Abstract
Breast carcinoma cells with the CD44+/CD24low phenotype have been reported to exhibit ‘cancer stem cell’ (CSC) characteristics on the basis of their enhanced tumorigenicity and self-renewal potential in immunodeficient mice. We used immunohistochemistry to study the expression of these proteins in whole tissue sections of human breast carcinoma. We found that the fraction of CD44v6+ cells is higher in estrogen receptor-positive carcinomas after neoadjuvant chemotherapy. We also performed double immunohistochemistry for CD44v6 and for the proliferation marker Ki67. We found that the relative number of quiescent carcinoma cells is higher in the CD44v6+ population than in the CD44v6− population in specific carcinoma subtypes. We then used quantum dots and spectral imaging to increase the number of antigens that could be visualized in a single tissue section. We found that anti-CD44v6 and CD24 antibodies that were directly conjugated to quantum dots retained their ability to recognize antigen in formalin-fixed, paraffin-embedded tissue sections. We then performed triple staining for CD44v6, CD24 and Ki67 to assess the proliferation of each sub-population of breast carcinoma cells. Our results identify differences between CD44v6-positive and CD44v6-negative breast carcinoma cells in vivo and provide a proof of principle that quantum dot-conjugated antibodies can be used to study specific sub-populations of cancer cells defined by multiple markers in a single tissue section.
Similar content being viewed by others
Main
Human tumors exhibit intra-tumoral heterogeneity at the genetic and epigenetic level. Functional heterogeneity between defined sub-populations of tumor cells has also been described in xenotransplantation studies. Numerous studies have used cell surface markers to isolate sub-populations of cancer cells, which exhibit enhanced tumorigenicity and self-renewal potential on injection into immunodeficient mice.1, 2 These sub-populations typically exhibit two distinct properties. First, they are more tumorigenic in mice on both primary inoculation and serial transplantation. Second, they appear to recapitulate the morphologic and immunophenotypic diversity of the tumor from which they were isolated. Cells exhibiting such properties have often been termed ‘cancer stem cells’ (CSCs). However, it is not yet known whether the behavior of these cells in mice recapitulates their behavior in humans.
Putative breast CSCs have been reported to reside in the CD44+CD24low fraction of a subset of human breast carcinomas.3, 4 CSCs have been hypothesized to preferentially survive cytotoxic chemotherapy by multiple possible mechanisms, including increased quiescence or expression of drug transporters. Two prospective studies of breast cancer patients receiving neoadjuvant chemotherapy have used flow cytometry to show that the percentage of CD44+CD24low cells was higher in post-chemotherapy tumor than in pre-treatment biopsy.5, 6 Preferential survival of CD44+CD24low cells after DNA damage has also been reported in cultured breast carcinoma cell lines.7, 8 The actual mechanisms by which this sub-population is more resistant to DNA damage remain to be elucidated.
Although ex vivo studies are essential for studying functional differences between defined subsets of cancer cells, there are limitations in the interpretation of these studies.9 It is difficult to know the extent to which the behavior of cells in mice or in tissue culture reflects their behavior in humans at primary or metastatic sites. Therefore, it will be essential to test hypotheses generated by ex vivo studies in primary, uncultured human tumors. For example, CD44+ and CD24+ cells freshly isolated from breast carcinomas exhibit significantly different gene expression profiles and distinct genetic changes.10
We sought to analyze the expression of markers associated with CSC behavior by using immunohistochemistry in primary breast carcinomas. It is difficult to visualize more than two antigens in one tissue section by traditional immunohistochemistry. We therefore used quantum dots and spectral unmixing to increase the number of antigens that could be identified in a single tissue section. Quantum dots are fluorescent semiconductor nanocrystals that have a constant excitation wavelength and a narrow emission spectrum that is proportional to their size.11 Quantum dots can be either directly conjugated to antibodies and oligonucleotide probes, or indirectly linked by strategies, such as the streptavidin–biotin interaction. Several antigens or mRNAs can then be assessed in a single tissue section using multiple quantum dots with different emission spectra.12, 13, 14 The signals from the quantum dots are deconvoluted by spectral unmixing to generate a composite image of each signal. Spectral unmixing also separates autofluorescence from quantum dot signals, an important step when studying formalin-fixed, paraffin-embedded (FFPE) samples. We have used traditional and quantum dot-mediated immunohistochemistry to define the properties of specific subsets of breast carcinoma cells in their native environment.
MATERIALS AND METHODS
Primary Tumor Samples
FFPE tumors were obtained from the archives of the Department of Pathology at the Brigham and Women's Hospital. In total, breast carcinomas from 48 different patients were evaluated, and 17 of these included pre- and post-chemotherapy material. Studies were carried out in accordance with protocols approved by the Institutional Review Board.
Immunohistochemistry
Four-micron thick sections of the FFPE tissue were passed through xylenes and graded alcohols and rehydrated. Heat-induced antigen retrieval was performed in a microwave for 15 min with 1 mM of EDTA (pH 8) for Id1 and Id3 antibodies or with 10 mM of citrate solution (pH 6) for all other antibodies. After antigen retrieval, peroxidase (Dako, Denmark), avidin (Vector, Burlingame, CA, USA), biotin (Vector) and protein (Dako) blocking were performed sequentially. Sections were incubated with primary antibody overnight at 4°C (for CD24) or for 1–2 h at room temperature (all others). Sections were then incubated with a biotinylated secondary antibody (Biogenex, San Ramon, CA, USA) for 20 min followed by HRP-conjugated streptavidin (Biogenex). The enzymatic activity was detected using DAB or VIP substrates (Vector) and slides were counterstained with hematoxylin or methyl green. For double immunohistochemistry, slides were transferred from chromagen to the peroxidase-blocking step and the protocol was repeated with another primary antibody. For quantum dot-conjugated antibodies, rehydration and antigen retrieval were performed as described above. Slides were then incubated with a protein blocking solution followed by primary antibody overnight at 4°C . The next day, slides were counterstained with Hoechst and mounted with Vectashield (Vector). For indirect detection of antibodies with quantum dots, peroxidase blocking was eliminated from the standard protocol and streptavidin-coated quantum dots (Invitrogen, Carlsbad, CA, USA) were substituted for HRP streptavidin.
Antibodies
Primary antibodies to the following proteins were used for immunohistochemistry: CD44, mouse monoclonal 156-3C11, 1:100 (LabVision, Fremont, CA, USA); CD44v6, mouse monoclonal VFF18, 1:400 (Millipore, Billerica, MA, USA); CD24, mouse monoclonal, 1:100 (LabVision); Ki67, rabbit polyclonal, 1:2000 (Vector); ER, rabbit polyclonal HC-20, 1:100 (Santa Cruz Biotechnology, Santa Cruz, CA, USA); PR, rabbit monoclonal SP2, 1:200 (LabVision); ALDH1, mouse monoclonal, 1:100 (BD Biosciences, San Jose, CA, USA); and Id1 and Id3, rabbit monoclonal, 1:50 (CalBioreagents, San Mateo, CA, USA). CD44v6 and CD24 antibodies were used at concentrations of 1:50 to 1:25 after direct conjugation to quantum dots.
Quantum Dot-Antibody Conjugation
Primary antibodies were conjugated to quantum dots using the Qdot antibody conjugation kit (Invitrogen) following the manufacturer's protocol. Antibodies were denatured with dithiothreitol, desalted and conjugated to activated quantum dots for 1 h at room temperature. The reaction was quenched with 2-mercaptoethanol, and the antibody quantum dot conjugate was purified in a phosphate-buffered saline by size exclusion chromatography.
Spectral Imaging
Spectral imaging was performed as previously described.12, 14 The imaging system comprised a Leitz Diaplan fluorescence microscope (Leitz, Germany) with a 490-nm excitation filter and a CRI Nuance spectral analyzer (Cambridge Research and Instrumentation Inc., Woburn, MA, USA). The CRI Nuance analyzer contains a liquid crystal tunable filter (LCTF) and a CCD (charged-coupled device) camera. The LCTF allows only light of a certain wavelength to pass to the camera, which acquires images. For each slide, a series of images were collected at 5-nm wavelength intervals from 450 to 720 nm. Each image contains the complete spectral information for every pixel at that given wavelength. The analyzer aligns all images in a stack and creates a high-resolution three-dimensional optical profile, which can be saved to memory as a spectral ‘cube’. This cube has the dimensions x, y and wavelength. All spectral information is scaled to the brightest point within this cube. Analysis of the acquired spectral data is performed using a proprietary software program within the Nuance system. This program is based on a generic mathematical algorithm for spectral unmixing (least-squares fitting) and has been specifically modified for bioimaging. It allows the digital removal of autofluorescence from tissue and also that of other fluorescent markers, provided their spectral profiles are known. A spectral library comprising the spectra of tissue autofluorescence and that of each quantum dot used is created. Spectra of tissue autofluorescence were acquired from a negative control slide lacking fluorescent markers, and spectra of relevant QDs were acquired from QD crystals. The spectral library is then used in spectral unmixing, after which the separate spectral contributions to the data ‘cube’ are outputted as falsely colored intensity maps. These images represent the distribution of each of the quantum dots and autofluorescence in the tissue.
Quantitation of Immunostains
Tissue sections were first examined at low power to characterize the overall staining pattern and to identify representative areas for precise quantitation. Positive and negative cells were then manually counted at high power in multiple representative fields. Approximately 200–1500 cells were counted per tumor, depending on the amount of tissue present. For CD24, membranous staining was considered positive, but pure cytoplasmic staining was not. When evaluating triple stains, cells were manually scored in composite images in which the signal from each quantum dot had been assigned a different pseudo-color. Triple stains were also compared with traditional IHC (single stains) on the same tumor to corroborate that the pattern of staining observed with quantum dots was accurate. Normal tissue elements with known patterns of protein expression were used as internal positive controls whenever possible.
Statistical Analysis
A paired, two-sided Wilcoxon signed-rank test was used to assess significance. Differences were considered significant at P<0.05.
RESULTS
The Fraction of CD44v6+ Cells is Increased in ER-Positive Breast Carcinomas after Neoadjuvant Chemotherapy
Our initial immunostains for CD44 in breast carcinomas showed that CD44 was expressed in both carcinoma cells and in stromal cells, particularly infiltrating lymphocytes (Figure 1a, left). We therefore inquired whether an epithelial-specific isoform of CD44 could be used to enable a more accurate evaluation of the fraction of CD44+ carcinoma cells in tissue sections. We found that CD44v6 is expressed in both normal breast ducts (predominantly basal cells and occasional luminal cells) and in breast carcinoma in a pattern similar to that of CD44 (Figure 1a, middle). However, the majority of stromal cells were negative for CD44v6. We therefore proceeded with our analysis using a CD44v6-specific antibody.
Increase in CD44v6+ fraction after neoadjuvant chemotherapy in ER+ breast carcinomas. (a) Detection of CD44, CD44v6 and CD24 in normal epithelium (top row) and in invasive ductal carcinoma (bottom row). CD44 (left) and CD44v6 (middle) are both expressed in the basal layer of normal breast epithelium and in a subset of carcinoma cells. However, CD44 is expressed in the stroma and inflammatory cells, whereas CD44v6 is largely confined to the epithelial compartment. Original magnification: × 200; scale bars: 50 μm. (b) Immunostains for CD44v6 in ER-positive carcinoma (top) and triple negative carcinoma (bottom). Left panels: pre-chemotherapy biopsy. Right panels: post-chemotherapy mastectomy. Original magnification: × 100; scale bars: 100 μm. (c) Quantitation of the percentage of CD44v6-positive carcinoma cells before and after chemotherapy in ER+ (n=12) and triple negative breast tumors (n=5). *P=0.004.
To test whether CD44v6+ breast carcinoma cells are increased after chemotherapy, we collected a series of pre-chemotherapy biopsies and matched resections taken from the same patient after neoadjuvant chemotherapy. We collected tumors from 17 breast cancer patients who had received neoadjuvant chemotherapy, including 12 patients with ER+ carcinomas (7 invasive ductal and 5 invasive lobular carcinoma) and 5 with ‘triple negative’ carcinomas (negative for ER, PR and Her2 expressions). We did not identify enough patients with Her2-amplified carcinomas receiving neoadjuvant chemotherapy to include in this study. We used immunohistochemistry and manual counting to quantitate the percentage of CD44v6+ carcinoma cells in biopsies taken before chemotherapy and in subsequent mastectomy specimens. In six cases (three ER+ and three triple negative), the only pre-chemotherapy biopsy available was from a sentinel (axillary) lymph node. In these cases, we compared these biopsies with other axillary lymph node metastases resected during mastectomy.
In ER+ invasive carcinomas (both ductal and lobular), the fraction of CD44v6+ carcinoma cells was higher in the post-chemotherapy specimen than in pre-treatment biopsy in 9 out of 12 cases (Figure 1b). In the other three cases, the fraction of CD44v6+ carcinoma cells was essentially the same before and after chemotherapy, and a significant decrease in the fraction of CD44v6+ cells was not observed. Quantitation of CD44v6 expression showed that the percentage of positive cells increased significantly from 62% before chemotherapy to 85% after chemotherapy (P=0.004, Figure 1c). Importantly, this difference is significant even if the three cases involving comparisons between lymph node biopsies are excluded (P=0.015, data not shown).
In contrast to the ER+ breast carcinomas, we saw no difference in the CD44v6 expression in triple negative tumors before and after chemotherapy. In general, levels of CD44v6 were relatively high in untreated tumors, consistent with a previous report15 and did not change significantly after chemotherapy (Figure 1b and c). We therefore assessed other proteins whose expression has been reported to be associated with self-renewal in triple negative breast carcinomas, such as ALDH1, Id1 and Id3.4, 16 Expression of these proteins was found in triple negative carcinomas (Supplementary Figure 1), typically in a small subset of cells, but there was no detectable increase in the number of positive carcinoma cells after chemotherapy.
We also assessed CD24 expression in all 17 neoadjuvant cases. CD24+ cells can be purified from normal breast epithelium and they display a luminal phenotype.10 However, CD24 is only focally detectable in normal breast epithelium by immunohistochemistry on the FFPE tissue.17 We therefore optimized CD24 immunostaining using the fallopian tube epithelium as a positive control (Figure 1a, top right), which exhibited a consistent apical membrane positivity.18 Under these conditions, we found that the cellular localization of CD24 staining varied substantially in breast carcinomas. In some tumors, CD24 staining was predominantly membranous, either in a circumferential (Figures 1a and 2c, left) or in an apical distribution (Figure 2c, right). In other tumors, CD24 staining was diffusely cytoplasmic (data not shown), which has been observed by others.15, 17 Previous functional studies of CD44+/CD24− cells have used fluorescence-activated cell sorting, which assesses surface protein expression; hence, the relevance of diffuse CD24 positivity is unclear. The variable pattern of immunostaining precluded precise quantitation of the percentage of CD24-positive cells, but there was no obvious difference in levels of membranous CD24 between pre- and post-chemotherapy specimens. These data suggest that further studies are needed to determine how the CD24 expression in live cells correlates with the results of immunostaining.
Increased quiescence in CD44v6+ carcinoma cells in specific subtypes of breast carcinoma. (a) Double immunostain for Ki67 (brown, nuclear) and CD44v6 (purple, membranous). Left panel: normal breast epithelium. Middle and right panels: invasive ductal carcinoma (Her2 amplified and triple negative, respectively). Original magnification: × 400; scale bars: 25 μm. (b) Quantitation of Ki67 rate in CD44v6+ and CD44v6− invasive carcinoma cells in four major subtypes of breast cancer. *P<0.01. **P=0.032. (c) Double immunostain for Ki67 (brown, nuclear) and CD24 (purple, membranous) in Her2-amplified breast carcinomas. Both circumferential (left) and purely apical (right) membranous staining of CD24 is noted. Original magnification: × 400; scale bars: 25 μm. (d) Quantitation of the Ki67 rate in CD24+ and CD24− invasive ductal carcinoma cells in Her2-amplified cases (n=6). *P=0.03.
CD44v6+ Cells are More Quiescent Than CD44v6− Cells in ER+ and HER2-Amplified Invasive Ductal Carcinomas
Stem cells are often more quiescent than progenitor cells in normal tissues. CSCs have been hypothesized to spend more time in quiescence compared with other cells in a tumor. There is evidence for this hypothesis in some malignancies, such as CML.19
To determine whether CD44v6+ cells are more quiescent than CD44v6− cells in breast carcinomas, we assembled an independent series of 31 additional untreated invasive breast carcinomas. We included the four major subtypes delineated by a routine pathological evaluation: ER+ ductal (n=11), Her2 amplified (n=7), triple negative (n=6) and ER+ invasive lobular (n=7). We performed single and double immunohistochemistry on whole tissue sections for CD44v6 and for the proliferation marker Ki67, a nuclear antigen that is expressed in cells that are in the cell division cycle (G1/S/G2/M), but not in quiescent cells. We then manually quantitated the Ki67 rate for CD44v6+ and CD44v6− cells in each tumor.
Evaluation of single stains showed that nearly all tumors in our study exhibited at least focal CD44v6 expression. Consistent with previous reports,15 the percentage of CD44v6+ cells in Her2-amplified tumors was generally lower than that in triple negative and ER+ tumors (data not shown). As expected, Ki67 rates correlated positively with tumor grade. Double stains showed that in normal breast ducts and lobules, CD44v6+ cells were predominantly Ki67 negative and were located along the basal aspect of the epithelium, whereas Ki67-positive cells were typically luminal and CD44v6− (Figure 2a, left). In contrast, breast carcinomas showed a range of double staining patterns. In some tumors, CD44v6 and Ki67 positivity was generally mutually exclusive, whereas in others, there were a large number of cells positive for both markers (Figure 2a, middle and right panels).
We observed significant differences in Ki67 rates between CD44v6-positive and CD44v6-negative sub-populations for some tumor types but not for others (Figure 2b). In Her2-amplified IDC (n=7), Ki67 rates were significantly lower in CD44v6+ cells than in CD44v6− cells (15 vs 32%, P=0.03). ER+ IDC (n=11) also showed a significant difference between CD44v6+ and CD44v6− fractions (13 vs 22%, P<0.005). In contrast, the Ki67 rate in CD44v6-positive and CD44v6-negative cells was nearly identical in invasive lobular carcinoma (n=7) and in triple negative IDC (n=6). These results suggest that in specific subtypes of human breast carcinoma, a greater fraction of CD44v6+ cells are quiescent than CD44v6− cells.
Previous reports have shown that mRNA levels of the estrogen receptor and progesterone receptor genes are lower in CD44+ breast cells (both normal and neoplastic) than in CD24+ cells.10 We therefore performed double immunohistochemistry for CD44v6 and either ER or PR in ER+ tumors. In normal ducts, we found that the percentage of hormone receptor positive cells was higher in the CD44v6-negative population, similar to our observations with Ki67 (Supplementary Figure 2). In contrast, we observed that both CD44v6-positive and CD44v6-negative carcinoma cells (ductal and lobular subtypes) exhibited high levels of ER and PR expressions (Supplementary Figure 2).
As previously noted, the pattern of CD24 immunostaining in breast carcinomas was more variable than that of CD44v6, with many tumors exhibiting cytoplasmic and/or membranous positivity. We found that most Her2-amplified tumors (86 or 6/7) exhibited at least focal membranous staining. Evaluation of the remaining tumors in this study (untreated resections or pre-chemotherapy biopsies) showed that the membranous CD24 expression could be identified in 67% (20/30) of ER-positive carcinomas and in 36% (4/11) of triple negative carcinomas (data not shown). We therefore restricted our quantitative analysis of CD24 and Ki67 expressions to the Her2-amplified subset.
We performed double immunohistochemistry for CD24 and Ki67 in Her2-amplified cases (n=6, Figure 2c), and quantitated the Ki67 rate in each sub-population. We found that the Ki67 rate in CD24+ cells was significantly lower (P=0.03) than in CD24− cells (15% vs 29%, Figure 2c). These data, in combination with the CD44v6/Ki67 results, raise the question of which carcinoma sub-population contains the highest fraction of cycling cells. We therefore turned to quantum dot-mediated multiplex immunohistochemistry to assess multiple markers in one tissue section.
Conjugation of Quantum Dots to CD44v6 and CD24 Antibodies
To link antibodies and quantum dots covalently, we treated antibodies with a strong reducing agent (dithiothreitol) and incubated them with activated quantum dots to allow disulfide bond formation. Antibodies to CD44v6 and CD24 were conjugated to quantum dots with an emission maxima of 655 and 605 nm, respectively. We first used the normal tissue to determine whether conjugated antibodies retained the ability to recognize their cognate antigens in the FFPE tissue. Incubation of normal breast epithelium with CD44v6-655-conjugated antibody resulted in membranous staining of basal cells, whereas most luminal cells were negative (Figure 3a, left). This staining pattern was consistent with the results of traditional immunohistochemistry with this antibody (Figure 1a). The CD24-605-conjugated antibody exhibited an apical membranous staining pattern on the fallopian tube epithelium (Figure 3a, right), also consistent with traditional immunohistochemistry (Figure 1a). Single immunostains of breast carcinomas with CD44v6-655 and CD24-605 also yielded staining patterns similar to those observed in unconjugated antibodies (data not shown). We therefore conclude that reduction and quantum dot conjugation of antibodies to CD44v6 and CD24 leaves their ability to recognize antigen intact.
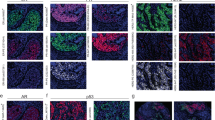
Detection of CD44v6 and CD24 in normal epithelium and in invasive ductal carcinoma by quantum dot-conjugated antibodies. (a) Left: breast epithelium stained with a CD44v6-655-nm quantum dot-conjugated antibody. Right: fallopian tube epithelium stained with CD24-605-nm quantum dot-conjugated antibody. Images represent live view of tissue section (before deconvolution). Green: Hoechst counterstain. (b) Dual stain of Her2-amplified carcinoma with CD44v6-655- and CD24-605-conjugated antibodies. Left panel: live view. Middle panels: signal from individual quantum dots after spectral unmixing. Right panel: composite image. Red: CD44v6, green: CD24, blue: Hoechst counterstain. (c) Composite images showing the heterogeneity of CD44v6 and CD24 expression in a single Her2-amplified tumor. Left panel: area containing primarily CD44v6+/CD24− (red) cells with occasional CD44v6+/CD24+ (yellow) and CD44v6−/CD24+ cells (green). Middle: CD44v6+/CD24+ cells (yellow) with occasional CD44v6−/CD24+ cells (green, arrow) and CD44v6+/CD24− cells (red, arrowhead). Right: CD44v6+/CD24− cells (red) and CD44v6−/CD24− cells. Blue: Hoechst counterstain. Original magnification: × 200 for all images; scale bars: 50 μm.
Thereafter, we performed double staining of breast carcinomas with CD44v6-655- and CD24-605-conjugated antibodies. As expected, the wavelengths emitted by the two quantum dots were too similar in color to be distinguished during live microscopy (Figure 3b, left panel). However, the signals from each quantum dot could be readily distinguished after spectral unmixing, which yielded grayscale images for each signal (Figure 3b, middle panels) and a pseudo-colorized composite image for both signals (Figure 3b, right panel). Using this technique, we were able to identify visually all four possible subsets of breast carcinoma cells (double positive, single positive for each antibody and double negative, Figure 3c). Quantitation of signal intensity using the Nuance software allowed for confirmation of visual assessments, and also enabled us to assign a numerical cutoff value for a cell to be considered positive or negative for each marker. We used this double stain to inquire whether a specific distribution of these markers could be observed in tumors. In some cases, the distribution of cell sub-populations within the tumor appeared random. In others, the CD24 expression was the highest in the central portion of the tumor cell population, whereas the CD44v6 expression was the highest in the peripheral tumor cells adjacent to the stroma (eg, Figure 4a, left panel). This pattern partially recapitulates the normal structure of the breast epithelium.
Quantitation of the Ki67 rate in sub-populations of breast carcinoma cells identified by quantum dot-conjugated antibodies. (a) Composite images of two Her2-amplified invasive ductal carcinomas triple stained for Ki67 (yellow, nuclear), CD44v6 (red) and CD24 (green). Ki67 was detected indirectly by biotinylated secondary antibody and streptavidin-conjugated 565-nm quantum dots. CD44v6 and CD24 were detected by antibodies directly conjugated to 655- and 605-nm quantum dots, respectively. Original magnification: × 200 for all images; scale bars: 50 μm. (b) Quantitation of Ki67 rates in four sub-populations defined by CD44v6 and CD24 expressions in Her2-amplified invasive ductal carcinomas (n=5). Although the double negative population exhibited the highest Ki67 rate, this difference did not reach statistical significance.
Triple Staining of HER2-Amplified Tumors for CD44v6, CD24 and Ki67 with Quantum Dots
Given that the four sub-populations of breast carcinoma cells could be identified using quantum dot-conjugated antibodies, we attempted to quantitate the Ki67 rate in each sub-population in a series of Her2-amplified tumors. We first conjugated a Ki67-specific antibody to 565-nm quantum dots. Although we obtained a nuclear signal from this conjugate in some control tissues, its intensity was not high enough in breast carcinomas to be readily visualized. We therefore used a streptavidin-linked 565-nm quantum dot to detect Ki67 indirectly and to amplify the quantum dot signal. This strategy enabled us to detect a specific nuclear Ki67 signal in breast carcinomas stained with Ki67 alone (data not shown) or in combination with CD44v6-655 and CD24-605 (Figure 4a).
We performed triple staining for Ki67, CD44v6 and CD24 on five Her2-amplified invasive ductal carcinomas and manually scored the Ki67 rate for all four sub-populations of carcinoma cells (eight possible combinations of protein expression). We found that the overall Ki67 rate was lower by this method than by standard immunohistochemistry, suggesting that this technique underestimates the absolute number of proliferating cells. Nevertheless, the proliferation rates of the tumors relative to each other were similar by either technique.
Quantitation of the Ki67 rate showed a trend in which the highest average proliferation rate was found in the CD44v6−/CD24− (double negative) sub-population, although this did not reach statistical significance (Figure 4b). When evaluating individual tumors, we found variations in the proliferation rates of the specific sub-populations. In one tumor, the Ki67 rate was very similar in the double negative and CD44v6+/CD24− populations. In another, the Ki67 rate was similar in the double negative and CD24+ populations and was the lowest in CD44v6+/CD24− cells. Thus, although the Ki67 rate was consistently high in the double negative population, it was sometimes just as high in other sub-populations. Evaluation of a larger cohort of tumors and functional studies of representative Her2-amplified breast carcinoma cell lines will be required to determine the significance of these variations in cell-cycle distribution.
DISCUSSION
The CD44+ fraction of breast cancer cells has been reported to contain most, if not all, of the self-renewing cells within the tumor.3, 4 Although these studies have of necessity been carried out in immunocompromised mice, it has been proposed that the CD44+ fraction also exhibits distinct properties in primary human tumors. We therefore sought to evaluate CD44+ and CD44− cells in tissue sections by traditional and quantum dot-mediated immunohistochemistry. Using an antibody to the epithelial-specific isoform CD44v6, we found significant differences between cellular sub-populations, but these differences were only apparent in specific pathologically defined types of breast carcinoma.
In normal tissues, specific cell types respond differently to chemotherapy and radiation. In the prostate, eg, radiation leads to atrophy and death of the luminal cells, whereas basal cells survive and even become hyperplastic.20 It has also been proposed that CSCs are differentially affected by cancer therapies. If CD44v6+ cells preferentially survive chemotherapy, the percentage of CD44v6+ cells should increase at some time points after treatment. We therefore examined the percentage of CD44v6+ cells in matched breast carcinomas for which both pre- and post-chemotherapy material was available. We found that in ER+ carcinomas (n=12), but not in triple negative carcinomas (n=5), the percentage of CD44v6+ cells was higher in post-chemotherapy tumors. These results are largely consistent with two other reports which used flow cytometry to analyze breast carcinomas before and after chemotherapy.5, 6 However, one of these reports found that the CD44+/CD24− fraction increased in triple negative carcinomas after chemotherapy, in contrast with our findings. This could be a result of the higher number of cases used in that study. It could also be the consequence of different means of assessing protein expression, or even be related to different chemotherapy protocols used in these patients. Although these results suggest preferential survival of CD44v6+ carcinoma cells, they could also be explained by other models. For example, chemotherapy itself might induce CD44 expression. If true, this would have to be a stable induction, as mastectomies are generally performed weeks after neoadjuvant treatment. DNA damage can lead to cytokine induction,21 and some cytokines can induce CD44 expression,22 suggesting one possible mechanism for such a model. In contrast, p53, which is activated by cytotoxic chemotherapy, can repress CD44 expression in human mammary epithelial cells.23 Thus, DNA damage has the potential to modulate CD44 levels through multiple pathways.
We found differences between breast carcinoma subtypes when evaluating the Ki67 rate in CD44v6-positive and CD44v6-negative cells. No significant difference between the Ki67 rate of CD44v6-positive and CD44v6-negative cells was observed in triple negative IDC or in invasive lobular carcinoma (Figure 2b). In contrast, the Ki67 rate was significantly lower in the CD44v6-positive cells of ER+IDC and Her2-amplified IDC. These results could mean that CD44v6+ cells spend a greater fraction of their time in quiescence, but only in specific types of breast cancer. These results could also be interpreted to mean that CD44v6 is a cell-cycle-regulated protein and that its levels fluctuate as cells transit between the cell cycle and quiescence. For example, levels of CD133, another reported marker of CSCs, fluctuate with the cell cycle in some cultured cell lines.24 It will be important to test this possibility in breast cancer cells that reflect the different subtypes evaluated in this study.
To analyze more than two markers at once, we directly conjugated CD44v6 and CD24 antibodies to quantum dots and used spectral unmixing to identify individual signals from each quantum dot. We found that conjugated CD44v6 and CD24 antibodies retained their ability to recognize their cognate antigens in both normal tissues (Figure 3a) and in tumors. The application of both antibodies allowed us to visualize all four possible cell populations defined by these two surface antigens (Figures 3b and c).
We attempted to use quantum dot-conjugated antibodies to further define which sub-populations were the most proliferative in breast carcinoma. We focused specifically on Her2-amplified cases because both CD44v6 and CD24 were readily detectable in most of these tumors. We first attempted to use a directly conjugated Ki67 antibody, but found that it was sensitive enough to detect its antigen in some tissue types (ie, germinal centers of lymph node), but not in breast carcinomas. We therefore used streptavidin-coated quantum dots to amplify the signal and detect Ki67 indirectly. With this strategy, we could readily detect Ki67 alone and in combination with CD44v6 and CD24 (Figure 4a). Nevertheless, the Ki67 rate was not as high as it was for the same tumors assessed by traditional immunohistochemistry. This indicates that additional technical improvements will be needed to reach maximum sensitivity for detection of certain antigens by quantum dots.
Using our triple stain, we manually quantitated the Ki67 rate of each sub-population in Her2-amplified IDC (Figure 4b). The spectral imaging software provides a quantitative assessment of the intensity of each quantum dots. This allowed us to set objective criteria for positivity for each marker and, removed much of the potential subjectivity that could be associated with manually counting multiple signals per cell. Although the differences did not reach significance, the double negative population on average contained the highest Ki67 rate, whereas the Ki67 rate in the putative CSC fraction (CD44v6+/CD24−) was lower in most cases. However, the analysis of individual cases showed that in one tumor, the Ki67 rate was virtually identical between these two populations. Additional tumors will need to be evaluated to determine how often the putative CSC fraction actually has a lower Ki67 rate than the bulk of the tumor. At this time, manual quantitation is labor intensive, but automation of this analysis is likely to become feasible. We expect that automation will bring the technology closer to the realm of flow cytometry with respect to the number of cells that can be analyzed per sample and the speed with which the analysis is carried out.
In summary, our results suggest that CD44v6+ cells have distinct properties in primary human breast carcinomas, but only in specific subtypes. Further work with cell lines and mouse models of each subtype will shed further light on some of these differences. Our results with quantum dot-conjugated cell surface markers provide a proof of principle and a starting point for additional analysis of specific sub-populations in unmanipulated primary human tumors.
References
Cho RW, Clarke MF . Recent advances in cancer stem cells. Curr Opin Genet Dev 2008;18:48–53.
Shipitsin M, Polyak K . The cancer stem cell hypothesis: in search of definitions, markers, and relevance. Lab Invest 2008;88:459–463.
Al-Hajj M, Wicha MS, Benito-Hernandez A, et al. Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci USA 2003;100:3983–3988.
Ginestier C, Hur MH, Charafe-Jauffret E, et al. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell 2007;1:555–567.
Li X, Lewis MT, Huang J, et al. Intrinsic resistance of tumorigenic breast cancer cells to chemotherapy. J Natl Cancer Inst 2008;100:672–679.
Yu F, Yao H, Zhu P, et al. Let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 2007;131:1109–1123.
Fillmore CM, Kuperwasser C . Human breast cancer cell lines contain stem-like cells that self-renew, give rise to phenotypically diverse progeny and survive chemotherapy. Breast Cancer Res 2008;10:R25.
Phillips TM, McBride WH, Pajonk F . The response of CD24(-/low)/CD44+ breast cancer-initiating cells to radiation. J Natl Cancer Inst 2006;98:1777–1785.
Fillmore C, Kuperwasser C . Human breast cancer stem cell markers CD44 and CD24: enriching for cells with functional properties in mice or in man? Breast Cancer Res 2007;9:303.
Shipitsin M, Campbell LL, Argani P, et al. Molecular definition of breast tumor heterogeneity. Cancer Cell 2007;11:259–273.
Alivisatos AP, Gu W, Larabell C . Quantum dots as cellular probes. Annu Rev Biomed Eng 2005;7:55–76.
Byers RJ, Di Vizio D, O’Connell F, et al. Semiautomated multiplexed quantum dot-based in situ hybridization and spectral deconvolution. J Mol Diagn 2007;9:20–29.
Fountaine TJ, Wincovitch SM, Geho DH, et al. Multispectral imaging of clinically relevant cellular targets in tonsil and lymphoid tissue using semiconductor quantum dots. Mod Pathol 2006;19:1181–1191.
Tholouli E, Hoyland JA, Di Vizio D, et al. Imaging of multiple mRNA targets using quantum dot based in situ hybridization and spectral deconvolution in clinical biopsies. Biochem Biophys Res Commun 2006;348:628–636.
Honeth G, Bendahl PO, Ringner M, et al. The CD44+/CD24- phenotype is enriched in basal-like breast tumors. Breast Cancer Res 2008;10:R53.
Gupta GP, Perk J, Acharyya S, et al. ID genes mediate tumor reinitiation during breast cancer lung metastasis. Proc Natl Acad Sci USA 2007;104:19506–19511.
Kristiansen G, Winzer KJ, Mayordomo E, et al. CD24 expression is a new prognostic marker in breast cancer. Clin Cancer Res 2003;9:4906–4913.
Kristiansen G, Denkert C, Schluns K, et al. CD24 is expressed in ovarian cancer and is a new independent prognostic marker of patient survival. Am J Pathol 2002;161:1215–1221.
Graham SM, Jorgensen HG, Allan E, et al. Primitive, quiescent, Philadelphia-positive stem cells from patients with chronic myeloid leukemia are insensitive to STI571 in vitro. Blood 2002;99:319–325.
Zhou M, Magi-Galluzzi C, Epstein JI . Genitourinary Pathology. Elsevier: Philadelphia, PA, 2007.
Biswas S, Guix M, Rinehart C, et al. Inhibition of TGF-beta with neutralizing antibodies prevents radiation-induced acceleration of metastatic cancer progression. J Clin Invest 2007;117:1305–1313.
Vincent T, Mechti N . IL-6 regulates CD44 cell surface expression on human myeloma cells. Leukemia 2004;18:967–975.
Godar S, Ince TA, Bell GW, et al. Growth-inhibitory and tumor-suppressive functions of p53 depend on its repression of CD44 expression. Cell 2008;134:62–73.
Jaksch M, Munera J, Bajpai R, et al. Cell cycle-dependent variation of a CD133 epitope in human embryonic stem cell, colon cancer, and melanoma cell lines. Cancer Res 2008;68:7882–7886.
Acknowledgements
We thank Joshua Rose and Stephen Finn for their assistance with quantum dots, Eyoung Shin for help with immunohistochemistry and Rebecca Gelman for her assistance with statistics. The work was funded by the Center for Molecular Oncologic Pathology and the Dana-Farber/Harvard Cancer Center SPORE in Breast Cancer.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website (http://www.laboratoryinvestigation.org)
Rights and permissions
About this article
Cite this article
Snyder, E., Bailey, D., Shipitsin, M. et al. Identification of CD44v6+/CD24− breast carcinoma cells in primary human tumors by quantum dot-conjugated antibodies. Lab Invest 89, 857–866 (2009). https://doi.org/10.1038/labinvest.2009.54
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2009.54
Keywords
This article is cited by
-
Comparison of mammosphere formation from stem-like cells of normal breast, malignant primary breast tumors, and MCF-7 cell line
Journal of the Egyptian National Cancer Institute (2022)
-
P-cadherin: a useful biomarker for axillary-based breast cancer decisions in the clinical practice
Modern Pathology (2017)
-
Regulation of stem cells-related signaling pathways in response to doxorubicin treatment in Hs578T triple-negative breast cancer cells
Molecular and Cellular Biochemistry (2015)
-
Cancer stem cells: The challenges ahead
Nature Cell Biology (2013)
-
Accurate Sensitivity of Quantum Dots for Detection of HER2 Expression in Breast Cancer Cells and Tissues
Journal of Fluorescence (2013)