Abstract
Advances in digital pathology, specifically imaging instrumentation and data management, have allowed for the development of computational pathology tools with the potential for better, faster, and cheaper diagnosis, prognosis, and prediction of disease. Images of tissue sections frequently vary in color appearance across research laboratories and medical facilities because of differences in tissue fixation, staining protocols, and imaging instrumentation, leading to difficulty in the development of robust computational tools. To address this challenge, we propose a novel nonlinear tissue-component discrimination (NLTD) method to register automatically the color space of histopathology images and visualize individual tissue components, independent of color differences between images. Our results show that the NLTD method could effectively discriminate different tissue components from different types of tissues prepared at different institutions. Further, we demonstrate that NLTD can improve the accuracy of nuclear detection and segmentation algorithms, compared with using conventional color deconvolution methods, and can quantitatively analyze immunohistochemistry images. Together, the NLTD method is objective, robust, and effective, and can be easily implemented in the emerging field of computational pathology.
Similar content being viewed by others
Main
Advances in imaging instrumentation and data management provide the foundation for computational approaches to analyze digitized images of tissue sections and derive objective, quantitative measurements at the tissue, cellular, subcellular, and molecular levels.1 Computational pathology approaches offer a cost-effective platform to increase throughput, accuracy, and reliability of diagnoses of tissue samples.2, 3 Further, the quantitative nature of computational pathology can be used in combination with other assays to improve pathologists’ knowledge of disease and help inform treatment strategies and further stratify patient prognosis. It has been shown that, by integrating information derived from computational pathology with a patient’s clinical data, a better prognostic model can be derived for many diseases, including prostate cancer,4, 5, 6 lung cancer,7 breast cancer,8, 9, 10, 11, 12 glioblastoma,13, 14 basal cell carcinoma,15, 16 and ovarian cancer.17, 18
One of central challenges of computational biology, which limits its large-scale applications, is that images of tissue sections frequently vary in color appearance across research laboratories and medical facilities due to differences in tissue fixation, staining protocols, and imaging instrumentation. The wide spectrum of image color appearance causes difficulty in robustly extracting the representative images of different tissue components, such as nuclei.19 Previous studies have shown that technician variance or technique differences can lead to marked differences in staining.20 For example, the conventional hematoxylin and eosin (H&E) staining techniques have been modified to reduce material use and processing time21 or to improve the contrast and detail in the digital image.22 These technique differences provide some advantage to the pathologist, and also lead to variation in the staining of slides for use in computational pathology approaches that must be addressed.
Several stain normalization computational approaches—including color deconvolution (CD),23 histogram equalization,24 and the use of the CMYK space25—have been developed to correct for the difference image appearance and facilitate the separation of tissue types.19, 20 Of these approaches, CD is the most commonly used approach to extract nuclear and cellular images in both H&E and immunohistochemically (3,3′-diaminobenzidine, DAB) stained images.2, 9, 23, 26, 27, 28 CD uses the method of singular value decomposition, which seeks to linearly separate the color space to identify regions rich in each particular dye. However, a major disadvantage of CD is the requirement of prior knowledge for each dye’s color spectrum to visualize accurately tissue components.29 Owing to color appearance difference between images, using the same stain vector across images will introduce variance in the representative image for each dye. Although there are automated methods to determine the stain vector for individual images, the additional processing step leads to significant increase in processing time across large image data sets.30 Furthermore, CD only decouples the concentration of dye in the histopathological image, and further processing is needed to separate individual tissue components such as blood, nuclei, and extracellular matrix- and cytoplasmic-rich regions for quantification.
In this work, we propose a novel nonlinear tissue-component discrimination (NLTD) method to register automatically the color space of histopathology images and obtain representative images for individual tissue components, such as the nuclei or cytoplasm, irrespective of perceptual color differences between images. We demonstrate that the nuclei image obtained from NLTD display consistent appearance for histopathology images—including those with distinct color differences—taken from different tissues types and prepared at different institutions, including The Cancer Genome Atlas project (TCGA, http://cancergenome.nih.gov/). Importantly, the processing time of NLTD is highly comparable to the CD for small images, and much more efficient for large images, notably whole slide images. Further, we demonstrated that the nuclei images derived using NLTD produce highly accurate nucleus tracing and counting, and NLTD allows for quantitative analysis of antigen presence in immunohistochemical images. Taken together, we show that NLTD is an effective approach to obtain quantitative tissue-component images that can be easily integrated in emerging computational pathology applications.
MATERIALS AND METHODS
The NLTD method consists of five main steps (Figure 1a), detailed further here: (1) color joint-histogram creation; (2) ridge detection; (3) ridge set registration; (4) transformation function creation; and (5) tissue component image creation.
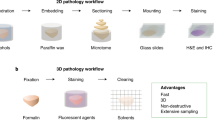
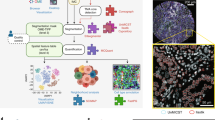
Brief overview of nonlinear tissue-component discrimination (NLTD) approach. (a) NLTD applied to an image of a hematoxylin and eosin (H&E)-stained section (top) and immunohistochemically (IHC) stained image (bottom). Shown are a typical H&E image of a small artery, exhibiting multiple tissue components (nuclei (N), extracellular matrix (ECM)-rich and cytoplasm (E), blood (B)) and a typical IHC image, stained for LINE-1 ORF1p expression,31 exhibiting two tissue components (antigen (A) and nuclei (N)). The NLTD method is schematically shown in the center. Briefly, the red-blue joint histogram is first segmented to identify each region in the red-blue color space. The x axis corresponds to each red color, the y axis shows each blue-color, and the color axis represents the frequency of each discrete color combination. Ridges for each tissue component are overlaid, on the red-blue color joint histogram (RBJH). The ridge set is registered and transformed to yield the pseudocolored transformation function for each component. The pseudocolored grayscale images are shown for the nuclei, non-nuclei, and blood components (purple, pink, and red, respectively) in the far right box. (b and c) Grayscale correlation values for the red-blue joint histogram, blue-green joint histogram, and red-green joint histogram, with a value of 1 corresponding to a completely correlated colorset. (b) Pancreatic cancer H&E data set (n=45). (c) Ovarian immunohistochemistry data set (n=81). (d) Separation of red-blue color space into individual tissue components: nuclei (purple box); ECM and cytoplasm-rich (pink box).
Color Joint-Histogram Creation
In a typical 8-bit tissue image, I, the color of an individual pixel, p, at location (xp, yp) is expressed by three intensities (rp,gp,bp), each of which ranges from 0 to 255, discretely, that is,

For example, if all the intensities of a pixel are zero, the resultant color is black; conversely, if all are 255, the resultant color is white. The color joint histogram is a three-dimensional histogram created by counting the occurrence of pixels at all different set of red, green, and blue (RGB) intensities in an image. However, calculating every color combination in RGB color space and analyzing three-dimensional RGB color joint histogram is a highly computationally intensive process. An 8-bit image can contain more than 16 million unique combinations. To reduce computational time, it would be advantageous to only consider two of the three color axes, reducing the number of unique combinations 256-fold.
In a cohort of 45 H&E images, we found that blue and green color components are highly correlated within individual images (Figure 1b). Furthermore, we also found that the red and green color components are highly correlated in a set of 81 immunohistochemically stained images (DAB)31 (Figure 1c). These observations show that in both H&E and DAB images green color channel encodes highly correlative information to other color channel and implies that the red-blue color joint histogram (RBJH) can be a representative simplification of the histopathological image RGB color space. The RBJH is a two-dimensional matrix, created by counting the frequency (n) of pixels at different red (r) and blue (b) intensity values in the image (I), that is,

The resultant RBJH can be visualized as a three-dimensional surface, with the x and y axes corresponding to the red and blue color space values, respectively, and the z axis corresponding to the incidence rate for each red-blue intensity combinations.
Ridge Detection
The RBJH shows the most abundant color combinations in the red-blue color space for an image. In the RBJH, distinct populations of red-blue combinations are readily observed, corresponding to different tissue components (Figure 1d). However, to detect and segment individual tissue components in the RBJH is challenging because of the elongated nonlinear distribution of red-blue color combinations that complicates separation of the populations and, consequentially, common approaches, such as Gaussian mixture models or k-means clustering, do not work well. Gaussian mixture models fail because most images’ RBJH do not follow strictly Gaussian distributions, often having one major peak, along with a long sparse tail. Similarly, k-means clustering is not effective at detecting sparse areas in the RBJH. Additionally, both models require prior knowledge of the numbers of components present, which we have found can vary from 1 to 3 in most images. Successful extraction of individual tissue components’ locations in the red-blue color space needs to incorporate the asymmetric and elongated distribution of red-blue color combinations that is observed in the RBJH. Therefore, we propose to identify locations of major tissue components in the red-blue color space by tracing the location of ridges of distinct population in the RBJH (Supplementary Figure S2).
We first identified the major orientation of signal in RBJH using weighted principal component analysis (PCA) (Supplementary Figure S1a). PCA is applied to the each red and blue index of the RBJH, with the frequency (nr,b) associated with each color combination used as a weight. The principal component provides the major direction of the RBJH color space, which can be combined with the location of the global maximum of the RBJH to create a major axis (v0). Next, we identify the local maxima tangent along the major axis in RBJH (Supplementary Figure S1b). To ensure the all local maximum is detected, this routine is repeated along two other vectors at angles ±15° of the major direction. A map of all local maxima identified is then created by counting the frequency of local maxima identified at each red-blue index (Supplementary Figure S1c). This map is further processed through morphological dilation and thinning operations to provide a binarized location of ridges for all distinct populations in RBJH (Supplementary Figure S1d).
Ridge Set Registration
To register the ridge set maps in RBJH to different tissue components, we developed a robust algorithm based on each ridge’s proximity to specified reference color combination (Supplementary Figure S2a). Four reference color combinations are used: Red (r=255, b=0), Blue (r=0, b=255), Black (r=0, b=0), and White (r=255, b=255). The Euclidean distance transform32 is calculated for each reference color combination, and the minimum distance along each ridge is found. The component with the smallest distance from a reference color combination is determined to be the closest. In H&E staining, in general, nuclei would be closer to black and blue, whereas extracellular matrix- and cytoplasm-rich areas would be closer to white. Similar logic can be applied to DAB chromagen staining, where antigen-rich areas are closer to red than nuclei, which are closer to blue.
In some cases, where the RBJH is more homogeneous, it is possible that only one ridge is found (Supplementary Figure S2b). For H&E staining, in general, each image contains at least two distinct components—nuclei or extracellular matrix and cytoplasm-rich components. The portion of ridge closer to white and red represents extracellular matrix and cytoplasm-rich areas and the remaining portion of the ridge represents the nuclei-rich regions; this necessitates further segmentation of the ridge. To segment the ridge, the intensity profile (ie, frequency of color combinations) of the RBJH along the ridge is first extracted. A peak, corresponding to the most frequently occurring color combination, commonly appears that represents the central location of extracellular matrix and cytoplasm-rich regions on the ridge. Thus, we fit this intensity profile with a Gaussian distribution to measure the distribution of the extracellular matrix and cytoplasm-rich areas along the ridge, and segment the single identified ridge using a distance of 2 s.d. from the peak into two distinct ridges corresponding to nuclei or extracellular matrix and cytoplasm-rich components. For DAB chromagen staining, the possibility of only one ridge being identified is most likely due to the lack of presence of antigen to probe in the tissue section and hence no further segmentation is needed.
Transformation Function Creation
We formulated tissue transformation functions (TF) to convert the red-blue color space to intensity of different tissue components (k=1,2,…,N). We assume the red-blue color space has different regions that exclusively correspond to different tissue components based on the proximity to each ridge in the ridge set. A watershed segmentation is applied to the ridge of the RBJH to identify regions of the red-blue color space that represent the unique tissue components (Supplementary Figure S3a). Additionally, the regions of the red-blue color space with the most absorption (ie, lower r and/or b indices) correspond to the strongest signal within each tissue region. For each particular tissue component, the red and blue indices that are closest to the tissue’s ridge indicate a higher likelihood of belonging to that tissue and also contribute to a stronger signal.
To account for these three factors, we developed a transformation function, TFk, that account for the tissue component’s region in red-blue color space (fregion), its absorption (fabsorption), and the distance from each tissue component’s ridge (fridge) (Supplementary Figure S3b), expressed by

where k=1,2,…,N tissue component.
The tissue region function, fregion, uses the watershed segmentation result as a basis to exclude any part of the red-blue color space not belonging to the same tissue component. The region of the red-blue color space corresponding to the kth tissue component is defined as Wk. A Gaussian filter, g, can be applied to the edge of the region to allow for a smoother transition between components (Supplementary Figure S3b(i)), that is,

The absorption function, fabsorption, for kth tissue component is obtained by first calculating the Euclidean distance transform32 (Ddark) of all points in red-blue color space from the point with highest absorption (ie, darkest) on the kth tissue-component ridge (Rk), defined as the point on the ridge closest to black (r=0, b=0). To scale the distance with level of absorption of dye, the absorption function (Supplementary Figure S3b(ii)), is expressed by

The ridge function, fridge, is derived from the Euclidean distance transform,32 D, which is the minimum distance of any point in the red-blue color space to a point on the ridge of the kth tissue component, Rk. To scale the likelihood with distance, the ridge function (Supplementary Figure S3b(iii)), is expressed by

Tissue Component Image Creation
To obtain the kth tissue-component image, Tk, the red and blue pixel intensities (rp, bp) at each location (xp,yp) in the original image, I, were used to create a grayscale image according to the transformation function, TFk, that is,

Sample Acquisition
Histopathological images were acquired from pathologists at the Johns Hopkins University. The tissue samples were formalin fixed and paraffin embedded. Tissue sections were fixed for 3 h in formalin on tissue processor, followed by 1–2 h of gross room fixation. Paraffin sections were cut at 5 μm thickness. Sections were then stained with H&E and digitized using a DP27 5MP color camera. Sections of pancreatic cancer, colon cancer, ovarian cancer, and glioblastoma were included. Immunohistochemically (DAB) stained tissue was acquired through an ovarian cancer tissue microarray, as described previously.31 Additional tissue images were acquired from The Cancer Genome Atlas project (http://cancergenome.nih.gov) and published sources.33, 34
Nuclei Detection
To perform a comparison of segmentation results between CD and the NLTD method, a publicly available data set,33 including both tissue images and ground-truth nuclei locations, was analyzed. For the NLTD method, the corresponding nuclei image was obtained and nuclei location is obtained using following procedure:
-
1)
Binarize each image using a dynamic threshold, calculated using Otsu’s method.35
-
2)
Remove small objects based on a size threshold of 50 pixels.
-
3)
Watershed segmentation to separate clusters of nuclei.
The same segmentation approach was used for the CD image corresponding to the hematoxylin dye. For each segmented nucleus identified, the nearest ground-truth nucleus was found. If two segmented nuclei were attached to the same ground-truth nuclei, the nearest would be counted as a true positive and the other would be counted as a false positive. Any segmented nucleus with the nearest ground-truth nuclei was more than one average cell diameter away was counted as a false positive. Conversely, any ground-truth nucleus that did not have any segmented nuclei within one average cell diameter was counted as a false negative.
Immunohistochemistry Scoring
A TMA of ovarian cancer tissue stained using an antibody for LINE-1 ORF1p31 was used to evaluate the utility of NLTD as an immunohistochemistry scoring aide. Each image in the TMA was separated into two images using the NLTD method, a nuclei- and an antigen-rich image (Supplementary Figure S5). Preprocessing steps were performed to only analyze nuclei-rich regions where antigen staining was present, and avoid background areas where no staining should occur. Briefly, the nuclei-rich image was segmented using Otsu’s thresholding technique.35 Small objects were removed from the image, followed by morphological opening and closing operations and another removal of small objects. After preprocessing, a transformation score was derived based on the ratio of antigen intensity to nuclei intensity (Equation (8)). Importantly, only antigen and nuclei intensity in the areas from the segmented, preprocessed image were counted.

Hardware and Software
All image processing was performed using MATLAB 2015 (Mathworks). To determine statistical significance, two-tailed t-tests were performed using Graphpad Prism 6. All computations were performed on Windows 7 Professional with an Intel Core i7-3820 processor and 16 GB RAM.
Statistics
To quantify the segmentation results, precision, recall, and F-score statistics were used.36 For this data set, it is not possible to assess accuracy or other statistics using true-negative counts, as the classification system has no negative result included and only positive occurrences (ie, nuclei) are identified. Each statistic is defined as follows:



RESULTS
Overview of the NLTD Method
The NLTD method presented in this work consists of five major steps, as illustrated with an H&E-stained image and an immunohistochemically stained image in Figure 1a. First, the RBJH is created. This joint histogram represents the frequency at which each red and blue pixel intensity combination occurs in a histopathological image, and serves as the basis for tissue-component discrimination. The RBJH is reduced to a set of curves representing the ridges, or local maxima, using an iterative approach. This ridge set is further registered with corresponding individual tissue components (eg, nuclei, extracellular matrix, and cytoplasm-rich, etc). Further, the ridge set serves as a basis for the creation of a set of transformation functions used to create individual, grayscale images from the original image representative of each tissue component present in the image (see more details in Materials and Methods section). The resulting set of tissue-component images can then be used for additional tissue processing and analysis, including nuclei detection algorithms and quantitative scoring of immunohistochemically stained samples. The MATLAB package is available upon request.
Robustness of NLTD
To demonstrate the robustness of the NLTD method, we applied NLTD to a set of histopathological images with wide range of apparent colors to show the uniformity in the images of nuclei extracted using the NLTD method (Figure 2). The image set spans multiple tissue types, along with several different image sources: the Johns Hopkins School of Medicine; images from previous studies performed at University of California, Santa Barbara34 and the University of Berlin;33 and publicly available images from the TCGA image database (http://cancergenome.nih.gov). The results show that even though the RBJH color space for different images have unique and different distributions, the NLTD method can successfully identify and register each tissue-component and extract nuclei images consistently and robustly.
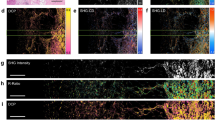
Application of nonlinear tissue-component discrimination (NLTD) across a wide variety of tissue types. The NLTD method is applied on many different types of tissue. The original tissue image, red-blue color joint histogram (RBJH), registered RBJH, and the nuclei component grayscale image (pseudocolored purple) are shown (left to right). The registered RBJH shows a purple line for the nucleus component, a pink line for the extracellular matrix (ECM)/cytoplasmic component, and a red line for the blood component. The sample tissue types are: (a) colon cancer, (b) kidney cancer, (c) ovarian cancer, (d) lung adenocarcinoma, (e) gastric mucosa, (f) astrocytoma, (g) skin cutaneous melanoma, and (h) breast cancer.
Processing Time
For most computational pathology applications, the time associated with processing each image and gleaning important information can quickly become a barrier with image size and/or lager cohort of images. Previous work has compared stain normalization processing time for smaller images (256 × 256, 512 × 512, and 1024 × 1024).29 Whole slide imaging, however, often results in much larger images (10 000 × 10 000 or greater), and it is important for image processing time to scale well with the size of each image. In our work, we compared CD, CD using Macenko’s color normalization method,37 and the NLTD approaches. Macenko’s approach involves an additional preprocessing step to determine each individual image’s optimal stain vector and uses the optimal stain vector for CD. Both CD approaches were faster than the NLTD method at small image size (up to 2500 × 2500), but, as the image size approached whole slide levels (15 000 × 15 000), the NLTD method was much faster than both CD-based methods and took only a quarter of the time to process each image (Figure 3). This result suggests that NLTD can more efficiently analyze larger images, which can be very useful for large data sets, such as the TCGA. Since CD is more time efficient at smaller sizes, it is possible to partition one large image into many smaller images (ie, one 10 000 × 10 000 image into one hundred 1000 × 1000 images). However, this additional processing step would still lead to an increase in processing time compared with the NLTD approach (3.8 s for NLTD on one 10 000 × 10 000 image, 11.85 s for CD on one hundred 1000 × 1000 images). Therefore, the NLTD method can be more efficiently applied to whole slide images and reduce the time needed to analyze large cohorts of images.
Processing time of nonlinear tissue-component discrimination (NLTD) and color deconvolution (CD). Comparison time for processing of images of various sizes using three different color normalization techniques: NLTD (squares), CD (circles), and CD using Macencko’s method of automated stain vector determination (MMCD, diamonds). Each image used was a three-dimensional red, green, and blue (RGB) image, with side lengths defined by the x axis. Processing time is shown on the y axis in seconds as the median of 10 runs for each method at each image size.
Improving Nuclei Detection with NLTD
Nuclei detection in histopathological images has been critical and often used in computational pathology approaches to develop prognostic and diagnostic models.7, 8, 9, 11, 12, 17, 33 Currently, CD is commonly used to extract a representative nuclei image (corresponding to the hematoxylin dye levels) to apply nuclei detection algorithms.9, 28, 29, 30, 33, 38 Here, we show that using the nuclei image derived from the NLTD method improves the detection of nuclei over the CD approach. We first evaluated the contrast of individual nuclei images created from both the NLTD and the CD method (Figures 4a–c). By examining the intensity profile along one axis across nuclei, we found that the nuclei image obtained from NLTD has a substantial decrease in intensity at the periphery of the nucleus compared with nuclei images from CD. This result suggests that the implementation of segmentation algorithms to the NLTD nuclei image would be less sensitive to the intensity threshold value and hence could lead to improvements in the accuracy and robustness of nuclei segmentation algorithms.
Evaluation of nonlinear tissue-component discrimination (NLTD) method. (a–c) Nuclei intensity comparison between NLTD and color deconvolution (CD) approaches. Representative nuclei from several tissue types are shown, along with the NLTD and CD nuclei transformations. The intensity of each color space is integrates along the dotted lines shown, with the NLTD intensity shown in purple and the CD intensity in brown. Intensity values are normalized linearly between 0 and 1, with 0 corresponding to the minimum value in the input image, and 1 corresponding to the maximum. (d) Typical breast cancer image.33 (e) Example of segmentation results from Otsu thresholding of the nuclei NLTD color space. Detected nuclei are overlaid on top of the image from panel (d). True positives are represented by a green dot, false positives by a red dot, and false negatives by a yellow dot. (f) Precision, (g) sensitivity, and (h) F-score values for segmentation results from 35 images. (i) Receiver-operating characteristic curve for change in segmentation parameterization (threshold value) for nuclei detection. Recall (sensitivity) is shown on the x axis, with precision shown on the y axis. Results from NLTD method are shown in black, with CD shown in gray.
To examine quantitatively the performance of the NLTD and CD methods in nuclei detection and segmentation, we applied previous proposed detection algorithm (see more details in Materials and Methods section) after applying both color normalization methods (NLTD and CD) to a published set of 35 images.33 This data set included nuclei locations that had been previously registered by a pathologist and were used as ground truth (Figures 4d and e).
To assess each method, the precision, sensitivity, and F-score were measured. High precision and sensitivity are both valuable in a nuclei detection system. A system that lacks precision will lead to unnecessary calculation and validation by an observer with too many nuclei identifies. Conversely, a system that is not sensitive will miss many nuclei and potentially distort the values of nuclei counts or exclude rare nuclei events, such as mitotic or atypical nuclei. The F-score provides the harmonic mean between sensitivity and precision and serves as an overall measure of how accurate the system is.
Among the 35 images tested, we found that, overall, the images normalized using NLTD have significantly higher sensitivity in detecting nuclei than the corresponding CD images (NLTD=0.868; CD=0.753), but slightly lower precision (NLTD=0.938; CD=0.976) (Figures 4f–h). The overall accuracy, as represented by the F-score, for NLTD images is 0.860 and is significantly higher than the CD images (F-score=0.805). The slightly lower precision in our NLTD system correlates to an overdetection, with more nuclei identified by the NLTD detection system than the ground truth. The higher sensitivity, however, means that the NLTD detection system leaves fewer ground truth nuclei undetected. Taken together, these results suggest that the NLTD method is able to provide more accurate nuclei segmentation results, compared with conventional CD methods.
NLTD for Quantitative Immunohistochemistry Analysis
In addition to providing a platform for image appearance normalization and nuclei detection, the NLTD method can be used as a companion diagnostic for analysis of immunohistochemical labeling quantitatively and objectively. The intensity level of DAB chromagen labeling is used to access the level of antigen presence in tissue sample by pathologists. We applied our method to an ovarian cancer tissue microarray cohort that had been immunolabeled for L1ORF1p, a cytoplasm-localizing protein associated with cancer31 (Figure 5 and Supplementary Figure S4). Each tissue sample in this cohort was scored by a trained pathologist using a discrete scoring system (0, 1, 2, or 3). A score of 0 indicates no significant protein expression, whereas a score of 3 was given for high expression. We applied our NLTD method to individual tissue images of the TMA to create component images for antigen- and nuclei-rich regions. These images were then used to calculate an overall score corresponding to the level of antigen, normalized by nuclei intensity (see more details in Materials and Methods section). Our results showed a strong correlation (Spearman’s ρ=0.8122) between our automated scoring platform and the scoring by the pathologist. Minor overlap exists between tissues with a score of 1 and 2, but both high expression (3) and very low expression (0) scores were well stratified. This result shows the utility of our NLTD method as a nonparametric tool to assess immunolabeling.
Nonlinear tissue-component discrimination (NLTD) method as a quantitative descriptor for immunohistochemistry (IHC). Ovarian tissue samples were stained with an antibody for LINE-1 ORF1p and manually scored by a pathologist31 on a discrete scale of 0 (no expression) to 3 (high expression). A quantitative score is calculated using the NLTD grayscale images. The scores correlate well, with a Spearman's p=0.8122.
DISCUSSION
CD,23 and other associated methods,9, 26, 27, 28, 29, 37 are routinely used for dye separation in histopathological images, but are limited by difference in dye appearance between images, potentially time-consuming automated image processing, and a need for further postprocessing to identify specific tissue components, such as the nuclei. The NLTD approach presented here is able to bypass these limitations, specifically the requirement of prior knowledge of color information for different batches of histopathological images. The NLTD approach makes no inherent assumptions about the histopathological image’s color space, and yields consistent, batch-invariant tissue component separation in histopathological images. We demonstrate that the NLTD method can successfully identify nuclei for a wide variety of histopathological images despite large variations in the perceptual color space (Figure 2). Importantly, no prior knowledge or user input is required, as our algorithm will automatically register locations of for each tissue-component, and the method can be used across multiple batches of images without additional user input. Therefore, NLTD method can be seamlessly integrated in computational pathology pipelines that aim to analyze large cohorts of images, such as the TCGA project (http://cancergenome.nih.gov/) or Human Protein Atlas Project.39 The TCGA project also provides the opportunity to link morphological features of the histopathological images with genomic information, with potential for better understanding of what effect the changes in gene expression can have on the morphology of the tissue.
The tissue component images created through the NLTD method can be readily analyzed to yield additional information, such as nuclei information and immunohistochemical grading. We found that the NLTD method performs nuclei segmentation better than the CD approach. The segmentation approach presented here based on a simple implementation of Otsu’s thresholding, but more refined approaches, as mentioned in reviews of computational pathology,19, 20 should lead to greater accuracy using tissue-component images from the NLTD method. We have demonstrated that the nuclei component images generated using the NLTD method have greater separation of signal from background compared to CD, suggesting simpler processes for nuclei edge detection can be used and lead to significantly reduced segmentation times. Nuclei detection requires very fast computation since an individual tissue image can have millions of nuclei, leading to large increases in total processing time with each additional nuclei detection step.
The field of computational pathology is rapidly growing, and there are many opportunities for computational approaches to provide additional prognostic and diagnostic information that cannot be provided by pathologists alone.40, 41, 42 The NLTD method presented here provides a framework that can be easily implemented for many different applications, including nuclei detection and immunohistochemistry grading. In addition to these applications, NLTD could be used as a visualization tool to normalize tissue appearance across batches, provide texture information for abundance of certain tissue components in a sample, or identify rare occurrences in whole slide images, such as mitotic nuclei. Further, the NLTD method requires no prior knowledge of an image’s color space and requires no parameterization from the user, which can allow for pathologists or medical technicians to apply this approach without requiring more sophisticated knowledge that may be needed for optimization methods or complex, linear algebraic approaches. Taken together, the proposed NLTD method presents an opportunity to establish a pipeline for classification and analysis of histopathological images that, in combination with pathologists’ expertise, can lead to better diagnosis and treatment planning for patients in the future.
References
Hipp J, Flotte T, Monaco J et al. Computer aided diagnostic tools aim to empower rather than replace pathologists: lessons learned from computational chess. J Pathol Inform 2011;2:25.
De Sousa EMF, Wang X, Jansen M et al. Poor-prognosis colon cancer is defined by a molecularly distinct subtype and develops from serrated precursor lesions. Nat Med 2013;19:614–618.
Beck AH, Sangoi AR, Leung S et al. Systematic analysis of breast cancer morphology uncovers stromal features associated with survival. Sci Trans Med 2011;3:108ra13.
Kwak JT, Hewitt SM, Sinha S et al. Multimodal microscopy for automated histologic analysis of prostate cancer. BMC Cancer 2011;11:62.
Doyle S, Feldman M, Tomaszewski J et al. A boosted Bayesian multiresolution classifier for prostate cancer detection from digitized needle biopsies. IEEE Trans Biomed Eng 2012;59:1205–1218.
Monaco JP, Tomaszewski JE, Feldman MD et al. High-throughput detection of prostate cancer in histological sections using probabilistic pairwise Markov models. Med Image Anal 2010;14:617–629.
Hamilton PW, Wang Y, Boyd C et al. Automated tumor analysis for molecular profiling in lung cancer. Oncotarget 2015;6:27938–27952.
Yuan Y, Failmezger H, Rueda OM et al. Quantitative image analysis of cellular heterogeneity in breast tumors complements genomic profiling. Sci Transl Med 2012;4:157ra143.
Veta M, Kornegoor R, Huisman A et al. Prognostic value of automatically extracted nuclear morphometric features in whole slide images of male breast cancer. Mod Pathol 2012;25:1559–1565.
Wang C, Pécot T, Zynger DL et al. Identifying survival associated morphological features of triple negative breast cancer using multiple datasets. J Am Med Inf Assoc 2013;20:680–687.
Wang L-W, Qu A-P, Yuan J-P et al. Computer-based image studies on tumor nests mathematical features of breast cancer and their clinical prognostic value. PLoS One 2013;8:e82314.
Dong F, Irshad H, Oh EY et al. Computational pathology to discriminate benign from malignant intraductal proliferations of the breast. PLoS One 2014;9:e114885.
Cooper LA, Kong J, Gutman DA et al. Novel genotype–phenotype associations in human cancers enabled by advanced molecular platforms and computational analysis of whole slide images. Lab Invest 2015;95:366–376.
Cooper LA, Kong J, Gutman DA et al. Integrated morphologic analysis for the identification and characterization of disease subtypes. J Am Med Inform Assoc 2012;19:317–323.
Diaz G, Romero E . Micro-structural tissue analysis for automatic histopathological image annotation. Microsc Res Tech 2012;75:343–358.
Haggerty JM, Wang XN, Dickinson A et al. Segmentation of epidermal tissue with histopathological damage in images of haematoxylin and eosin stained human skin. BMC Med Imag 2014;14:7.
Kothari S, Phan JH, Osunkoya AO et al In Proceedings of the ACM Conference on Bioinformatics, Computational Biology and Biomedicine 218-225 (ACM, Orlando, Florida, 2012).
Janowczyk A, Chandran S, Singh R et al. High-throughput biomarker segmentation on ovarian cancer tissue microarrays via hierarchical normalized cuts. IEEE Trans Biomed Eng 2012;59:1240–1252.
Irshad H, Veillard A, Roux L et al. Methods for nuclei detection, segmentation, and classification in digital histopathology: a review-current status and future potential. IEEE Rev Biomed Eng 2014;7:97–114.
McCann MT, Ozolek JA, Castro CA et al. Automated histology analysis: opportunities for signal processing. IEEE Signal Process Mag 2015;32:78–87.
Falkeholm L, Grant CA, Magnusson A et al. Xylene-free method for histological preparation: a multicentre evaluation. Lab Invest 2001;81:1213–1221.
Martina JD, Simmons C, Jukic DM . High-definition hematoxylin and eosin staining in a transition to digital pathology. J Pathol Inform 2011;2:45.
Ruifrok AC, Johnston DA . Quantification of histochemical staining by color deconvolution. Anal Quant Cytol Histol 2001;23:291–299.
Kothari S, Phan JH, Moffitt RA et al Automatic batch-invariant color segmentation of histological cancer images, in Biomedical Imaging: From Nano to Macro, 2011 IEEE International Symposium on, pp 657–660, March 30 2011-April 2 2011; doi:10.1109/ISBI.2011.5872492.
Pham N-A, Morrison A, Schwock J et al. Quantitative image analysis of immunohistochemical stains using a CMYK color model. Diagn Pathol 2007;2:8.
Khan AM, El-Daly H, Rajpoot NM . A Gamma-Gaussian mixture model for detection of mitotic cells in breast cancer histopathology images, in Pattern Recognition (ICPR), 2012 21st International Conference on, pp 149-152, 11-15 November 2012.
Veta M, Huisman A, Viergever MA et al Marker-controlled watershed segmentation of nuclei in H&E stained breast cancer biopsy images. In Biomedical Imaging: From Nano to Macro, 2011 IEEE International Symposium on, vol., no., pp.618-621, March 30 2011-April 2 2011; doi:10.1109/ISBI.2011.5872483.
Veta M, van Diest PJ, Kornegoor R et al. Automatic nuclei segmentation in H&E stained breast cancer histopathology images. PLoS One 2013;8:e70221.
Khan AM, Rajpoot N, Treanor D et al. A non-linear mapping approach to stain normalisation in digital histopathology images using image-specific colour deconvolution. IEEE Trans Biomed Eng 2014;61:6.
Hoffman RA, Kothari S, Wang MD . Comparison of normalization algorithms for cross-batch color segmentation of histopathological images. Conf Proc IEEE Eng Med Biol Soc 2014;2014:194–197.
Rodic N, Sharma R, Sharma R et al. Long interspersed element-1 protein expression is a hallmark of many human cancers. Am J Pathol 2014;184:1280–1286.
Fabbri R, Costa LDF, Torelli JC et al. 2D Euclidean distance transform algorithms: A comparative survey. ACM Comput Surv (CSUR) 2008;40:2.
Wienert S, Heim D, Saeger K et al. Detection and segmentation of cell nuclei in virtual microscopy images: a minimum-model approach. Sci Rep 2012;2:503.
Drelie Gelasca E, Obara B, Fedorov D et al. A biosegmentation benchmark for evaluation of bioimage analysis methods. BMC Bioinform 2009;10:368.
Otsu N . A threshold selection method from gray-level histograms. Automatica 1975;11:23–27.
Powers DMW . Evaluation: from precision, recall and F-meausure to ROC, informedness, markedness & correlation. J Mach Learn Technol 2011;2:37–63.
Macenko M, Niethammer M, Marron JS et al. A method for normalizing histology slides for quantitative analysis. 2009 IEEE International Symposium on Biomedical Imaging: From Nano to Macro, vols 1 and 2, 2009, pp 1107–1110.
Chen JM, Qu AP, Wang LW et al. New breast cancer prognostic factors identified by computer-aided image analysis of HE stained histopathology images. Sci Rep 2015;5:10690.
Uhlen M, Oksvold P, Fagerberg L et al. Towards a knowledge-based Human Protein Atlas. Nat Biotechnol 2010;28:1248–1250.
Bouzin C, Lamba Saini M, Khaing KK et al. Digital pathology: elementary, rapid and reliable automated image analysis. Histopathology 2015; http://onlinelibrary.wiley.com/doi/10.1111/his.12867/abstract.
Louis DN, Gerber GK, Baron JM et al. Computational pathology: an emerging definition. Arch Pathol Lab Med 2014;138:1133–1138.
Roth KA, Almeida JS . Coming into focus: computational pathology as the new big data microscope. Am J Pathol 2015;185:600–601.
Acknowledgements
We acknowledge funding from the National Cancer Institute (U54CA143868) and American Heart Association (12POST12050638).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website
The authors propose a novel non-linear tissue-component discrimination method to automatically register the color space of histopathology images and visualize individual tissue components, independent of color differences between images. They determine that this method is objective, robust and effective, and can be easily implemented in the emerging field of computational pathology.
Supplementary information
Rights and permissions
About this article
Cite this article
Sarnecki, J., Burns, K., Wood, L. et al. A robust nonlinear tissue-component discrimination method for computational pathology. Lab Invest 96, 450–458 (2016). https://doi.org/10.1038/labinvest.2015.162
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2015.162