Abstract
Follicular dendritic cell sarcoma is a rare malignant neoplasm of dendritic cell origin that is currently poorly characterized by genetic studies. To investigate whether recurrent genomic alterations may underlie the biology of follicular dendritic cell sarcoma and to identify potential contributory regions and genes, molecular inversion probe array analysis was performed on 14 independent formalin-fixed, paraffin-embedded samples. Abnormal genomic profiles were observed in 11 out of 14 (79%) cases. The majority showed extensive genomic complexity that was predominantly represented by hemizygous losses affecting multiple chromosomes. Alterations of chromosomal regions 1p (55%), 2p (55%), 3p (82%), 3q (45%), 6q (55%), 7q (73%), 8p (45%), 9p (64%), 11q (64%), 13q (91%), 14q (82%), 15q (64%), 17p (55%), 18q (64%), and 22q (55%) were recurrent across the 11 samples showing abnormal genomic profiles. Many recurrent genomic alterations in follicular dendritic cell sarcoma overlap deletions that are frequently observed across human cancers, suggesting selection, or an active role for these alterations in follicular dendritic cell sarcoma pathogenesis. In support of a tumor suppressor-driven biology, homozygous deletions involving tumor suppressor genes CDKN2A, RB1, BIRC3, and CYLD were also observed. Neither recurrent gains nor amplifications were observed. This genomic characterization provides new information regarding follicular dendritic cell sarcoma biology that may improve understanding about the underlying pathophysiology, provide better prognostication, and identify potential therapeutic markers for this rare disease.
Similar content being viewed by others
Main
Follicular dendritic cell sarcoma is a rare malignant neoplasm with diverse clinical presentation and unknown etiology. Follicular dendritic cell sarcoma arises from aberrant proliferation of follicular dendritic cells, stromal-derived cells of mesenchymal origin that reside in the germinal centers of lymph nodes and in other non-lymphoid (extranodal) tissues throughout the body, where they function to organize B-cell compartments and regulate B-cell-mediated immunity.1 Owing to their normal distribution, follicular dendritic cell sarcoma occurs most often in the lymph nodes, particularly of the head and neck region, although extranodal presentation accounts for up to a third of all cases, with diverse anatomical sites of involvement reported.2, 3, 4, 5 Epidemiologic studies indicate follicular dendritic cell sarcoma primarily affects adults, with a wide range of age at presentation observed (median age, fifth decade) and females and males equally affected.2, 3, 6 Risk factors for developing follicular dendritic cell sarcoma are largely unknown.
Diagnosis, determination of prognosis, and effective therapy for follicular dendritic cell sarcoma are challenged by the rarity and diverse clinical presentations of this disease. Diagnosis relies upon recognition of characteristic morphologic and immunohistochemical features, including expression of the follicular dendritic cell markers CD21, CD23, and CD35. Despite their relative specificity, these features may be missed or misinterpreted due to lack of disease context or biological factors such as small biopsy size, necrosis or dense lymphoid infiltrates obscuring typical morphologic features.2, 3, 7 Prognosis and choice of treatment strategy in follicular dendritic cell sarcoma vary, but typically take into account the age of the patient, staging, site(s) of involvement, tumor size, and morphologic factors. A number of recent reviews have examined prognosis, therapy and outcomes in follicular dendritic cell sarcoma.2, 4, 6, 7, 8 While surgical resection is practically universal, the use of adjuvant radiotherapy/chemotherapy, as well as associated outcomes, varies widely, and a consensus therapeutic approach for follicular dendritic cell sarcoma remains undefined. Furthermore, despite successful initial responses to treatment, local recurrences and metastases are common, and regardless of treatment strategy, ultimately approximately 10–20% of patients die of the disease.3, 6 Thus there is a need for better prognostic markers and therapeutic targets in follicular dendritic cell sarcoma.
The genetics underlying follicular dendritic cell sarcoma are poorly understood. Cytogenetic findings across 7 abnormal cases recently reviewed by Udayakumar et al9 included variable ploidy states (from hypodiploid to near-tetraploid) and mostly non-recurrent alterations, which may in part be due to technical challenges in obtaining sufficient, representative metaphase spreads from these tumor samples. On the molecular level, Go et al10 identified the BRAF V600E mutation, a potentially drug-targetable gain-of-function mutation that is variably present in other histiocytic and dendritic cell tumors, in a subset of follicular dendritic cell sarcoma tumors (18.5%, n=27). In a case report of a rare, high-grade follicular dendritic cell sarcoma tumor of the thyroid, Starr et al11 identified pathogenic loss-of-function mutations in PTEN (bi-allelic, nonsense) and TP53 (missense), and a novel missense mutation in RET using next-generation sequencing for a panel of 236 cancer genes. Griffin et al12 recently provided the most comprehensive genetic analysis of follicular dendritic cell sarcoma to date using a targeted next-generation sequencing assay to profile mutational and copy-number status across 309 cancer-related genes in 13 tumors. This analysis identified recurrent alterations of genes regulating NF-κB activation (NFKBIA and CYLD; 38% of cases), cell cycle progression (CDKN2A, RB1; 31% of cases), and immune evasion (CD274 and PDCD1LG2; 23% of cases). Of note, recurrent structural changes, which are common in many other sarcomas, were not detected.
Cytogenomic SNP microarray analysis has been widely used for genomic profiling of tumor samples, providing high-quality data for detection of copy-number alterations and loss-of-heterozygosity using limited DNA quantities that can be obtained from fresh, frozen or formalin-fixed, paraffin-embedded tissues.13 Here, we used molecular inversion probe array analysis, which is optimized for use on formalin-fixed, paraffin-embedded tissue samples,10, 11 to characterize the genomic profile of 14 archived follicular dendritic cell sarcoma tumors, including 6 samples previously studied by next-generation sequencing (5 with copy-number information).12 This analysis revealed a complex genomic landscape that is predominantly represented by hemizygous losses affecting multiple chromosomes, many of which overlap common cancer deletion regions and contain known or putative tumor suppressor genes functioning in similar cellular pathways. Homozygous deletions involving tumor suppressor genes CDKN2A, RB1, BIRC3, and CYLD were also observed. This analysis expands our knowledge of the genomic profile of follicular dendritic cell sarcoma and provides new insight into the biology of this rare disease.
Materials and methods
Patient Samples
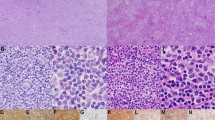
The diagnosis of follicular dendritic cell sarcoma was confirmed by experienced pathologists (JH, YK, AL, DO, and LW) based on standard WHO 2008 classification criteria.3 A total of 14 archived formalin-fixed, paraffin-embedded samples were obtained from several US institutions. Patient samples were de-identified and used in accordance with University of Utah Institutional Review Board regulations. Tumor content was estimated by pathologist review of H&E stained slides.
DNA Isolation, Processing, and Array Hybridization
Genomic DNA was extracted using the QIAamp DNA FFPE Tissue Kit (Qiagen, Valencia, CA, USA) and quantitated by Qubit PicoGreen (Life Technologies, Carlsbad, CA, USA). DNA yields varied depending on size and number of tissue slides available for extraction. The average yield was 1.5 μg (range 112 ng–6.4 μg). Approximately 80 ng genomic DNA was utilized for molecular inversion probe array analysis.
Genomic analysis was performed using the OncoScan FFPE Assay kit (Affymetrix, Santa Clara, CA, USA), using a modified protocol.14 Briefly, genomic DNA is hybridized to a series of specialized nucleic acid probes called molecular inversion probes, which target ~40 bp regions across the genome. These probes contain unique sequence identifier tags (barcodes), and are capable of circularization in order to selectively remove non-hybridized sequence (by exonuclease activity) and thereby enrich for genomic targets of interest. Following re-linearization, molecular inversion probe array sequences are selectively amplified, then digested to release the barcodes for subsequent hybridization to a high-density barcode-targeted microarray to determine both copy-number and allelic state across the genome.
The functional resolution of this assay varies based on probe density, clonal burden, and sample quality. The genome-wide limit of detection is approximately 300 kb for copy-number alterations and 3 Mb for loss-of-heterozygosity. In targeted regions containing known or putative cancer genes, copy-number alteration resolution is approximately 50–100 kb. The limit of detection for clonal alterations (mosaicism) varies based on alteration type, size, and sample quality; in general, the limit of detection for mosaicism is between 10–20%. Sequence-level alterations and balanced genomic rearrangements cannot be detected by this technology.
Data Processing and Analysis
Data processing and analysis were performed using Nexus Express Software for OncoScan 3.1 (Biodiscovery, Hawthorne, CA, USA) and with reference to human genome assembly GRCh37/hg19. All files were initially processed using the TuScan segmentation algorithm with default parameters. For two cases (samples 1 and 8), files were re-processed using the SNP-FASST2 segmentation algorithm (min. 20 probes/segment) and the copy-number reference region was manually adjusted (re-centered) in order to compensate for ploidy changes or to best fit the data to a user-defined diploid region, respectively. Sample quality was assessed using the OncoScan median absolute pairwise difference score, a global measure of variation in the log2 ratio across the genome.
All segmentation calls were manually reviewed by geneticists with experience in interpretation of molecular inversion probe array data for research and clinical purposes (EA and SS). False, inaccurate, unconvincing, or missing calls made due to poor probe performance or misinterpretation of genomic context by the software were manually edited or omitted. Non-mosaic alterations were evaluated as possibly constitutional and omitted when clinical significance was considered uncertain, likely benign or benign, or unrelated to the indication for testing. A minimum size threshold of 3 Mb was used for loss-of-heterozygosity calls. Note that for samples with a male chromosome complement, losses affecting the Y chromosome, including pseudoautosomal regions mapped to chromosome X, may be age-related and not necessarily due to the disease process in these patients.
Clonal burden was estimated using either the OS-% Aberrant Cells algorithm in Nexus or manually by averaging levels of mosaicism across calls within each sample. Level of mosaicism (% mosaicism) for each call was determined from genotype data (B-allele frequency and/or allele difference values) using calculations described by Conlin et al.15 Percent (%) mosaicism was not determined for calls in case 1 due to extensive genomic complexity, and for any high copy-number gains (calls with greater than two copies gained), unless automatically estimated by the software using the OS-% Aberrant Cells algorithm. Clonal diversity was defined when ≥3 alterations were present at distinct levels of mosaicism (≥2-fold difference) in the sample. When clonal diversity was evident, the subset of calls with higher % mosaicism was used to calculate clonal burden. Percent genome altered was determined from total % genome changed (by copy-number alterations) and % loss-of-heterozygosity, measured in Nexus.
Recurrently altered genomic regions were defined by the presence of similar overlapping copy-number and loss-of-heterozygosity calls in at least 45% (5/11) abnormal cases. Naming of genomic regions was based on the relative size (focal/non-focal) of alterations observed across the majority of abnormal cases. Regions containing ≥50% focal alterations were further subdivided into chromosome arm regions (distal, medial, proximal), otherwise whole-arm designations were used. Alterations spanning <75% of a chromosome arm were considered focal.
Smallest regions of overlap for calls within recurrently altered genomic regions were delineated manually, using genomic coordinates from individual calls to define the proximal and distal boundaries. Regions containing the most overlapping single copy loss or loss-of-heterozygosity calls across cases were used. Some cases showed multiple noncontiguous calls, which led to delineation of a broader smallest region of overlap. Homozygous deletion breakpoints were not used to delineate smallest region of overlap.
Known and putative tumor suppressor genes within recurrently altered and homozygously deleted regions were identified using several resources including: the Cancer Gene Census list (v73, accessible online at http://cancer.sanger.ac.uk/census/), which contains 571 mutated genes that are causally implicated in human cancer,16 with selection for genes with annotated loss-of-function type mutations (deletions, nonsense, frameshift mutations); the Mut-driver genes list, which contains 138 high-confidence cancer drivers (74 total tumor suppressor genes) identified from analysis of mutations in 18 000 genes within the COSMIC database17 using the ‘20/20 rule’, whereby genes are classified as tumor suppressor genes when >20% of recorded mutations are inactivating/truncating, and including recurrent homozygously deleted genes;18 and the TUSON Explorer genes list, a manually curated list of 1000 cancer-associated genes predicted to have tumor suppressor gene-like signatures by a computational method that incorporates mutation types, positions and relative proportions of different mutation classes to predict gene function, using genes ranked with manual confidence scores of 3 or 4 (208 genes).19
Pan-cancer deletion peaks overlapping recurrently altered regions in follicular dendritic cell sarcoma were determined by comparison to a list of 86 peak deleted regions identified by GISTIC analysis20 across 10844 cancer samples from 33 cancer types, available from the TCGA Copy Number Portal, Broad Institute (analysis version: 2015-06-01 stddata__2015_04_02 regular peel-off, cancer type: all_cancers), accessible online at www.broadinstitute.org/tcga/. 21
Recurrent, homozygous deleted regions were defined by the presence of at least 2 overlapping homozygous (or for case 1, 3-copy) deletions and delineated similarly to smallest regions of overlap, using breakpoints from individual calls to define boundaries.
Tumor suppressor gene functional annotations were performed using various resources, including the Cancer Gene Census list, NCBI Gene (www.ncbi.nlm.nih.gov/gene), OMIM (www.ncbi.nlm.nih.gov/omim), KEGG (www.genome.jp/kegg/) and PubMed (www.ncbi.nlm.nih.gov/pubmed). Actionability/drug sensitivity/targetability and potential therapies were investigated using the following sources: the Drug Gene Interaction Database (DGIdb; v2.09—sha1 c138873);22, 23 Genomics of Drug Sensitivity in Cancer (GDSC) resource (data version 5, 70 genes);24 My Cancer Genome Pathways (www.mycancergenome.org/content/molecular-medicine/pathways/); and the Tumor alterations relevant for genomics-driven therapy (TARGET) database (v3_02142015, www.broadinstitute.org/cancer/cga/target).
Concordance evaluation for molecular inversion probe array and next-generation sequencing panel calls was performed for all likely pathogenic calls listed in Figure 1 of Griffin et al.12 One call (Case 9) was excluded from comparison due to poor probe coverage on the molecular inversion probe array. Discordance was defined as any difference in copy-number state for copy-number alterations across platforms. Due to the complex near-triploid genomic profile of Case 1, discordance was not counted for two- versus -three copy-number losses as these alterations are functionally concordant (nullisomic). For two cases (Case 6 and 9) with apparent discordance due to normalization error by next-generation sequencing, a corrected concordance evaluation was also performed.
Genomic alterations in 14 follicular dendritic cell sarcoma samples. (a) Whole genome view. Vertical bars/alternating coloration delineate boundaries for chromosomes 1–22, X and Y. Top panel: aggregate (stacked calls) of all copy-number calls in this data set (red=single copy loss, dark red=two or greater copy loss, blue=single copy gain, dark blue=two or greater copy gain); y-axis: percentage of cases (n=14) with copy-number calls (only). SROs: brackets indicate smallest regions of overlap for genomic regions recurrently altered by losses/loss-of-heterozygosity in ≥45% abnormal cases (see also Table 2). Bottom panel: Copy-number and loss-of-heterozygosity (yellow) calls within individual cases. ≥2 copy-number alterations are displayed as stacked bars. (b) Chromosome Summary view with aggregate of copy-number/loss-of-heterozygosity calls by chromosome. Left side: losses/loss-of-heterozygosity, right side: gains. Asterisks: recurrently altered regions.
Tables and Figures
Tables were generated and calculations performed in Microsoft Excel. Images were created in Nexus Express Software and modified using Adobe Photoshop. Array data are displayed using the log2 ratio, B-allele frequency, and/or Allele peaks, which display copy-number and genotype data. B-allele frequency assigns each allele a value 1 and calculates B/(A+B). Allele peaks assigns each allele a value 0.5 and calculates A-B.
Results
Clinical and genomic characteristics for each case are summarized in Table 1. Mean patient age was 45 years (range: 19 to 77). Six patients were female and eight were male. Tumor site was nodal in five cases and extranodal in the remaining nine cases. The greater proportion of extranodal tumors in this sample set is largely accounted for by the subset of cases initially described in Griffin et al,12 attributed to the authors’ soft tissue tumor pathology expertise. Follow-up information was available for seven patients: one patient was alive with no evidence of disease at 61 months following diagnosis, one patient was alive with disease at 37 months, five patients died from their disease, on average 30 months following diagnosis (range: 15–37 months).
Molecular inversion probe array analysis revealed genomic alterations in 11 of 14 (79%) samples (Table 1; Figure 1a; Figure 2). The majority of abnormal cases showed extensive genomic complexity with ≥20 alterations observed in 8/11 cases. No general correlations between clinical and genomic data were recognized in this data set. The mean number of alterations per sample was 26 (range: 4–69) and average proportion of the genome altered was 35% (range: 14–62%), consistent with the presence of high genomic instability. Most samples (2–11) had near-diploid <2n> genomic profiles. Sample 1 showed a pattern consistent with near-triploidy, with copy-number alterations considered relative to <3n> (Figure 2c). Analysis of alteration types across abnormal cases showed a predominance of losses (72%, n=290); loss-of-heterozygosity and copy-number gains accounted for 13 and 15% of all calls, respectively (Supplementary Table 1). High copy-number gains (>2 copies gained) were rarely observed (1.0%) while homozygous losses-or for case 1, three copy losses-accounted for 6.5% of all copy-number calls. In three cases (12–14), no clinically significant acquired alterations were detected.
Representative follicular dendritic cell sarcoma cases. (a–c) Whole genome views of samples 14 (a), 4 (b) and 1 (c). Top panels: log2 ratio, middle panels: B-allele frequency, bottom panels: Allele peaks. Vertical lines delineate chromosome boundaries. Arrows: recurrently altered regions. (a) Case 14: normal genomic profile. (b) Case 4: complex genomic pattern with clonal diversity (eg, compare ~50% gain of chr. 20 to ~25% gain of chr. 12). (c). Case 1: complex, near-triploid genomic pattern.
Tumor content across all 14 samples ranged from 40–90% (mean: 70%) (Table 1). Clonal burden in samples 2–11 ranged from 20–80% (mean: 45.5%) (Table 1; Supplementary Table 1). For sample 1, clonal burden could not be determined due to the complex, polyploid genomic profile. A significant difference between average tumor content and clonal burden was recognized (P<0.01, t-test). Since clonal burden is a more direct assessment of tumor DNA in the sample, the possible failure to detect alterations in samples with lower clonal burden or normal genomic profiles due to overestimation of tumor content by H&E cannot be excluded.
Clonal diversity, which is defined as the presence of alterations at different levels of mosaicism in a sample, is suggestive of clonal evolution and is another indicator of genomic complexity. Clonal diversity was evident in three samples (2, 4, and 6) and in 5 others, the presence of clonal diversity was suspected but uncertain due to genomic complexity (sample 1) and/or the presence of only one or two alterations at a distinct level in the sample (samples 7–9, 11) (Table 1; Supplementary Table 1; Figure 2). Three samples (3, 5, and 10) did not show evidence of clonal diversity. Although detection of clonal diversity in some samples may have been limited by clonal burden, its presence in a subset illustrates further the genomic complexity of follicular dendritic cell sarcoma.
Fifteen genomic regions were recurrently altered by losses/loss-of-heterozygosity in 45% (5 cases) or more (Table 2; Figure 1). Approximately half of these regions were defined by the presence of overlapping focal alterations in ≥50% cases, allowing regional specification within a chromosome arm (i.e. distal, medial, or proximal). The most commonly altered region was chromosome 13q in 91% (10/11) cases, followed by chromosomes 3p and 14q in 82% (9/11) of cases. The majority of these alterations (7/10 for 13q, 7/9 for 14q, 5/9 for 3p) were non-focal, including large or whole chromosome losses. Losses/loss-of-heterozygosity in medial 7q (73%) and in medial 9p, medial 11q, proximal 15q and 18q (each observed in 64%) were also frequent. Other chromosome regions with recurrent losses/loss-of-heterozygosity included proximal 1p, medial 2p, 6q, 17p, and distal 22q, each observed in 6/11 (55%) cases; and distal 3q and 8p in 5/11 (45%) cases. Neither recurrent gains nor amplifications were observed.
Smallest regions of overlap, defined by the minimal region of overlapping calls, were delineated within each recurrently altered genomic region (Table 2; Figure 1a). These regions are likely to contain genes contributing toward follicular dendritic cell sarcoma biology, particularly when the majority of calls across cases are focal. For example, the presence of multiple overlapping interstitial deletions in the 1p proximal, 9p medial, and 15q proximal regions suggests that important genes may reside within these chromosome arm regions, while regions such as 13q, 14q, and 18q may contain additional contributory genes outside the smallest regions of overlap due to the predominance of whole-arm/chromosome alterations across samples.
The majority of alterations in recurrently altered regions were losses, suggesting the potential contribution of tumor suppressor genes within these regions. We analyzed the genomic content of smallest regions of overlap for the presence of known or putative tumor suppressor genes using genes lists from the Cancer Gene Census,16 the Mut-driver genes list,18 and the TUSON Explorer manually curated genes list.19 Most of the smallest regions of overlap contained at least one gene that has been implicated as a tumor suppressor gene (Table 2). For several regions (9p medial, 11q medial, 13q, 17p, 18q, and 22q distal), genes annotated in all three tumor suppressor gene lists were contained by the smallest region of overlap, providing strong support for cancer drivers in these regions. Tumor suppressor genes were not identified within the smallest regions of overlap for regions 3p and 8p, the former of which was among the most commonly altered regions. This may be due to the presence of driver genes outside the smallest regions of overlap, which is supported by the presence of non-focal 3p and 8p alterations across the majority of cases; however it is also possible that these regions contain novel tumor suppressor genes or act as passenger alterations that do not directly contribute to follicular dendritic cell sarcoma oncogenesis. Additionally, as our analysis was limited to subsets from these gene lists, it is possible some tumor suppressor genes remain unrecognized within the smallest regions of overlap.
Somatic copy-number alterations are present at a significantly higher frequency compared with background rates in cancer, suggesting these regions may be subject to positive selection.20 To further investigate the significance of recurrently altered genomic regions in follicular dendritic cell sarcoma, we compared the smallest regions of overlap with a list of recurrent focal hemizygous deleted regions in cancer (pan-cancer deletion peaks).21 The majority of these regions (11/15) showed overlap with at least one pan-cancer deletion peak, several of which contained known or putative tumor suppressor genes: 9p medial (CDKN2A), 11q medial (SDHD, USP28), 13q (RB1), 15q proximal (MGA), 17p (TP53 and MAP2K4), and 18q (SMAD4) (Table 2; Figure 1; Figure 3a and c). Previous studies have shown that pan-cancer deletion peaks are enriched for known or putative tumor suppressor genes and depleted of oncogenes and genes essential to cellular function.19, 21, 25 This nonrandom pattern of gene distribution is consistent with an active role for these regions in cancer initiation and/or progression. Thus, other overlapping follicular dendritic cell sarcoma/pan-cancer regions (1p proximal, 3p, 3q distal, 7q medial, 18q, 22q distal) may contain as yet unidentified contributory genes or genomic regions. Four of these regions did not overlap the pan-cancer deletion peaks (2p medial, 6q, 8p, 14q), and may represent cancer-specific or cancer-type-limited alterations, particularly the frequent 14q losses/loss-of-heterozygosity.
Selected recurrent genomic alterations in follicular dendritic cell sarcoma. (a–d) Top panels: aggregate of all copy-number calls; x-axis: genomic position; y-axis: percentage of abnormal cases (n=11) with copy-number calls (only). Middle panels: smallest region of overlap (SRO), pan-cancer deletion peaks (Peak), and Cancer Gene Census genes (Genes) are shown (brackets). Bottom panels: copy-number/loss-of-heterozygosity calls within individual cases; ≥2 copy-number alterations are displayed as stacked bars; flanking dotted lines: smallest regions of overlap; arrows: genes of interest. (a) Chromosome 9 view shows 2.0 Mb smallest region of overlap for loss/loss-of-heterozygosity calls across 7 cases, 4 with homozygous deletions (smallest region of overlap: 21,928,114 [1]-21,974,064 [1]; 53 kb, overlapping CDKN2A). Genes CD274 and PDCD1LG2 in 9p24 are shown. (b) Chromosome 11 view shows 15.0 Mb smallest region of overlap for loss/loss-of-heterozygosity calls across 7 cases, 4 with homozygous deletions (smallest region of overlap: 102,166,344 [4]- 102,296,024 [9]; 130 kb, overlapping BIRC3). (c) Chromosome 13 view shows 22.6 Mb smallest region of overlap for loss/loss-of-heterozygosity calls across 10 cases, 2 with homozygous deletions (smallest region of overlap: 48,920,908 [1] - 49,060,447 [1]; 140 kb, overlapping RB1). (d) Chromosome 16: 2 homozygous deletions (smallest region of overlap: 49,852,987 [1] - 51,757,339 [1]; 1.9 Mb, overlapping CYLD).
As recurrent homozygous deletions and amplifications often signify the localization and function of cancer genes, we interrogated the genomic content of any region showing high copy-number gains or ≥2 copy-number nullisomic losses in two cases or more. Four genomic regions with overlapping homozygous deletions were identified (Figure 3), three of which resided within smallest regions of overlap (9p21.3, 11q22.2, and 13q14.2). The fourth region (16q12.1) did not qualify as being recurrently altered by single copy losses/loss-of-heterozygosity at the 45% cut-off, although the deletion patterns and presence of whole chromosome deletion/loss-of-heterozygosity in two additional cases suggest this region may also contain important genes in follicular dendritic cell sarcoma. All four recurrent homozygously deleted regions contained tumor suppressor genes: CDKN2A in 9p21.3, BIRC3 in 11q22.2, RB1 in 13q14.2 and CYLD in 16q12.1. Accordingly, CDKN2A, RB1, and CYLD are known to act in a recessive manner.16 Hemizygous BIRC3 inactivation (by mutation or deletion) is recurrent in B-cell neoplasms, including splenic marginal zone lymphoma26 and in chemorefractory CLL;27 to the best of our knowledge, homozygous deletion of BIRC3 has not yet been reported; loss of two copies may have additional or distinct pathogenic effects.
Overall, 37 known or putative tumor suppressor genes were identified within recurrently altered regions (Table 3). This association suggests that at least some of these genes have roles in follicular dendritic cell sarcoma pathogenesis. Consistent with their tumor suppressor function, common associated molecular functions and cellular pathways for these genes were recognized, including cell cycle regulation, DNA maintenance and repair, regulation of gene expression (transcriptional and epigenetic), protein turnover, and signal transduction. Interestingly, BIRC3, which was not covered in the 309 cancer gene panel used by Griffin et al,12 also participates in negative regulation of NF-κB signaling,26 a pathway highlighted in their study. 15 tumor suppressor genes are currently classified as actionable due to their prognostic significance and/or drug sensitivity/targetability. These findings support the pathogenicity of these regions and identify potential key genes and pathways that may provide therapeutic targets in follicular dendritic cell sarcoma in the future.
Our sample set included six tumor specimens that were also studied by next-generation sequencing,12 five of which had copy-number data (Table 1). Concordance was examined for 17 of the 18 likely pathogenic copy-number alterations identified by next-generation sequencing, which had adequate array probe coverage for comparison (Supplementary Table 2). Overall, 16/17 (94%) calls were concordant (Table 1). Two cases (6 and 9) with initial one-step discrepancies due to next-generation sequencing normalization were resolved using genome-wide ploidy state determination from the molecular inversion probe array data. Sample 1, with a near-triploid pattern, showed one-step, but functionally concordant calls for nullisomy of CDKN2A, RB1, and CYLD. The single unresolved discrepant call in sample 4 (9p21.3, CDKN2A) was a one-step discordance between loss copy-number state (nullisomy versus monosomy). These cases illustrate the overlapping, but complementary capabilities of copy-number determination by molecular inversion probe array relative to next-generation sequencing.
Discussion
Analysis of 14 follicular dendritic cell sarcoma samples by molecular inversion probe array revealed a genomic landscape characterized by extensive complexity, with losses, gains and loss-of-heterozygosity affecting numerous chromosomes and regions. Compared to previous cytogenetic and next-generation sequencing-based studies, this analysis provides a higher degree of resolution for copy-number changes across the entire genome, with identification of several new recurrently altered regions. The overall predominance of loss-type alterations in these samples suggests a tumor suppressor-driven etiology for follicular dendritic cell sarcoma initiation and/or progression. In support, analysis of the 15 recurrently altered genomic regions, all of which were defined by losses and/or loss-of-heterozygosity, showed overlap with recurrent deletion peaks across cancer and revealed known tumor suppressor genes within the majority of these regions. This type of analysis has not previously been performed for follicular dendritic cell sarcoma, but has been applied to discover key genes with causal roles in oncogenesis across a variety of cancer types. Recurrent homozygous deletions of tumor suppressor genes CDKN2A, RB1, and CYLD were also observed. Homozygous deletion of BIRC3 is an apparently novel finding not only in follicular dendritic cell sarcoma, but also across different cancer types. Although balanced structural and sequence-level alterations were not interrogated in this analysis, the absence of recurrent gains/amplifications from this data set supports our hypothesis. In concordance, a recent next-generation sequencing-based study which included a subset of samples analyzed here (5 analyzed for copy-number), showed similar results.12
Though much of our analysis focused on candidate tumor suppressor genes within the smallest regions of overlap of recurrent alterations, the contribution of large alterations and overall genomic complexity to follicular dendritic cell sarcoma biology should be noted. Whole chromosome and segmental aneuploidy is often driven by defects in the cell cycle, mitosis, and DNA replication and repair.28 In support, many genes classified as tumor suppressors that were identified within recurrently altered regions are involved in these processes. As yet, it is uncertain whether these recurrently altered genomic regions represent primary or secondary alterations in follicular dendritic cell sarcoma. While a clear pattern of clonal progression cannot be outlined from the genomic profiles observed in this set of cases, the most common alterations in this sample set, whole-arm/-chromosome losses of 3p, 13, and 14, may represent early events in follicular dendritic cell sarcoma pathogenesis, potentially involving a large number of genes. The presence of focal alterations in all regions helps to narrow down critical genomic loci that may provide additional markers for genomic characterization and for future studies with clinical data, potential correlates to disease progression and/or outcomes.
Current diagnosis and management of follicular dendritic cell sarcoma is based upon clinical, morphologic and immunohistochemical findings,3 while genetic analysis offers the potential to provide new pathogenetic insights and future therapies. Although karyotypes in follicular dendritic cell sarcoma have mostly been inconsistent, likely due to technical difficulties in obtaining representative metaphases through culturing,9 molecular inversion probe array identified novel, recurrent genomic alterations, highlighting the utility of this analysis for genomic characterization. Future work to expand upon these findings and incorporate standard clinical and pathologic criteria with copy-number and mutational status, including rearrangement detection is needed. Although the identification of novel therapies is beyond the scope of this study, the presence of recurrent alterations of known tumor suppressor genes in follicular dendritic cell sarcoma provides insight into the molecular and cellular pathways that drive this rare tumor type, and identifies important new avenues of therapeutic investigation.
References
Heesters BA, Myers RC, Carroll MC . Follicular dendritic cells: dynamic antigen libraries. Nat Rev Immunol 2014;14:495–504.
Saygin C, Uzunaslan D, Ozguroglu M et al, Dendritic cell sarcoma: a pooled analysis including 462 cases with presentation of our case series. Crit Rev Oncol Hematol 2013;88:253–271.
Chan JKC, Pileri SA, Delsol G et al, Follicular dendritic cell sarcoma. In: Swerdlow SH, Campo E, Harris NL et al, (eds). WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues, 4th ed. IARC: Lyon, 2008, pp 363–364.
Li J, Zhou ML, Zhou SH . Clinical and pathological features of head and neck follicular dendritic cell sarcoma. Hematology 2015;20:571–583.
Youens KE, Waugh MS . Extranodal follicular dendritic cell sarcoma. Arch Pathol Lab Med 2008;132:1683–1687.
Perkins SM, Shinohara ET . Interdigitating and follicular dendritic cell sarcomas: a SEER analysis. Am J Clin Oncol 2013;36:395–398.
Pang J, Mydlarz WK, Gooi Z et al, Follicular dendritic cell sarcoma of the head and neck: Case report, literature review, and pooled analysis of 97 cases. Head Neck 2016;38:E2241–E2249.
Gounder M, Desai V, Kuk D et al, Impact of surgery, radiation and systemic therapy on the outcomes of patients with dendritic cell and histiocytic sarcomas. Eur J Cancer 2015;51:2413–2422.
Udayakumar AM, Al-Bahri M, Burney IA et al, Follicular dendritic cell sarcoma: cytogenetics and pathological findings. Sultan Qaboos Univ Med J 2015;15:e411–e414.
Go H, Jeon YK, Huh J et al, Frequent detection of BRAF(V600E) mutations in histiocytic and dendritic cell neoplasms. Histopathology 2014;65:261–272.
Starr JS, Attia S, Joseph RW et al, Follicular dendritic cell sarcoma presenting as a thyroid mass. J Clin Oncol 2015;33:e74–e76.
Griffin GK, Sholl LM, Lindeman NI et al, Targeted genomic sequencing of follicular dendritic cell sarcoma reveals recurrent alterations in NF-kappaB regulatory genes. Mod Pathol 2016;29:67–74.
Wang Y, Cottman M, Schiffman JD . Molecular inversion probes: a novel microarray technology and its application in cancer research. Cancer Genet 2012;205:341–355.
Paxton CN, Rowe LR, South ST . Streamlining the OncoScan(R) array procedure for use in a clinical laboratory. J Assoc Genet Technol 2015;41:61–65.
Conlin LK, Thiel BD, Bonnemann CG et al, Mechanisms of mosaicism, chimerism and uniparental disomy identified by single nucleotide polymorphism array analysis. Hum Mol Genet 2010;19:1263–1275.
Futreal PA, Coin L, Marshall M et al, A census of human cancer genes. Nat Rev Cancer 2004;4:177–183.
Forbes SA, Beare D, Gunasekaran P et al, COSMIC: exploring the world's knowledge of somatic mutations in human cancer. Nucleic Acids Res 2015;43:D805–D811.
Vogelstein B, Papadopoulos N, Velculescu VE et al, Cancer genome landscapes. Science 2013;339:1546–1558.
Davoli T, Xu AW, Mengwasser KE et al, Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell 2013;155:948–962.
Beroukhim R, Mermel CH, Porter D et al, The landscape of somatic copy-number alteration across human cancers. Nature 2010;463:899–905.
Zack TI, Schumacher SE, Carter SL et al, Pan-cancer patterns of somatic copy number alteration. Nat Genet 2013;45:1134–1140.
Griffith M, Griffith OL, Coffman AC et al, DGIdb: mining the druggable genome. Nat Methods 2013;10:1209–1210.
Wagner AH, Coffman AC, Ainscough BJ et al, DGIdb 2.0: mining clinically relevant drug-gene interactions. Nucleic Acids Res 2016;44:D1036–D1044.
Yang W, Soares J, Greninger P et al, Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res 2013;41:D955–D961.
Solimini NL, Xu Q, Mermel CH et al, Recurrent hemizygous deletions in cancers may optimize proliferative potential. Science 2012;337:104–109.
Rossi D, Deaglio S, Dominguez-Sola D et al, Alteration of BIRC3 and multiple other NF-kappaB pathway genes in splenic marginal zone lymphoma. Blood 2011;118:4930–4934.
Rossi D, Fangazio M, Rasi S et al, Disruption of BIRC3 associates with fludarabine chemorefractoriness in TP53 wild-type chronic lymphocytic leukemia. Blood 2012;119:2854–2862.
Burrell RA, McGranahan N, Bartek J et al, The causes and consequences of genetic heterogeneity in cancer evolution. Nature 2013;501:338–345.
Acknowledgements
Support for this research was provided by the ARUP Institute for Clinical and Experimental Pathology. The authors thank Xinjie Xu from ARUP Laboratories for assembly of some of the genes lists used in this analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Supplementary information
Rights and permissions
About this article
Cite this article
Andersen, E., Paxton, C., O'Malley, D. et al. Genomic analysis of follicular dendritic cell sarcoma by molecular inversion probe array reveals tumor suppressor-driven biology. Mod Pathol 30, 1321–1334 (2017). https://doi.org/10.1038/modpathol.2017.34
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2017.34
This article is cited by
-
Chimeras in Merlot grapevine revealed by phased assembly
BMC Genomics (2023)
-
Recurrent PDGFRB mutations in unicentric Castleman disease
Leukemia (2019)