Abstract
Distinction of oligodendrogliomas from other gliomas is clinically important, but the histologic diagnosis of oligodendroglioma has been a difficult and notoriously subjective task. Testing for loss of heterozygosity (LOH) on chromosomal arms 1p and 19q, the genetic signature of oligodendroglioma, has emerged as a useful, objective adjunct to the traditional histologic evaluation. However, LOH testing of glioma specimens has not yet been widely implemented, presumably because of a lack of a practical LOH assay. We describe a 1p and 19q LOH assay suitable for routine diagnostics. In contrast to traditional microsatellite-based LOH analysis, we show that detection of LOH is usually possible even without normal tissue or blood from the same patient. A small area of tumor on a single paraffin section is sufficient for the assay. The assay protocol consists of a one-step DNA extraction, multiplex PCR for microsatellites on 1p and 19q, and capillary electrophoresis of the PCR products. LOH is detected by analysis of the allelic patterns and by integration of data from multiple highly polymorphic microsatellites. In a validation study on 19 gliomas, the results were concordant with results obtained by established methods and correlated well with histologic diagnoses. Because only a paraffin section is required, the pathologist can perform both the traditional histopathologic evaluation and this supporting molecular assay from the material at hand.
Similar content being viewed by others
Introduction
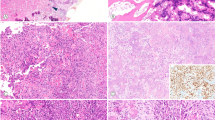
Accurate diagnosis of an oligodendroglioma is important because oligodendrogliomas, unlike astrocytomas, are often chemosensitive (Cairncross et al, 1998). Oligodendrogliomas are also associated with a longer average patient survival than astrocytomas (Leighton et al, 1997). However, the histologic distinction between oligodendroglioma and astrocytoma is often highly subjective (Fig. 1), and there has been significant interobserver variation in the diagnosis (Coons et al, 1997).
Because loss of heterozygosity (LOH) on 1p (short arm of chromosome 1) and 19q (long arm of chromosome 19) has clinical prognostic value and is strongly associated with classic histologic features of oligodendroglioma, testing for LOH is useful in the diagnosis. Specifically, LOH on 1p predicts that the tumor is chemosensitive, and LOH on both 1p and 19q predicts an extended duration of chemotherapeutic response and patient survival (Cairncross et al, 1998; Ino et al, 2001; Smith et al, 2000). A relationship between LOH and morphologic features of oligodendroglioma is suggested by the finding that increasing the stringency of the histologic criteria in the diagnosis of oligodendroglioma leads to a higher proportion of oligodendrogliomas with LOH. When an oligodendroglioma is defined based on either stringent histologic criteria or a unanimous diagnosis by four neuropathologists, more than 85% of oligodendrogliomas have LOH on 1p and 19q (Burger et al, 2001; Sasaki et al, 2002; Ueki et al, 2002).
The objective of the present study was to develop a 1p and 19q LOH assay that would overcome the common limitations of small sample size and lack of normal and frozen tissue, thereby allowing the routine analysis of paraffin-embedded tissue. Such an assay would also be ideal for retrospective studies using archival specimens. To meet these requirements, we developed and validated an assay based on PCR amplification of highly polymorphic microsatellite repeats and subsequent capillary electrophoresis. This assay has been recently approved for clinical use at our institution.
Results
Performance of the Assay on Gliomas Dissected from Paraffin Sections
The reported assay uses multiplex PCR amplification of microsatellite loci followed by high-resolution PCR product sizing by capillary electrophoresis (Figs. 2 and 3, Table 1). The assay was used to successfully determine 1p and 19q LOH in 19 tumors previously assessed by other methods for this alteration. The reported assay uses formalin-fixed, paraffin-embedded tissue that is dissected from standard unstained histologic slides. After assay optimization using the first 10 tumors, the assay was used to successfully determine the 1p 19q status on the initial attempt for the remaining 9 tumors (Table 2).
An overview of the 1p and 19q LOH assay method and the suggested approach for interpreting results. The first and often the only PCR to be performed is a single-tube multiplex reaction with primers for three microsatellite loci on 1p (primer mix A). If two or more of the three amplified loci show a heterozygous allelic pattern, it is concluded that the tumor does not have LOH on 1p. Otherwise, more evidence for LOH is sought by two additional PCRs (primer mix B and C) containing primers for three loci on 19q and two additional loci on 1p.
Genomic tumor DNA for analysis was extracted from tumor areas (3 to 20 mm2) on 5-μm–thick unstained paraffin sections. The amount of genomic DNA isolated from these tumor regions was sufficient for at least two complete assays (six multiplex PCR reactions). During assay development, we found that larger or smaller areas of tissue did not work as consistently as those ranging from 3 to 20 mm2.
The unstained paraffin sections were used for DNA extraction within 2 weeks of cutting the sections. We also analyzed DNA extracted from unstained sections that had been cut several years earlier. DNA extracted from these slides produced interpretable results in most cases, although occasional artifacts were identified. Characteristic artifacts associated with the use of old slides included the apparent presence of three or more alleles, simulating microsatellite instability. The extra alleles were not observed when freshly cut sections and frozen tissue from the same tumors were assessed. The age of the paraffin block itself did not seem to matter; freshly cut sections from paraffin blocks as old as 12 years were analyzed successfully, and these results agreed with data generated from frozen tissue from the same tumor.
Allelic Patterns in Samples Without LOH
Inspection of the capillary electropherograms revealed that, as expected, two prominent alleles were identified for most microsatellite loci in the cases that had been determined not to have LOH on 1p and 19q by the reference methods (eg, Fig. 4, Case T250). The presence of two prominent alleles at a given locus was defined as the heterozygous pattern and indicated by “2” in Table 2. The peak height (intensity) of the longer allele was observed to always be less than that of the shorter allele, presumably because of PCR bias favoring complete replication of the shorter allele during the first PCR cycle. When the two alleles were separated in length by more than two nucleotides, the peak height of the longer allele ranged from 22% to 85% of the peak height of the shorter allele. When the two alleles were separated by exactly two nucleotides, the height of the longer allele was 26% to 60% of the height of the shorter allele (Fig. 5). As expected, some loci in samples without LOH showed evidence of only one allele, a finding consistent with homozygosity at that locus (ie, two copies of the same allele). This pattern, termed the single allele pattern, is indicated by “1” in Table 2.
Representative capillary electropherograms of two PCR-amplified microsatellites from an oligodendroglioma and an astrocytoma. Arrows point to alleles. The alleles of each locus are separated by size. The alleles of different loci are automatically shown in different panels based on the different-colored fluorescence associated with each forward primer and the corresponding PCR product. A locus with one allele can mean either homozygosity or LOH, whereas two alleles indicate heterozygosity. The lower-intensity peaks at −2 nucleotide intervals from each allele are stutter peaks. They confirm the specificity of the PCR, because they only occur when a dinucleotide repeat is amplified. The intensity of the longer allele is normally less than that of the shorter allele (22–85% of the height of the short allele). For T250 (right), the allelic ratios (height of the long allele/height of the short allele) are 0.45 and 0.29 at D1S226 and D1S162, respectively. Based on the results for these two loci and their heterozygosity scores, the likelihood of LOH on 1p in the oligodendroglioma is approximately 98% or 1−(1−0.84)*(1−0.86). However, this tumor also showed only one allele at three additional loci on 1p (Table 2), bringing the likelihood of LOH to 99.99%.
An example of the characteristic pattern of peaks obtained when the sample is heterozygous and the size of the two alleles differs by two nucleotides. The arrows points to the longer allele, whose stutter peaks are not visible as they overlap the shorter allele and its stutter peaks. The relative heights of the two alleles were not significantly affected by the number of PCR cycles. The allelic ratio is 0.59 after 35 PCR cycles and 0.61 after 42 PCR cycles.
Allelic Patterns in Samples with LOH
Three allelic patterns were observed in the tumors known to have LOH on 1p and 19q. In the first pattern, only one allele was seen (Fig. 4, Case T197). This pattern was indistinguishable from the single allele pattern observed in some loci of samples without LOH and was therefore also indicated by “1” in Table 2. The second pattern, termed LOH pattern A, consisted of a shorter allele (the allele measuring fewer nucleotides in length) with a relatively high peak and a longer allele with a diminutive peak (Fig. 6). The height of the longer allele was never more than 12% of the height of the shorter allele. Thus, there was no overlap between LOH pattern A and the heterozygous pattern seen in samples without LOH. To avoid overcalling LOH pattern A because of small nonspecific peaks, we required the diminutive long allele to have at least two shorter stutter peaks at −2 and −4 nucleotides, which are specific indicators of amplified dinucleotide repeats. In the third pattern, termed LOH pattern B, the intensity of the shorter allele was less than that of the longer allele (Fig. 7). LOH pattern B was also never seen unless LOH was present. In Table 2, LOH patterns A and B are represented by “L.”
In this oligodendroglioma, the diminutive long allele (arrows) likely represents contamination from normal brain tissue (eg, blood vessels), providing strong evidence for LOH even when no normal tissue is available for comparison. The relative heights of the two alleles were not significantly affected by the amount of template DNA or the number of PCR cycles. The allelic ratios are 0.09 (8 μl of DNA extract, 35 PCR cycles) and 0.10 (4 μl of DNA extract, 40 PCR cycles).
As demonstrated by comparison with normal brain in this example, the finding of a short allele (arrows) of lower intensity than the long allele was always associated with LOH. Thus, the short allele in this tumor sample likely represents contamination from normal brain or blood vessels. The allelic ratios were 1.91 and 0.50 in the tumor and normal brain, respectively.
Comparison with Normal Tissue Was Not Necessary for the Detection of LOH
The single allele pattern (only one allele present) was the only allelic pattern seen in both tumors with LOH and in tumors without LOH. Of the 19 gliomas analyzed, one tumor (T197) showed the single allele pattern at all five microsatellite loci on 1p (Table 2). In theory, all these five loci could represent homozygosity caused by two copies of the same allele rather than LOH. However, based on the heterozygosity scores of the individual alleles (Table 1), the likelihood of this being the case is only approximately 1 in 18,000 (0.005%). Thus, no normal tissue was required to conclude that this tumor has a high likelihood of LOH on 1p.
Among the nine tumors with LOH on 19q, there were three tumors that showed only the single allele pattern on all three 19q loci (T35, 77, and 246; Table 2). Based on the heterozygosity scores of the three microsatellites on 19q (Table 1), the likelihood of all three loci being homozygous without having LOH is 1 in 1020 (0.1%). Considering that 77% to 86% of gliomas with LOH on 1p also have LOH on 19q (Smith et al, 2000; Ueki et al, 2002) and that LOH on 19q only needs to be assessed when there is LOH on 1p, the likelihood of these tumors not having LOH on 19q drops to 1 in 4400 to 7300 (0.01–0.02%). Thus, even in these three cases, no normal tissue was required to conclude that these tumors had a high likelihood of LOH on 19q.
Relative Intensities of the Alleles Were Generally Unaffected by PCR Conditions
In most cases, the relative intensities (peak heights) of the alleles remained relatively constant from one experiment to another, regardless of the amount of template DNA or the number of PCR cycles (Figs. 5 and 6).
Assay Validation
The results of the assay correlated well with the results of the established (reference) methods (Table 2), with 18 of 19 tumors showing LOH of 1p and 19q by all methods. The single lesion showing discordant results is discussed below. To further validate the reported approach, we dissected and isolated DNA from histologically identifiable areas of normal or predominately normal brain tissue in six of nine cases that had shown evidence of LOH. These areas of normal brain were subjected to the same assay procedure that had been used for the areas of tumor. In all six cases, LOH was confirmed at one or more loci by detecting allelic imbalance (Skotheim et al, 2001) in tumor compared with normal brain (Fig. 7).
Discussion
In this study, we have developed and validated a microsatellite assay for the detection of 1p and 19q LOH in paraffin sections. Our data on 19 gliomas suggest that LOH can be detected by this approach even when no normal tissue is available for comparison (Table 2). In traditional microsatellite-based LOH assays, LOH is defined as the relative loss of an allele in tumor compared with normal tissue or blood from the same patient. In our approach, data from several highly polymorphic microsatellite loci are combined to conclude that there is a high likelihood of LOH when all of the microsatellite loci in a tumor show the presence of only one allele (single allele pattern) in tumor tissue. In addition, our results suggest that the allelic pattern consisting of a single prominent allele and a diminutive second allele, designated by us as the LOH pattern (“L” in Table 2), is only found when LOH is present.
Precise dissection of areas containing the highest tumor cell density in the tissue section was possible using the method of Turbett et al (1996), in which the areas of interest are covered with a starch-based glue, peeled off, and submitted for DNA extraction. The yield and specificity of the multiplex PCR reactions were increased by diluting the forward primers relative to the reverse primers (Berg et al, 2000). Use of capillary electrophoresis capable of multicolor fluorescent detection of the PCR products facilitated multiplexing of several primer pairs into a single PCR tube. Capillary electrophoresis also has significantly better size resolution than traditional solid gel-based methods, leading to more precise resolution and identification of the tumor tissue allelic pattern.
Of the 19 tumors assessed by the reported method and the reference methods, the results were discordant for only 1 tumor (T142, Table 2). In this case, the reference methods reported no LOH on 1p or 19q, whereas the reported assay identified LOH on 19q but not on 1p. The cause of the discrepancy is unknown, but a subsequent comparison of tumor and normal brain from the same patient demonstrated that the tumor sample used for the present assay does in fact have LOH on 19q (Fig. 7).
The microsatellites selected for this assay are located in the regions extending from 1p22 to 1p36.22 and from 19q13.31 to 19q13.32 (Fig. 2). LOH outside these regions would not be detected by this assay. However, this region on 1p was entirely involved by LOH in all of the 97 oligodendrogliomas with LOH on 1p analyzed in previous studies (Bello et al, 2000; Bigner et al, 1999; Smith et al, 1999). Similarly, the region on 19q assessed for LOH using the reported microsatellites was involved in all of the 59 oligodendrogliomas with combined 1p and 19q LOH identified in previous studies (Bigner et al, 1999; Smith et al, 1999).
In the present study, one tumor (T246) was found to have partial LOH on 1p, a finding confirmed by comparison with the allelic pattern derived from normal tissue dissected from this case (Table 2). In this tumor, heterozygosity was only preserved at one locus of eight assessed on 1p and 19q. In light of previous studies (Bello et al, 2000; Bigner et al, 1999; Smith et al, 1999), this tumor may represent a rare aberration from the usual extent of LOH on 1p in oligodendrogliomas. Although the clinical significance of this finding of partial LOH is not known, the tumor is probably best classified as having a high likelihood of clinically relevant LOH on 1p, considering that heterozygosity was preserved at only one locus and that the tumor was histologically an oligodendroglioma. Of note, the present assay strongly suggested partial LOH in this particular tumor (T246) even without comparison with normal tissue, because three microsatellite loci on 1p showed the allelic pattern designated as the LOH pattern (Table 2). Thus, the possibility of a partial LOH was not found to be a serious limitation in this material, even when no normal tissue was used for the analysis. In any case, caution is advised when interpreting partial LOH, because some high-grade astrocytomas have partial LOH secondary to their widespread genomic instability (Leung et al, 2000; Smith et al, 1999). The only genetic change that has been established as the “molecular signature” of oligodendroglioma is the combined LOH involving the entire chromosomal arms 1p and 19q (Bello et al, 2000; Bigner et al, 1999; Smith et al, 1999).
Ideally, the area to be dissected should contain solid tumor with as few contaminating normal cells as possible. However, in our experience, LOH can be reliably detected without comparison with normal tissue even when a few scattered, entrapped normal cells are seen in every high-powered field (×400), corresponding to a tumor cell content of approximately 80% to 90% (Fig. 8D). In all 19 tumor specimens, areas containing >90% tumor cells and measuring >3 mm2 were successfully identified and analyzed. In our subsequent analysis of more than 60 additional in-house and consult cases, we have not so far encountered a case in which normal cell contamination has been so extensive as to preclude analysis. A careful examination of hematoxylin and eosin (HE) slides from all available paraffin blocks has been sometimes necessary to find an area containing a sufficiently high density of tumor cells. In rare cases, the minimum of 3 mm2 of tumor required for the assay has been obtained only by pooling of two to four tumor foci. Specimens containing only infiltrative tumor with extensive normal brain contamination are not suitable for analysis using the present assay, which we recognize as a limitation of the assay. It may be possible to analyze such specimens using fluorescence in situ hybridization, provided that the infiltrating tumor cells can be reliably identified.
Relationship between the ratio of tumor cells to normal cells and the allelic ratio in an oligodendroglioma with LOH on 1p. Representative histology (×200) is shown in five different areas (A to E) dissected from a single slide of the oligodendroglioma specimen. The dissected areas measured 3 to 9 mm2. The figure demonstrates that even when scattered, entrapped normal cells are present among the tumor cells (D), the resulting allelic pattern is clearly different from the heterozygous pattern seen in normal brain (A). Areas such as D and E can be confidently analyzed even when no normal tissue is available for comparison. Note the diminutive peak (black arrow) in the electropherogram from E, originating from the capillary endothelial cells of the otherwise solid tumor. An area containing approximately equal numbers of tumor and normal cells (C) can be analyzed if an area of normal brain (A) can also be dissected and analyzed for comparison. If the specimen contains only sparse infiltrating tumor cells (B), the specimen is not suitable for LOH analysis using the reported assay.
Another potential problem with DNA-based analysis of microsatellite loci could be the possible difference in heterozygosity scores at the loci tested in different ethnic populations. In the worst case, allele frequency discrepancies could lead to false-positive results, because of higher than expected rates of allele homozygosity. Although we believe this possibility to be of fairly low likelihood, we nonetheless recommend that in cases showing a single allele pattern for all loci on 1p (as opposed to LOH pattern at some loci), LOH be confirmed by a subsequent analysis of normal tissue or blood from the same patient. Among the 19 cases in the present study, this recommendation would have applied to only 1 case (T197).
This method addresses the three main drawbacks that have been attributed to previous microsatellite-based LOH assays: the logistical challenge of obtaining a blood sample from each patient, the concern of false-negative results caused by normal cell contamination, and the difficulty of extracting DNA from a small amount of formalin-fixed tissue. Reportedly, these are the reasons why some laboratories have chosen fluorescence in situ hybridization instead of a microsatellite LOH assay (Titus, 2002). The most important conceptual advantage of the reported assay compared with previous microsatellite LOH assays is that most specimens can be analyzed without normal tissue from the same patient. Even when normal tissue is required, the high sensitivity of the assay allows for the use of a small, histologically normal area of brain (if present) instead of a blood sample. The high sensitivity of the assay also makes it easier to avoid areas of normal cell contamination, because the dissected area can be small (3 mm2) and the tissue can be dissected from a slide rather than a block of tissue. Finally, the method of DNA extraction incorporated in this assay has proven to be simple and robust. In summary, this is a relatively straightforward 1p and 19q LOH assay that can be performed as a part of the routine practice of diagnostic neuropathology.
Materials and Methods
Tissue Samples
Nineteen formalin-fixed, paraffin-embedded glioma specimens were used in this study. These specimens were selected because they were among the 21 Johns Hopkins Hospital glioma specimens that, as a part of a multicenter study (Smith et al, 1999), had been previously tested for 1p and 19q LOH by at least two of the following three established (reference) methods: comparative genomic hybridization, fluorescent chromosomal in situ hybridization, and PCR-based microsatellite analysis with comparison to normal DNA from the same patient. The cases for this previous study were selected based solely on availability of frozen tissue in our tumor bank. The identification numbers of the tumors in the multicenter study (Smith et al, 1999) were the same as the identification numbers used in this article. Two of the 21 cases were excluded before the start of the assay development and validation process, because inconsistent results had been reported by the three reference methods. One additional case was excluded because it could not be located in our archives. It was replaced by another Johns Hopkins case (T272) from a different study (Burger et al, 2001); this tumor also had been analyzed by the same reference methods. Ten glioma specimens were used for assay development and nonblinded validation, whereas nine specimens were used for blinded validation. This study was granted an exemption from review by the Johns Hopkins Joint Committee on Clinical Investigation.
Dissection of Tissue and Extraction of DNA
Three 5-μm–thick unstained sections were cut from the paraffin block. To minimize effects of DNA degradation resulting from exposure to air, only the deepest (third) section was used for the assay. The second section was stained with HE. The first section was not used. The area(s) with the highest density of tumor cells was identified by microscopic examination and marked on the HE slide (Fig. 3). The areas of tumor were required to contain >90% tumor cells and to measure 3 to 20 mm2. In all 19 cases, areas of tumor meeting these requirements were successfully identified. The cut-off value of 90% (minimum ratio of tumor cells to normal cells) was chosen conservatively. It was based on the allelic ratios that we have observed in specimens with different tumor/normal ratios, as well as on the range of allelic ratios that has been observed in heterozygous samples by us and other investigators (Skotheim et al, 2001). Areas of normal brain, if present, were also marked on the slide. The third section was placed in a slide mailer and deparaffinized by warming the slide at 60° C for 5 minutes, followed by incubations at room temperature on a shaking platform in xylene for 2 × 20 minutes, 100% ethanol for 2 minutes, and 95% ethanol for 2 minutes. The slide was then air dried for 5 to 10 minutes while protected from dust. The dissection and DNA extraction were performed as follows, according to the method of Turbett et al (1996), with small modifications. The HE slide (or its photocopy) served as a guide for dissection. Using a micropipette tip, the area of interest on the unstained, deparaffinized section was covered with Pinpoint solution (Zymo Research, Orange, California), a starch-based glue containing xylene cyanol to facilitate its visualization. The slide was air dried for 30 to 45 minutes. The area of interest was then peeled from the slide using a clean razor blade and placed in a PCR tube. DNA was released from the tissue fragment by an incubation in 55 μl of a proteinase K solution, prepared from kit components according to the manufacturer's instructions (Zymo Research), for 8 hours at 55° C in a thermocycler, followed by 10 minutes at 97° C to inactivate the proteinase K. The DNA extract was stored at 4° C up to several weeks until used in aliquots of 8 μl for PCR reactions. No purification of the DNA was performed, because this was found to result in a failure of the subsequent PCR.
PCR Primers
The PCR primer sequences for microsatellites on 1p and 19q were retrieved from the Genome Database (http://www.gdb.org/gdb/regionSearch.html). Initially, nine microsatellite loci on 1p and seven on 19q were selected based on their high heterozygosity scores (>0.83). Using two tumor samples, the corresponding 16 primer pairs were screened for a high yield and lack of nonspecific products in multiplex PCR. Five loci on 1p and three on 19q, all dinucleotide repeats, were then selected for the final assay (Fig. 2). Of note, all five loci on 1p and one of the three loci on 19q were also included in an LOH assay previously developed by the laboratory of Dr. Robert B. Jenkins (Burger et al, 2001). To facilitate consistent addition of a 3′ adenosine nucleotide to the forward strand during PCR, the sequences of some of the reverse primers were modified by adding a guanosine nucleotide to their 5′ ends (Magnuson et al, 1996) (Table 1). After synthesis, each forward primer was 5′-labeled with a fluorescent phosphoramidite (TET, HEX, or 6-FAM), as indicated in Table 1 (Oligos Etc, Bethel, Maine). Purification of the primers by HPLC was found to be necessary. Primer stock solutions (100 μm) were prepared by adding Tris EDTA buffer, pH 7.4, and stored at −20° C. Working solutions of forward (5 μm) and reverse primers (20 μm) were prepared from stock solutions by adding molecular biology grade distilled water and stored at 4° C.
Multiplex PCR
The primers were multiplexed into three PCR reactions: mix A (D1S162, D1S226, D1S199), mix B (D19S112, D1S186, D1S312), and mix C (D19S918, D19S206). In this study, all three reactions (A to C) were performed for all samples for validation purposes. In routine use, PCR with mix A can serve as an initial screen and PCRs with mix B and C need only be performed if LOH on 1p cannot be ruled out by the PCR with mix A (Fig. 3). To enhance the specificity and yield of the multiplex reactions, the concentrations of the forward primers were optimized at lower levels than those of the reverse primers (Berg et al, 2000) (Table 1). The PCR reactions contained GeneAmp 10× PCR6 buffer II (1× final), MgCl2 (1.5 mm final), dNTPs (0.3 mm final of each), AmpliTaq Gold DNA polymerase (0.625 U/25 μl final), primers (Table 1), DNA extract (8 μl), and distilled water up to 25 μl (all reagents from Applied Biosystems, Foster City, California). PCR cycling conditions were 95° C for 9 minutes once, 42 cycles (mix A and B) or 33 cycles (mix C) at 94° C for 45 seconds, 55° C for 45 seconds, and 72° C for 60 seconds, followed by 60° C for 45 minutes once and 4° C indefinitely.
Capillary Electrophoresis
Fluorescently labeled PCR products were detected by capillary electrophoresis using the ABI 310 Genetic Analyzer and GeneScan Collection software (Applied Biosystems). The loading solution contained 2.8 μl of the PCR product, 17.7 μl of deionized formamide, and 0.5 μl of TAMRA size standards. The following run parameters were used: GS STR POP-4 C 1 ml run module, 5 to 15 seconds of injection time, 15 kV injection and electrophoresis voltage, column temperature of 60° C, and a 20-minute electrophoresis time.
Interpretation of Electropherograms During Assay Validation
Examination of the allelic patterns in the 10 tumors used for assay development revealed that the ranges of intensity (peak heights) of the less intense allele (when present) relative to the more prominent allele did not overlap between cases known to have LOH (eg, Case T197 in Fig. 4, Fig. 6) and cases known not to have LOH (eg, Case T250 in Fig. 4). These observed ranges of relative peak heights were used to devise written guidelines on interpreting the results (for more details, see “Results”). These interpretation guidelines were then applied in a blinded validation study of the assay using an additional nine tumor specimens (Table 2). The technical steps of the assay and interpretation of the results in the blinded validation phase were performed by persons who had no knowledge of the results of the reference methods or the histologic diagnoses and who had not been directly involved in the initial assay development.
References
Bello MJ, de Campos JM, Vaquero J, Ruiz-Barnes P, Kusak ME, Sarasa JL, and Rey JA (2000). hRAD54 gene and 1p high-resolution deletion-mapping analyses in oligodendrogliomas. Cancer Genet Cytogenet 116: 142–147.
Berg KD, Glaser CL, Thompson RE, Hamilton SR, Griffin CA, and Eshleman JR (2000). Detection of microsatellite instability by fluorescence multiplex polymerase chain reaction. J Mol Diagn 2: 20–28.
Bigner SH, Matthews MR, Rasheed BK, Wiltshire RN, Friedman HS, Friedman AH, Stenzel TT, Dawes DM, McLendon RE, and Bigner DD (1999). Molecular genetic aspects of oligodendrogliomas including analysis by comparative genomic hybridization. Am J Pathol 155: 375–386.
Burger PC, Minn AY, Smith JS, Borell TJ, Jedlicka AE, Huntley BK, Goldthwaite PT, Jenkins RB, and Feuerstein BG (2001). Losses of chromosomal arms 1p and 19q in the diagnosis of oligodendroglioma: A study of paraffin-embedded sections. Mod Pathol 14: 842–853.
Cairncross JG, Ueki K, Zlatescu MC, Lisle DK, Finkelstein DM, Hammond RR, Silver JS, Stark PC, Macdonald DR, Ino Y, Ramsay DA, and Louis DN (1998). Specific genetic predictors of chemotherapeutic response and survival in patients with anaplastic oligodendrogliomas. J Natl Cancer Inst 90: 1473–1479.
Coons SW, Johnson PC, Scheithauer BW, Yates AJ, and Pearl DK (1997). Improving diagnostic accuracy and interobserver concordance in the classification and grading of primary gliomas. Cancer 79: 1381–1393.
Ino Y, Betensky RA, Zlatescu MC, Sasaki H, Macdonald DR, Stemmer-Rachamimov AO, Ramsay DA, Cairncross JG, and Louis DN (2001). Molecular subtypes of anaplastic oligodendroglioma: Implications for patient management at diagnosis. Clin Cancer Res 7: 839–845.
Leighton C, Fisher B, Bauman G, Depiero S, Stitt L, MacDonald D, and Cairncross G (1997). Supratentorial low-grade glioma in adults: An analysis of prognostic factors and timing of radiation. J Clin Oncol 15: 1294–1301.
Leung SY, Yuen ST, Chan TL, Chan AS, Ho JW, Kwan K, Fan YW, Hung KN, Chung LP, and Wyllie AH (2000). Chromosomal instability and p53 inactivation are required for genesis of glioblastoma but not for colorectal cancer in patients with germline mismatch repair gene mutation. Oncogene 19: 4079–4083.
Magnuson VL, Ally DS, Nylund SJ, Karanjawala ZE, Rayman JB, Knapp JI, Lowe AL, Ghosh S, and Collins FS (1996). Substrate nucleotide-determined non-templated addition of adenine by Taq DNA polymerase: Implications for PCR-based genotyping and cloning. Biotechniques 21: 700–709.
Sasaki H, Zlatescu MC, Betensky RA, Johnk LB, Cutone AN, Cairncross JG, and Louis DN (2002). Histopathological-molecular genetic correlations in referral pathologist-diagnosed low-grade “oligodendroglioma.” J Neuropathol Exp Neurol 61: 58–63.
Skotheim RI, Diep CB, Kraggerud SM, Jakobsen KS, and Lothe RA (2001). Evaluation of loss of heterozygosity/allelic imbalance scoring in tumor DNA. Cancer Genet Cytogenet 127: 64–70.
Smith JS, Alderete B, Minn Y, Borell TJ, Perry A, Mohapatra G, Hosek SM, Kimmel D, O'Fallon J, Yates A, Feuerstein BG, Burger PC, Scheithauer BW, and Jenkins RB (1999). Localization of common deletion regions on 1p and 19q in human gliomas and their association with histological subtype. Oncogene 18: 4144–4152.
Smith JS, Perry A, Borell TJ, Lee HK, O'Fallon J, Hosek SM, Kimmel D, Yates A, Burger PC, Scheithauer BW, and Jenkins RB (2000). Alterations of chromosome arms 1p and 19q as predictors of survival in oligodendrogliomas, astrocytomas, and mixed oligoastrocytomas. J Clin Oncol 18: 636–645.
Titus K (2002). Brainteaser: 1p/19q deletions in gliomas. CAP Today (Sep):1.
Turbett GR, Barnett TC, Dillon EK, and Sellner LN (1996). Single-tube protocol for the extraction of DNA or RNA from paraffin-embedded tissues using a starch-based adhesive. Biotechniques 20: 846–850, 852–853.
Ueki K, Nishikawa R, Nakazato Y, Hirose T, Hirato J, Funada N, Fujimaki T, Hojo S, Kubo O, Ide T, Usui M, Ochiai C, Ito S, Takahashi H, Mukasa A, Asai A, and Kirino T (2002). Correlation of histology and molecular genetic analysis of 1p, 19q, 10q, TP53, EGFR, CDK4, and CDKN2A in 91 astrocytic and oligodendroglial tumors. Clin Cancer Res 8: 196–201.
Acknowledgements
We thank Ms. Cynthia L. Glaser and Mr. Michael Hafez for technical assistance in the blinded validation of the assay, Ms. Patricia Goldthwaite for managing the brain tumor archive, and Dr. Charles G. Eberhart for valuable suggestions.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work was supported by National Cancer Institute Grant CA62475 (to PCB) and an American Brain Tumor Association Grant (to KJH).
Rights and permissions
About this article
Cite this article
Hatanpaa, K., Burger, P., Eshleman, J. et al. Molecular Diagnosis of Oligodendroglioma in Paraffin Sections. Lab Invest 83, 419–428 (2003). https://doi.org/10.1097/01.LAB.0000059948.67795.EF
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1097/01.LAB.0000059948.67795.EF
This article is cited by
-
Relative oxygen extraction fraction (rOEF) MR imaging reveals higher hypoxia in human epidermal growth factor receptor (EGFR) amplified compared with non-amplified gliomas
Neuroradiology (2021)
-
The genomic landscape of TERT promoter wildtype-IDH wildtype glioblastoma
Nature Communications (2018)
-
IDH-mutant glioma specific association of rs55705857 located at 8q24.21 involves MYC deregulation
Scientific Reports (2016)
-
High expression of the stem cell marker nestin is an adverse prognostic factor in WHO grade II–III astrocytomas and oligoastrocytomas
Journal of Neuro-Oncology (2014)
-
Extraventricular anaplastic ependymoma with metastasis to scalp and neck
Journal of Neuro-Oncology (2011)