and 12 HER2+/ER− invasive breast carcinomas identified by gene microarray analysis were evaluated under light microscopy. The array subtype was blinded at the time of histologic review. Tumors were evaluated for grade (1–3) using the modified Scarff–Bloom–Richardson grading system comprising an architectural grade, nuclear grade, and mitotic grade. Mitotic counts were performed using the × 40 objective on an Olympus BX51 microscope. Tumors were also evaluated for tumor histologic type, presence of geographic necrosis, quantity of apoptotic tumor cells, border appearance, lymphocytic stromal response, nucleoli, nuclear chromatin pattern, apocrine features, metaplastic features, large central acellular zone, and medullary features. The histopathologic features of basal-like, luminal/ER+, and HER2+/ER− tumors were compared.
Statistical Methods
Fisher's exact test was used for data categorized into 2 × 2 contingency tables. The nonparametric Jonckheere–Terpstra method was used to test for ordered differences over categories of interest (such as tumor array subtype by associated histologic features or expression of tested immunohistochemical markers). With this test, the null hypothesis is that the distribution of the response does not differ across the ordered categories. Statistical analyses were performed with SAS statistical software, Version 9.1, SAS Institute Inc., Cary, NC, USA.
Results
Light Microscopic Findings
A detailed morphological analysis of known basal-like tumors has not been described; therefore, we performed a morphologic evaluation of 23 basal-like carcinomas identified by microarray analysis. All basal-like tumors showed an overall modified Scarff–Bloom–Richardson grade of 3 (Table 2). The majority of tumors showed solid architecture with no tubule formation and a high density of tumor cells with scant intervening stroma between tumor cells. In all, 14/23 (61%) tumors showed a pushing border and 13/23 (56%) had some degree of a stromal lymphocytic infiltrate at the tumor edge. Four tumors demonstrated histologic features similar to those described for atypical medullary carcinoma (Figure 1a), including a predominantly pushing border, syncytial arrangement of tumor cells, and a marked stromal lymphocytic infiltrate at the tumor edge. Also, 5/23 (22%) tumors showed regions of ribbon-like architecture with associated central necrosis (Figure 1b). Most of the tumor cells contained scant cytoplasm and round to oval nuclei producing a high nuclear/cytoplasmic ratio. The nuclear chromatin pattern ranged from coarse to vesicular. Nucleoli were present in 20/23 (87%) cases and ranged from inconspicuous to prominent. A high mitotic rate was identified in all tumors, ranging from 26 mitoses/10 high-power fields to 92 mitoses/10 high-power fields (average 45 mitoses/10 high-power fields). Geographic zones of necrosis were identified in 17/23 (74%) (Figure 1c) tumors. Apoptotic tumor cells were numerous in each of the cases with geographic necrosis. Two tumors showed metaplastic components, including squamous metaplasia (Figure 1d) and matrix production. None of the tumors showed appreciable apocrine differentiation or a large, central acellular zone.
Morphologic features of basal-like carcinomas. (a) Some tumors showed medullary features, including solid architecture with a pushing margin and brisk stromal lymphocytic response. (b) Other tumors showed characteristic ribbon-like architecture associated with central necrosis. (c) Foci of geographic tumor necrosis were a frequent finding in these tumors. (d) One of the two metaplastic carcinomas showed areas of squamous metaplasia. Original magnification × 200 (a–c) and × 400 (d).
Histologic differences were identified among the basal, luminal, and HER2+/ER− subtypes. The rate of grade 3 histology seen in the basal-like tumors (23/23, 100%) was significantly higher than that observed in the luminal tumors (2/23, 9%), P<0.0001. When the histologic features of all tumor subtypes were compared, markedly elevated (>25/10 high-power fields) mitotic index (P<0.001), geographic tumor necrosis (P=0.0003), pushing border of invasion (P=0.0001), stromal lymphocytic response (P=0.01), and atypical medullary features (P=0.03) were all associated with the basal-like subtype. The basal-like and luminal tumors were dissimilar in nearly all evaluated histologic features. There was overlap in the histology of basal-like and HER2+/ER− tumors (Table 2); however, metaplastic and medullary tumor features were limited to the basal-like subtype.
Immunohistochemical Findings
Immunohistochemical stains for ER, HER2, EGFR, SMA, p63, CD10, cytokeratin 8/18, vimentin, and cytokeratin 5/6 were performed on 18 basal-like, 16 luminal, and 12 HER2+/ER− invasive breast carcinomas identified by gene microarray analysis (Table 3). The basal-like tumors were ER− (0/18, 0%) and HER2− (0/18, 0%), while the luminal subtype tumors were ER+ (18/18, 100%) and HER2− (0/18, 0%), and the HER2+/ER− subtype tumors were ER− (0/12, 0%) and HER2 overexpressed (11/12, 92%). The rate of EGFR expression seen in the basal-like (13/18, 72%) tumors was significantly higher than that observed in luminal tumors (0/23 0%), P<0.0001. Most HER2+/ER− (7/12, 58%) subtypes also expressed EGFR; thus, in this study, EGFR expression was seen exclusively in the basal-like and HER2+/ER− subtypes.
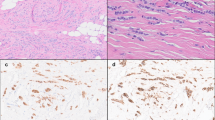
Immunoreactivity for myoepithelial markers within the group of basal-like tumors was highly variable (Table 3). In all, 4/18 (22%) of the basal-like tumors showed immunoreactivity for SMA (Figure 2a), with tumor cell immunoreactivity being distinguished from the adjacent myofibroblast immunoreactivity. SMA expression, although infrequent, was associated with the basal-like subtype (P=0.02). None of the luminal or HER2+/ER− tumors showed immunoreactivity for SMA. Also, 4/18 basal-like tumors showed immunoreactivity for p63 (Figure 2b). Immunoreactivity was generally weak to moderate and present in <5% of the tumor cells. Similar weak positivity for p63 was also seen in 2/16 (12%) of luminal tumors and in 4/12 (33%) of HER2+/ER− tumors. Further, 2/18 (11%) basal-like tumors showed weak immunoreactivity for CD10, while CD10 expression was also seen in 1/12 (8%) HER2+/ER− tumors and 0/16 (0%) luminal tumors, and 11/18 (61%) basal-like tumors showed immunoreactivity for cytokeratin 5/6. Cytokeratin 5/6 expression in tumors was significantly associated with the basal-like subtype, P=0.0001. None of the luminal tumors showed cytokeratin 5/6 expression and 1/12 HER2+/ER− tumors showed weak cytokeratin 5/6 expression. Most tumors, regardless of subtype, demonstrated expression of luminal cytokeratin 8/18, including the basal-like tumors, which showed expression of cytokeratin 8/18 in 15 of 18 (83%) cases (Figure 2c). Strong cytokeratin 8/18 expression was identified in all luminal and HER2+/ER− subtype tumors. In total, 11/18 (61%) basal-like tumors showed coexpression of both basal and luminal cytokeratins, which suggests that many of these tumors show characteristics of both luminal and basal/myoepithelial cell lineages.
Immunophenotype of basal-like carcinomas. (a) SMA and (b) p63 positivity was present in a minority of tumors, with immunoreactivity being detected in only a small percentage of tumor cells. (c) Most tumors showed strong luminal cytokeratin 8/18 expression throughout the tumor. (e) Vimentin positivity was characterized by strong and diffuse immunoreactivity. Original magnification × 400.
Vimentin expression in tumors was strongly associated with a basal-like phenotype, P<0.0001, being present in 17/18 (94%) basal-like tumors. The positive tumors showed strong, near-diffuse immunoreactivity (Figure 2d). In contrast, 1/16 (6%) luminal tumors and 1/12 (8%) HER2+/ER− tumors showed weak vimentin expression. The expression pattern seen in these two tumors was significantly more focal and weaker than that seen in the basal-like tumors, suggesting that strong vimentin staining could be useful for the identification of basal-like tumors in breast cancers. The overall protein expression patterns seen in basal-like carcinomas were heterogeneous. The most common immunophenotype for the basal-like tumors was ER−, HER2−, vimentin positive, EGFR+, cytokeratin 5/6 positive, and cytokeratin 8/18 positive (Table 4). Strong vimentin and cytokeratin 5/6 expressions were the most specific markers for the basal-like subtype.
Discussion
A significant amount of data exists concerning the histological characteristics of ER− tumors and the immunohistochemical patterns of expression of nonluminal cell markers (like SMA, p63, vimentin) in breast carcinomas. The precise relationship between these features and basal-like tumors, however, is not known because the only reliable method to identify basal-like tumors is by microarray analysis. Therefore, we positively identified a set of basal-like tumors using microarray analysis and then performed a histological and immunohistochemical review of their properties.
The morphologic appearance of basal-like tumors was variable, but some consistent and unique histologic features were identified when these tumors were compared to luminal and HER2+/ER− tumors. Features strongly associated with the basal-like subtype included markedly elevated mitotic rate, geographic tumor necrosis, pushing margin of invasion, atypical medullary features, and a stromal lymphocytic response. The basal-like tumors contained scant cytoplasm producing a high nuclear/cytoplasmic ratio in which nuclei of adjacent cells touched or overlapped. The basal-like tumors were remarkably solid tumors with little intervening stroma between tumor cells, producing a densely cellular lesion. The high mitotic index observed in these tumors is consistent with their uniformly high expression of the ‘proliferation cluster’ of genes shown in DNA microarray studies.1, 2, 3 Apocrine differentiation, which is frequently associated with ER-negative status, was not identified in these tumors.
This study demonstrated that there is overlap in the histologic features of basal-like and HER2+/ER− tumors. Both subtypes share histologic features that have been previously reported as characteristic of ER− breast carcinomas.20 These features include grade 3 histology and geographic necrosis. Although there are some histologic features that appear to be distinctive of basal-like carcinomas, immunohistochemical studies are clearly needed for subtyping of high-grade breast carcinomas. Additionally, the morphologic and immunophenotypic features observed in the basal-like carcinomas (grade 3 histology, ER/HER2-negative status, and medullary features) are similar to those reported for BRCA1-related carcinomas.21, 22 These features may eventually play a role in risk assessment for BRCA1 mutation screening. The patient BRCA1 mutation status was not directly addressed in this study, but all the tumors evaluated were sporadic in nature, and it is likely that most of the basal-like tumors were not associated with a BRCA1 mutation.
Basal-like breast carcinomas have been characterized by gene expression analysis as having a gene expression pattern similar to basal/myoepithelial cells found in normal breast tissue (1–3), and patterns similar to squamous cell carcinomas of the lung.19 Normal breast myoepithelial cells show immunoreactivity for basal cytokeratins (ie cytokeratins 5, 6, and 17), SMA, CD10, vimentin, and p63. Studies have shown basal cytokeratin expression in a subset of invasive breast cancers,12, 14 and it is presumed that most of these tumors are basal-like tumors; however, these studies did not formally demonstrate this relationship and other classic myoepithelial markers were typically not examined.
All of the 18 known basal-like carcinomas in this study showed an incomplete basal/myoepithelial cell phenotype. Many, but not all, basal-like tumors showed expression of cytokeratin 5/6 and EGFR; however, the majority of basal-like tumors were negative for myoepithelial markers, including SMA, CD10, and p63. SMA immunoreactivity, when present, was only seen in basal-like carcinomas, but it was infrequent even among this subtype (4/18, 22%). Immunoreactivity for p63 was seen in 22% of basal-like tumors, but the degree of positivity was only weak to moderate and typically involved <5% of tumor cells. It should be noted that p63 expression was also identified in 2/16 (12%) luminal breast tumors and 4/12 (33%) HER2+/ER− tumors, and the immunoreactivity patterns were similar to the basal-like tumors. SMA, p63, and CD10, therefore, appear to be of limited value in the immunohistochemical detection of basal-like tumors.
Tumor basal cytokeratin expression was specific for the basal-like subtype in this study, but 7/18 (39%) of the basal-like tumors were negative for cytokeratin 5/6 protein expression, as was shown before.12, 13 This finding indicates a need for additional immunohistochemical markers to identify all basal-like tumors, and a candidate identified in this study may be vimentin, which was expressed in 17/18 basal-like tumors. This appeared to be a distinguishing feature from luminal and HER2+/ER− subtypes, suggesting that vimentin could be added to the current panel of antibodies (ER, EGFR, HER2, and cytokeratin 5/6) used to identify basal-like tumors; however, the precise combinations and scoring criteria may need to be re-evaluated and validated.
Vimentin expression in breast carcinomas may also have prognostic value and an association with an aggressive tumor phenotype.23, 24, 25, 26, 27 Previous studies have shown an association between tumor vimentin expression, a poor prognosis, and ER-negative status.23, 24 Cattoretti et al23 described a group of ER-negative tumors that often showed coexpression of vimentin, p53 and EGFR, which is a phenotype that is very consistent with the basal-like subtype currently described. We have shown here and before that many basal-like tumors are EGFR+, and Sorlie et al2 showed that most basal-like tumors are p53 mutated. Domagala et al demonstrated an inverse relation between vimentin expression and survival for node-negative breast cancers (P<0.0001).
The origin of vimentin expression in invasive basal-like carcinomas is unclear. The possible mechanisms include direct derivation of these tumors from myoepithelial cells, epithelial-to-mesenchymal transition as a result of dedifferentiation, and derivation of these tumors from stem cells. The heterogeneous expression pattern and frequent lack of expression of myoepithelial markers in these tumors do not support a direct derivation of these tumors from myoepithelial cells. Additional evidence against a myoepithelial cell origin is the cytokeratin 8/18 expression pattern seen in basal-like tumors. Cytokeratin 8/18 is a marker typically expressed in the luminal epithelial cells of the breast and is not expressed in normal myoepithelial cells. Strong expression of luminal cytokeratin 8/18 was observed in most basal-like tumors (83%) in this study; 10/18 basal-like tumors also showed coexpression of both basal/myoepithelial and luminal cytokeratins, indicative of a mixed basal/luminal phenotype, and all of these tumors were ER-negative. The coexpression of basal/myoepithelial and luminal cytokeratins raises the possibility that these tumors may arise from stem cells that subsequently undergo variable degrees of basal and luminal differentiation. Studies of the precursor lesions for these tumors, including hyperplasias and in-situ carcinomas, will be important in further evaluating this possibility. These data should assist in the identification of basal-like tumors in formalin-fixed, paraffin-embedded clinical specimens, which will be needed for future studies aimed at evaluating clinical outcomes, treatment responses, and the epidemiology of this group of tumors.
References
Perou CM, Sorlie T, Eisen MB, et al. Molecular portraits of human breast tumours. Nature 2000;406:747–752.
Sorlie T, Perou CM, Tibshirani R, et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 2001;98:10869–10874.
Sorlie T, Tibshirani R, Parker J, et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci USA 2003;100:8418–8423.
Sotiriou C, Powles TJ, Dowsett M, et al. Gene expression profiles derived from fine needle aspiration correlate with response to systemic chemotherapy in breast cancer. Breast Cancer Res 2002;4:R3.
van 't Veer LJ, Dai H, van de Vijver MJ, et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 2002;415:530–536.
van de Vijver MJ, He YD, van't Veer LJ, et al. A gene expression signature as a predictor of survival in breast cancer. N Engl J Med 2002;347:1999–2009.
Ma XJ, Wang Z, Ryan PD, et al. A two-gene expression ratio predicts clinical outcome in breast cancer patients treated with tamoxifen. Cancer Cell 2004;5:607–616.
Huang E, Cheng SH, Dressman H, et al. Gene expression predictors of breast cancer outcomes. Lancet 2003;361:1590–1596.
Sotiriou C, Neo SY, McShane LM, et al. Breast cancer classification and prognosis based on gene expression profiles from a population based study. Proc Natl Acad Sci USA 2003;100:10393–10398.
Perou CM, Jeffrey SS, van de Rijn M, et al. Distinctive gene expression patterns in human mammary epithelial cells and breast cancers. Proc Natl Acad Sci USA 1999;96:9212–9217.
Foulkes WD, Stefansson IM, Chappuis PO, et al. Germline BRCA1 mutations and a basal epithelial phenotype in breast cancer. J Natl Cancer Inst 2003;95:1482–1485.
van de Rijn M, Perou CM, Tibshirani R, et al. Expression of cytokeratins 17 and 5 identifies a group of breast carcinomas with poor clinical outcome. Am J Pathol 2002;161:1991–1996.
Nielsen TO, Hsu FD, Jensen K, et al. Immunohistochemical and clinical characterization of the basal-like subtype of invasive breast carcinoma. Clin Cancer Res 2004;10:5367–5374.
Abd El-Rehim DM, Pinder SE, Paish CE, et al. Expression of luminal and basal cytokeratins in human breast carcinoma. J Pathol 2004;203:661–671.
Kesse-Adu R, Shousha S . Myoepithelial markers are expressed in at least 29% of oestrogen receptor negative invasive breast carcinoma. Mod Pathol 2004;17:646–652.
Korsching E, Packeisen J, Agelopoulous K, et al. Cytogenetic alterations and cytokeratin expression patterns in breast cancer: integrating a new model of breast differentiation into cytogenetic pathways of breast carcinogenesis. Lab Invest 2002;82:1525–1533.
Tsuda H, Takarabe T, Hasegawa T, et al. Myoepithelial differentiation in high-grade invasive ductal carcinomas with large central acellular zones. Hum Pathol 1999;10:1134–1139.
Tsuda H, Takarabe T, Hasegawa F, et al. Large, central acellular zones indicating myoepithelial tumor differentiaton in high-grade invasive ductal carcinomas as markers of predisposition to lung and brain metastases. Am J Surg Pathol 2000;24:197–202.
Chung CH, Bernard PS, Perou CM . Molecular portraits and the family tree of cancer. Nat Genet 2002;32:533–540.
Putti TC, Abd El-Rehim DM, Rakha EA, et al. Estrogen receptor-negative breast carcinomas: a review of morphology and immunophenotypical analysis. Mod Pathol 2005;18:26–35.
Chang J, Hilsenbeck SG, Sng JH, et al. Pathologic features and BRCA1 mutation screening in premenopausal breast cancer patients. Clin Cancer Res 2001;7:1739–1742.
Quenneville LA, Phillips KA, Ozcelik H, et al. HER2/neu status and tumor morphology of invasive breast carcinomas in Ashkenazi women with known BRCA1 mutation status in the Ontario Familial Breast Cancer Registry. Cancer 2002;95:2068–2075.
Cattoretti G, Andreola S, Clemente C, et al. Vimentin and p53 expression in epidermal growth factor receptor-positive, oestrogen receptor-negative breast carcinomas. Br J Cancer 1988;57:353–357.
Domagala W, Lasota J, Dukowicz A, et al. Vimentin expression appears to be associated with poor prognosis in node-negative ductal NOS breast carcinomas. Am J Pathol 1990;137:1299–1304.
Sommers CL, Walker-Jones D, Heckford SE, et al. Vimentin rather than keratin expression in some hormone-independent breast cancer cell lines and in oncogene-transformed mammary epithelial cells. Cancer Res 1989;49:4258–4263.
Hendrix MJ, Seftor EA, Seftor RE, et al. Experimental co-expression of vimentin and keratin intermediate filaments in human breast cancer cells results in phenotypic interconversion and increased invasive behavior. Am J Pathol 1997;150:483–495.
Dandachi N, Hauser-Kronberger C, More E, et al. Co-expression of tenascin-C and vimentin in human breast cancer cells indicates phenotypic transdifferentiation during tumour progression: correlation with histopathological parameters, hormone receptors, and oncoproteins. J Pathol 2001;193:181–189.
Acknowledgements
This work was supported by an award to the University of North Carolina for a Breast Cancer Specialized Program of Research Excellence (SPORE) from the National Cancer Institute (CA58223), and by the NCI (RO1-CA-101227-01) to CMP.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Livasy, C., Karaca, G., Nanda, R. et al. Phenotypic evaluation of the basal-like subtype of invasive breast carcinoma. Mod Pathol 19, 264–271 (2006). https://doi.org/10.1038/modpathol.3800528
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3800528
Keywords
This article is cited by
-
Endosomal protein expression of γ1-adaptin is associated with tumor growth activity and relapse-free survival in breast cancer
Breast Cancer (2024)
-
Loss of function of GATA3 regulates FRA1 and c-FOS to activate EMT and promote mammary tumorigenesis and metastasis
Cell Death & Disease (2023)
-
FBXL2 promotes E47 protein instability to inhibit breast cancer stemness and paclitaxel resistance
Oncogene (2023)
-
Triple-negatives Mammakarzinom
Die Pathologie (2023)
-
Recent advances in therapeutic strategies for triple-negative breast cancer
Journal of Hematology & Oncology (2022)