Abstract
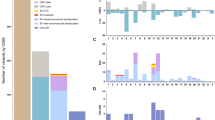
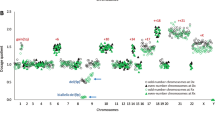
Progress in the management of patients with myelodysplastic syndromes (MDS) has been hampered by the inability to detect cytogenetic abnormalities in 40–60% of cases. We prospectively analyzed matched pairs of bone marrow and buccal cell (normal) DNA samples from 51 MDS patients by single nucleotide polymorphism (SNP) arrays, and identified somatically acquired clonal genomic abnormalities in 21 patients (41%). Among the 33 patients with normal bone marrow cell karyotypes, 5 (15%) had clonal, somatically acquired aberrations by SNP array analysis, including 4 with segmental uniparental disomies (UPD) and 1 with three separate microdeletions. Each abnormality was detected more readily in CD34+ cells than in unselected bone marrow cells. Paired analysis of bone marrow and buccal cell DNA from each patient was necessary to distinguish true clonal genomic abnormalities from inherited copy number variations and regions with apparent loss of heterozygosity. UPDs affecting chromosome 7q were identified in two patients who had a rapidly deteriorating clinical course despite a low-risk International Prognostic Scoring System score. Further studies of larger numbers of patients will be needed to determine whether 7q UPD detected by SNP array analysis will identify higher risk MDS patients at diagnosis, analogous to those with 7q cytogenetic abnormalities.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Corey SJ, Minden MD, Barber DL, Kantarjian H, Wang JC, Schimmer AD . Myelodysplastic syndromes: the complexity of stem-cell diseases. Nat Rev Cancer 2007; 7: 118–129.
Estey E . Acute myeloid leukemia and myelodysplastic syndromes in older patients. J Clin Oncol 2007; 25: 1908–1915.
Nimer SD . Myelodysplastic syndromes. Blood 2008; 111: 4841–4851.
de Witte T, Oosterveld M, Muus P . Autologous and allogeneic stem cell transplantation for myelodysplastic syndrome. Blood Rev 2007; 21: 49–59.
Haase D, Germing U, Schanz J, Pfeilstocker M, Nosslinger T, Hildebrandt B et al. New insights into the prognostic impact of the karyotype in MDS and correlation with subtypes: evidence from a core dataset of 2124 patients. Blood 2007; 110: 4385–4395.
List A, Kurtin S, Roe DJ, Buresh A, Mahadevan D, Fuchs D et al. Efficacy of lenalidomide in myelodysplastic syndromes. N Engl J Med 2005; 352: 549–557.
Ebert BL, Pretz J, Bosco J, Chang CY, Tamayo P, Galili N et al. Identification of RPS14 as a 5q- syndrome gene by RNA interference screen. Nature 2008; 451: 335–339.
Kantarjian H, Oki Y, Garcia-Manero G, Huang X, O'Brien S, Cortes J et al. Results of a randomized study of 3 schedules of low-dose decitabine in higher-risk myelodysplastic syndrome and chronic myelomonocytic leukemia. Blood 2007; 109: 52–57.
Silverman LR, Demakos EP, Peterson BL, Kornblith AB, Holland JC, Odchimar-Reissig R et al. Randomized controlled trial of azacitidine in patients with the myelodysplastic syndrome: a study of the cancer and leukemia group B. J Clin Oncol 2002; 20: 2429–2440.
Heinrichs S, Look AT . Identification of structural aberrations in cancer by SNP array analysis. Genome Biol 2007; 8: 219.
Gondek LP, Dunbar AJ, Szpurka H, McDevitt MA, Maciejewski JP . SNP array karyotyping allows for the detection of uniparental disomy and cryptic chromosomal abnormalities in MDS/MPD-U and MPD. PLoS ONE 2007; 2: e1225.
Gondek LP, Haddad AS, O'Keefe CL, Tiu R, Wlodarski MW, Sekeres MA et al. Detection of cryptic chromosomal lesions including acquired segmental uniparental disomy in advanced and low-risk myelodysplastic syndromes. Exp Hematol 2007; 35: 1728–1738.
Gondek LP, Tiu R, Haddad AS, O'Keefe CL, Sekeres MA, Theil KS et al. Single nucleotide polymorphism arrays complement metaphase cytogenetics in detection of new chromosomal lesions in MDS. Leukemia 2007; 21: 2058–2061.
Gondek LP, Tiu R, O'Keefe CL, Sekeres MA, Theil KS, Maciejewski JP . Chromosomal lesions and uniparental disomy detected by SNP arrays in MDS, MDS/MPD, and MDS-derived AML. Blood 2008; 111: 1534–1542.
Mohamedali A, Gaken J, Twine NA, Ingram W, Westwood N, Lea NC et al. Prevalence and prognostic significance of allelic imbalance by single-nucleotide polymorphism analysis in low-risk myelodysplastic syndromes. Blood 2007; 110: 3365–3373.
Bennett JM, Catovsky D, Daniel MT, Flandrin G, Galton DA, Gralnick HR et al. Proposals for the classification of the myelodysplastic syndromes. Br J Haematol 1982; 51: 189–199.
Greenberg P, Cox C, LeBeau MM, Fenaux P, Morel P, Sanz G et al. International scoring system for evaluating prognosis in myelodysplastic syndromes. Blood 1997; 89: 2079–2088.
Harris NL, Jaffe ES, Diebold J, Flandrin G, Muller-Hermelink HK, Vardiman J et al. World Health Organization classification of neoplastic diseases of the hematopoietic and lymphoid tissues: report of the Clinical Advisory Committee meeting-Airlie House, Virginia, November 1997. J Clin Oncol 1999; 17: 3835–3849.
Lin M, Wei LJ, Sellers WR, Lieberfarb M, Wong WH, Li C . dChipSNP: significance curve and clustering of SNP-array-based loss-of-heterozygosity data. Bioinformatics 2004; 20: 1233–1240.
Frazer KA, Ballinger DG, Cox DR, Hinds DA, Stuve LL, Gibbs RA et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 2007; 449: 851–861.
Liu TX, Becker MW, Jelinek J, Wu WS, Deng M, Mikhalkevich N et al. Chromosome 5q deletion and epigenetic suppression of the gene encoding alpha-catenin (CTNNA1) in myeloid cell transformation. Nat Med 2007; 13: 78–83.
Boultwood J, Fidler C, Strickson AJ, Watkins F, Gama S, Kearney L et al. Narrowing and genomic annotation of the commonly deleted region of the 5q- syndrome. Blood 2002; 99: 4638–4641.
Le Beau MM, Espinosa III R, Neuman WL, Stock W, Roulston D, Larson RA et al. Cytogenetic and molecular delineation of the smallest commonly deleted region of chromosome 5 in malignant myeloid diseases. Proc Natl Acad Sci USA 1993; 90: 5484–5488.
Poppe B, Vandesompele J, Schoch C, Lindvall C, Mrozek K, Bloomfield CD et al. Expression analyses identify MLL as a prominent target of 11q23 amplification and support an etiologic role for MLL gain of function in myeloid malignancies. Blood 2004; 103: 229–235.
Zatkova A, Schoch C, Speleman F, Poppe B, Mannhalter C, Fonatsch C et al. GAB2 is a novel target of 11q amplification in AML/MDS. Genes Chromosomes Cancer 2006; 45: 798–807.
McEachern MJ, Haber JE . Break-induced replication and recombinational telomere elongation in yeast. Annu Rev Biochem 2006; 75: 111–135.
Harada H, Harada Y, Niimi H, Kyo T, Kimura A, Inaba T . High incidence of somatic mutations in the AML1/RUNX1 gene in myelodysplastic syndrome and low blast percentage myeloid leukemia with myelodysplasia. Blood 2004; 103: 2316–2324.
Imai Y, Kurokawa M, Izutsu K, Hangaishi A, Takeuchi K, Maki K et al. Mutations of the AML1 gene in myelodysplastic syndrome and their functional implications in leukemogenesis. Blood 2000; 96: 3154–3160.
Conrad DF, Andrews TD, Carter NP, Hurles ME, Pritchard JK . A high-resolution survey of deletion polymorphism in the human genome. Nat Genet 2006; 38: 75–81.
Khaja R, Zhang J, MacDonald JR, He Y, Joseph-George AM, Wei J et al. Genome assembly comparison identifies structural variants in the human genome. Nat Genet 2006; 38: 1413–1418.
Redon R, Ishikawa S, Fitch KR, Feuk L, Perry GH, Andrews TD et al. Global variation in copy number in the human genome. Nature 2006; 444: 444–454.
Zhang J, Feuk L, Duggan GE, Khaja R, Scherer SW . Development of bioinformatics resources for display and analysis of copy number and other structural variants in the human genome. Cytogenet Genome Res 2006; 115: 205–214.
Knudson Jr AG . Mutation and cancer: statistical study of retinoblastoma. Proc Natl Acad Sci USA 1971; 68: 820–823.
Fitzgibbon J, Smith LL, Raghavan M, Smith ML, Debernardi S, Skoulakis S et al. Association between acquired uniparental disomy and homozygous gene mutation in acute myeloid leukemias. Cancer Res 2005; 65: 9152–9154.
Acknowledgements
We thank Martin H Nguyen and Michael Fernandez (MDACC) as well as Nikki Flores and Elena Wang (USCF) for excellent technical assistance and we are grateful to John Gilbert for editorial review and helpful discussions. We also thank Lorna Mangus for administrative support. The work was supported in part by NIH P01 Grant CA-108631 from the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Leukemia website (http://www.nature.com/leu)
Supplementary information
Rights and permissions
About this article
Cite this article
Heinrichs, S., Kulkarni, R., Bueso-Ramos, C. et al. Accurate detection of uniparental disomy and microdeletions by SNP array analysis in myelodysplastic syndromes with normal cytogenetics. Leukemia 23, 1605–1613 (2009). https://doi.org/10.1038/leu.2009.82
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2009.82
Keywords
This article is cited by
-
Clinical Applications of Chromosomal Microarray Testing in Myeloid Malignancies
Current Hematologic Malignancy Reports (2020)
-
Impact of copy neutral loss of heterozygosity and total genome aberrations on survival in myelodysplastic syndrome
Modern Pathology (2018)
-
Identify latent chromosomal aberrations relevant to myelodysplastic syndromes
Scientific Reports (2017)
-
Combined comparative genomic hybridization and single-nucleotide polymorphism array detects cryptic chromosomal lesions in both myelodysplastic syndromes and cytopenias of undetermined significance
Modern Pathology (2016)
-
Submicroscopic deletion of 5q involving tumor suppressor genes (CTNNA1, HSPA9) and copy neutral loss of heterozygosity associated with TET2 and EZH2 mutations in a case of MDS with normal chromosome and FISH results
Molecular Cytogenetics (2014)