Abstract
Analysis of the microRNA (miRNA) expression signature of lung squamous cell carcinoma (lung-SCC) revealed that the expression levels of miR-133a were significantly reduced in cancer tissues compared with normal tissues. In this study, we focused on the functional significance of miR-133a in cancer cell lines derived from lung-SCC and the identification of miR-133a-regulated novel cancer networks in lung-SCC. Restoration of miR-133a expression in PC10 and H157 cell lines resulted in significant inhibition of cell proliferation, suggesting that miR-133a functions as a tumor suppressor. We used genome-wide gene expression analysis to identify the molecular targets of miR-133a regulation. Gene expression data and web-based searching revealed several candidate genes, including transgelin 2 (TAGLN2), actin-related protein2/3 complex, subunit 5, 16kDa (ARPC5), LAG1 homolog, ceramide synthase 2 (LASS2) and glutathione S-transferase pi 1 (GSTP1). ARPC5 and GSTP1 likely represent bona fide targets as their expression is elevated in lung-SCC clinical specimens. Furthermore, transient transfection of miR-133a, repressed ARPC5 and GSTP1 mRNA and protein levels. As cell proliferation was significantly inhibited in lung-SCC cells following RNAi knock down of either gene, ARPC5 and GSTP1 may function as oncogenes in the development of lung-SCC. The identification of a tumor suppressive miRNA and the novel cancer pathways it regulates could provide new insights into potential molecular mechanisms of lung-SCC carcinogenesis.
Similar content being viewed by others
Introduction
Lung cancer is clearly the primary cause of cancer-related deaths worldwide. In Japan, it accounts for about 60 000 deaths every year, and is the leading cause of cancer-related deaths in men and the third leading cause in women. Approximately 80% of lung cancers are classified histopathologically as non-small cell lung cancers (NSCLC). NSCLC is subdivided into four major histological subtypes with distinct pathological characteristics: adenocarcinoma, squamous cell carcinoma, large cell carcinoma and neuroendocrine cancer.1, 2 Curative outcomes are found only in patients with early stage disease who undergo surgery, with a 5-year overall survival rate of 50–60%,1, 2 whereas patients in advanced stages rarely survive more than 5 years despite aggressive chemotherapy, molecular-targeted therapy or chemoradiotherapy.3, 4
It is well known that altered expression of cell surface growth factor receptors, including the receptor tyrosine kinase (RTK) family, has a major role in the development of many types of cancer, including NSCLC.5 Over-expression of epidermal growth factor receptor (also known as Erb1) and other members of the ErbB family of receptor tyrosine kinases (ErbB2 and ErbB3) is often observed in many cancers and contributes to increased cell proliferation.6 On the basis of this fact, several therapeutic agents have been designed in lung cancers, such as gefitinib and erlotinib for the mutation of epidermal growth factor receptor gene, and crizotinib for the EML4-ALK fusion gene.7 However, such a molecular-targeted strategy has not been developed for squamous cell carcinoma, but only for adenocarcinoma of the lung. The new genome-wide RNA analysis of lung squamous cell carcinoma (lung-SCC) may provide new avenues for research, and the development of novel diagnostic approaches and therapeutics for lung-SCC.
For the past several decades, many genes (such as RAS, MYC, PTEN and EGFR) have been identified and implicated in lung cancer that also have a role in normal cell growth and differentiation. Recent studies have revealed that the normal regulatory mechanisms in these cancer pathways are disrupted by aberrant expression of microRNA (miRNA).8 MiRNAs are a class of small non-coding RNA molecules consisting of 19–22 nucleotides that regulate gene expression through translational repression and mRNA cleavage,8 and have an important role in a variety of biological processes, including development, differentiation, apoptosis and cell proliferation. Bioinformatic predictions indicate that miRNAs regulate more than 30% of protein coding genes.9 Currently, 1424 human miRNAs have been registered at miRBase release 17.0 (http://microrna.sanger.ac.uk/).
Signaling by the RAS oncogene pathway has a central role in lung cancer and approximately 30% of lung cancers have activating mutations in this cascade.10 A growing body of studies has revealed that aberrant expression of miRNAs is associated with the initiation and development of NSCLC,11 and alterations of RAS signaling. For example, downregulation of the let-7 family of miRNAs, which normally reduces RAS expression levels by directly binding to the 3′UTR of RAS mRNA,12 was observed in cancer tissues from a NSCLC patient. Restoration of let-7 expression inhibited the growth of cancer cells, suggesting that let-7 reduction leads to over-expression of RAS and is a primary mediator of NSCLC carcinogenesis.
Recent studies from our group have identified tumor suppressive miRNAs and the molecular pathways they control in various human cancers.13, 14, 15, 16, 17, 18 As there are few studies about the role of miRNAs in lung-SCC, we sought to identify tumor suppressive miRNAs based on our lung-SCC expression signature. We focused on miR-133a, which was the most significantly downregulated miRNA in our signature, and we provide evidence that it functions as a tumor suppressor that inhibits cancer cell proliferation. To identify the target genes regulated by miR-133a, we also performed genome-wide gene expression analysis and found several candidate genes such as actin-related protein2/3 complex, subunit 5, 16 kDa (ARPC5) and glutathione S-transferase pi 1 (GSTP1). Insights into the association between tumor suppressive miR-133a and their target oncogene networks could enhance our understanding of the molecular mechanism of lung-SCC carcinogenesis.
Materials and methods
Clinical lung-SCC specimens and lung-SCC cell culture
A total of 25 pairs of primary lung-SCC tissue and corresponding normal tissue samples were obtained from patients in Chiba University Hospital (Chiba, Japan) from 2004 to 2007 (clinical features of patients with lung-SCC are shown in Table 1). Normal tissues were obtained far from the center of the cancer in surgical specimens. No cancer cells were detected in neighboring formalin-fixed paraffin-embedded tissues. Written consent was obtained from each patient authorizing tissue donation for research purposes before surgery, following a protocol approved by the Institutional Review Board of Chiba University. The specimens were immersed in RNAlater (Qiagen, Valencia, CA, USA) and stored at −20 °C until RNA was extracted. Human lung-SCC cell lines, PC10 and H157, were grown in RPMI1640 medium supplemented with 10% fetal bovine serum in a humidified atmosphere containing 5% CO2 at 37 °C.
RNA isolation
Total RNA was isolated using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. RNA concentrations were determined using a UV spectrophotometer and molecule integrity was checked by gel electrophoresis. RNA quality was confirmed using an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA).
miRNA expression signature of lung-SCC
Tissue specimens used for miRNA screening of a low-density array were obtained from five lung-SCC patients (Table 1; patient number 1–5). miRNA expression patterns were evaluated using the TaqMan LDA Human microRNA Panel v2.0 (Applied Biosystems, Foster City, CA, USA). The assay was composed of two steps: generation of cDNA by reverse transcription and a TaqMan real-time PCR assay. Description of real-time PCR and the list of human miRNAs can be found on the company's website (http://www.appliedbiosystems.com). Analysis of relative miRNA expression data was performed using GeneSpring GX version 7.3.1 software (Agilent Technologies) according to the manufacturer's instructions. A cutoff P value of <0.05 was used to narrow down the candidates after global normalization of the raw data. After global normalization, additional normalization was done with RNU48.
Mature miRNA transfection and small interfering RNA treatment
Mature miRNA molecules, Pre-miR miRNA precursors; (hsa-miR-133a; Pre-miR ID: AM10413 and negative control miRNA; P/N: AM17111, Applied Biosystems), small interfering RNA; si-ARPC5 (P/N: HSS145450 and HSS145450, Invitrogen), si-GSTP1 (P/N: s194475 and s194476, Applied Biosystems) and negative control siRNA (Stealth RNAi Negative Control Medium GC Duplex; 12935-300, Invitrogen) were incubated with Opti-MEM (Invitrogen) and Lipofectamine RNAiMax reagent (Invitrogen). Transfection efficiency of Pre-miR in cell lines was confirmed on the basis of downregulation of PTK9 mRNA following transfection with miR-1 as described previously.13, 14
Cell proliferation assays
Cells were transfected with 10 nM miRNA or siRNA by reverse transfection and plated into 96-well plates at 3 × 103 cells per well. After 72 and 96 h, cell proliferation was determined by the XTT assay, using the Cell Proliferation Kit II (Roche Molecular Biochemicals, Mannheim, Germany).14, 15 Triplicate wells were measured for cell viability in each treatment group.
Target gene search for miR-133a
To identify target genes of miR-133a, a genome-wide screen was performed in two lung-SCC cell lines, PC10 and H157, which were transfected with miR-133a. Expression profiles were generated using Oligo-microarray Human 44 K (Agilent Technologies) from the transfected cells and compared with a miRNA-negative-control sample. Hybridization and wash steps were performed as previously described.19 The arrays were scanned using a Packard GSI Lumonics ScanArray 4000 (Perkin Elmer, Boston, MA, USA). The data were analyzed by means of DNASIS array software (Hitachi Software Engineering, Tokyo, Japan), which converted the signal intensity for each spot into text format. The log2 ratios of the median-subtracted background intensity were analyzed. Data from each microarray study were normalized by a global normalization method.19
Predicted target genes and their miRNA binding site seed regions were investigated using TargetScan (release 5.1, http://www.targetscan.org/). The sequences of the predicted mature miRNAs were confirmed using miRBase (release 17.0, http://microrna.sanger.ac.uk/).
Real-time quantitative RT-PCR
First-strand cDNA was synthesized from 1 μg total RNA using High Capacity cDNA Reverse Transcription Kit (Applied Biosystems). Gene-specific PCR products were assayed continuously using a 7900-HT Real-Time PCR System according to the manufacturer's protocol. The initial PCR step consisted of a 10-min hold at 95 °C, followed by 40 cycles of a 15-sec denaturation at 95 °C and a 1-min annealing/extension at 63 °C. TaqMan probes and primers for ARPC5 (P/N: Hs00271722_m1), GSTP1 (P/N: Hs00168310_m1) and GUSB (P/N: Hs99999908_m1) as an internal control were obtained from Applied Biosystems (Assay-On-Demand Gene Expression Products). The expression levels of miR-133a (Assay ID: 002246) were analyzed by TaqMan quantitative real-time PCR (TaqMan MicroRNA Assay; Applied Biosystems) and normalized to RNU48 (Assay ID: 001006). All reactions were performed in triplicate and included negative control reactions that lacked cDNA.
Western blot
Cells were harvested 72 h after transfection and lysates were prepared. Using SDS-PAGE, 50 μg protein from each lysate was separated on Mini-PROTEAN TGX (Tris-Glycine eXtended) precast gels (Bio-Rad, Hercules, CA, USA) and transferred to PVDF membranes. Immunoblotting was performed with diluted (1:500) monoclonal ARPC5 antibody (ab51243; Abcam, Cambridge, UK), monoclonal GSTP1 antibody (#3369; Cell Signaling, Danvers, MA, USA) or GAPDH antibody (ab8245, Abcam) used as an internal control. The membrane was washed and incubated with goat anti-rabbit or anti-mouse IgG (H+L)–HRP conjugate (Bio-Rad). Specific complexes were visualized by echochemiluminescence (Immun-Star WesternC Chemiluminescence Kit #170-5070, Bio-Rad) and the expression level of each protein was evaluated by ImageJ software (ver.1.44; http://rsbweb.nih.gov/ij/index.html).
Statistical analysis
The relationships between the two groups and the numerical values obtained by real-time RT-PCR were analyzed using the nonparametric Mann–Whitney U test or the paired t-test. The relationship among three variables and numerical values was analyzed using the Bonferroni-adjusted Mann–Whitney U test; a non-adjusted statistical level of significance of P<0.05 corresponded to a Bonferroni-adjusted level of P<0.0167. Spearman's rank test was used to evaluate the relationships among the relative expression levels of miR-133a, ARPC5 and GSTP1 mRNA. All analyses were performed using Expert StatView (version 4, SAS Institute Inc., Cary, NC, USA).
Results
Identification of downregulated miRNAs in lung-SCC by miRNA expression signatures
We evaluated mature miRNA expression levels of five pairs of normal tissues and lung-SCC by miRNA expression signatures. A total of 24 significantly downregulated miRNAs were selected after the normalization using RNU48, and then 21 miRNAs were selected as their fold change were less than 0.5 (Table 2). Among them, miR-133a was the most downregulated and was selected for further study.
Expression of miR-133a in lung-SCC clinical specimens and effect of miR-133a transfectants on cell proliferation in PC10 and H157
Quantitative stem-loop RT-PCR demonstrated that the expression level of miR-133a was significantly reduced in 20 lung-SCC specimens (Table 1; patient number 6–25) in comparison with normal tissues (P<0.0001, Figure 1a).
miR-133a functions as a tumor suppressor in lung-SCC. (a) The expression level of miR-133a was significantly downregulated in tumor tissues compared with non-tumor tissues. RNU48 was used as an internal control. (b) Cancer cell proliferation was determined by XTT assay in PC10 and H157 cell lines. The XTT assay was performed 72 and 96h after transfection of 10nM miR-133a or miR-control and showed significant inhibition of cell proliferation in miR-133a transfectants in comparison with controls. *P<0.05.
To investigate the functional role of miR-133a, we performed gain-of-function studies in miRNA transfectants. The XTT assay showed significant inhibition of cell proliferation in miR-133a transfectants in comparison with the miR-control transfectants (% of cell viability, 72 h; PC10; 67.1±2.3 and 100.0±2.7, H157; 54.2±3.6 and 100.0±3.9, respectively, 96 h; PC10; 54.9±2.3 and 100.0±2.7, H157; 62.2±3.6 and 100.0±3.1, respectively, P<0.05, Figure 1b).
Expression profiling identifies downregulated genes in miR-133a transfectants
To gain further insight into which genes are regulated by miR-133a, we performed gene expression analysis following miR-133a transfection compared with negative controls in PC10 and H157 cells. Signal values less than 1000 from the raw data in miR-control transfectants were not further considered. A total of 116 genes were downregulated with log2 ratio scores less than −1.0 in miR-133a transfectants compared with the controls (top 30 genes are shown in Table 3). The TargetScan program showed that 14 genes had putative target sites of miR-133a in their 3′UTR among the top 30 downregulated genes. Entries from the microarray data were approved by the Gene Expression Omnibus and were assigned Gene Expression Omnibus accession number, GSE26032.
Expression levels of candidate target genes of miR-133a in lung-SCC clinical specimens
We measured the expression levels of mRNAs of the top four candidate genes in 20 lung-SCC clinical specimens by quantitative real-time RT–PCR. Two genes, actin-related protein2/3 complex, subunit 5, 16kDa (ARPC5) and glutathione S-transferase pi 1 (GSTP1) were significantly upregulated in the tumor region (P=0.019 and P=0.0022, respectively, Figure 2a and b, upper). The other two genes (TAGLN2 and LASS2) were not upregulated in the tumor region of lung-SCC (data not shown). There was a trend but no significant inverse correlation between ARPC5—miR-133a (r=−0.23 and P=0.16, Figure 2a, lower) and a significant inverse correlation between GSTP1—miR-133a (r=−0.65 and P<0.0001, Figure 2b, lower).
Increased expression levels of ARPC5 and GSTP1 mRNA in lung-SCC clinical specimens. (a) The mRNA expression levels of ARPC5 in tumor tissues and non-tumor tissues in 20 clinical specimens (upper). Expression of ARPC5 mRNA was significantly elevated in tumor tissues. The correlation between ARPC5 and miR-133a expression was investigated in clinical specimens (lower). The trend of inverse correlation was recognized between ARPC5 and miR-133a expression. (b) The mRNA expression levels of GSTP1 in tumor tissues and non-tumor tissues in 20 clinical specimens (upper). Expression of GSTP1 mRNA was significantly elevated in tumor tissues. The correlation between GSTP1 and miR-133a expression was investigated in clinical specimens (lower). A significant inverse correlation was recognized between GSTP1 and miR-133a expression.
Silencing of ARPC5 and GSTP1 by miR-133a transfection
We performed gain-of-function studies using miR-133a transfectants, and the ARPC5 and GSTP1 mRNA and protein expression levels were markedly downregulated in comparison with the controls (Figures 3 and 4).
miR-133a regulates ARPC5 expression in lung-SCC cell lines. (a) Both mRNA and protein expression levels of ARPC5 were significantly repressed in miR-133a-transfected PC10 cells. An internal control was GUSB for RT-PCR, and GAPDH for western blot analysis. (b) Both mRNA and protein expression levels of ARPC5 were significantly repressed in miR-133a-transfected H157 cells. *P<0.05. Predicted target sites of miR-133a in ARPC5 in the 3′UTR region are shown (right).
miR-133a regulates GSTP1 expression in lung-SCC cell lines. (a) Both mRNA and protein expression levels of GSTP1 were significantly repressed in miR-133a-transfected PC10 cells. An internal control was GUSB for RT-PCR, and GAPDH for western blot analysis. (b) Both mRNA and protein expression levels of GSTP1 were significantly repressed in miR-133a-transfected H157 cells. *P<0.05. Predicted target sites of miR-133a in GSTP1 in the 3′UTR region are shown (right).
Effect of silencing of ARPC5 and GSTP1 on cell proliferation in PC10
To examine the functional role of ARPC5 and GSTP1, we performed loss-of-function studies in PC10 cells using two different si-RNAs for each gene. The ARPC and GSTP1 levels of mRNA and protein expression were markedly repressed by each si-RNA treatment (Figures 5a, b, 6a and b). The XTT assay revealed significant inhibition of cell proliferation in each of the si-RNA transfectants in comparison with controls (% of cell viability, si-ARPC5; 55.6±4.9, 35.6±2.1 and 100.0±9.3, respectively, P<0.0001; Figure 5c, si-GSTP1; 60.9±4.6, 69.3±7.6 and 100.0±9.0, respectively, P<0.0001; Figure 6c).
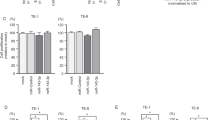
Effects of ARPC5 silencing in lung-SCC cell line PC10. (a) Expression of ARPC5 mRNA was significantly repressed by two si-ARPC5 transfectants. GUSB was used as an internal control. (b) Expression of ARPC5 protein was repressed in two si-ARPC5 transfectants. GAPDH was used as an internal control. (c) Cancer cell proliferation was determined with the XTT assay 72h post transfection. The XTT assay showed significant inhibition of cell proliferation in two si-ARPC5 transfectants. *P<0.0001.
Effects of GSTP1 silencing in lung-SCC cell line PC10. (a) Expression of GSTP1 mRNA was significantly repressed in two si-GSTP1 transfectants. GUSB was used as an internal control. (b) Expression of GSTP1 protein was repressed in two si-GSTP1 transfectants. GAPDH was used as an internal control. (c) Cancer cell proliferation was determined with the XTT assay after 72h post transfection. The XTT assay showed significant inhibition of cell proliferation in two si-GSTP1 transfectants. *P<0.0001.
Discussion
MiRNAs are a new class of small RNAs that regulate the expression of many genes, and a large number of studies have shown that miRNAs are aberrantly expressed in many types of human cancers.20 MiRNA expression profiles of NSCLC have also been reported by many researchers.21, 22 However, few studies describe similar analysis of lung-SCC, one of the most lethal malignancies in the world, with a 5-year survival rate of approximately 60% after curative surgery. Increased understanding of the molecular pathways involved in lung-SCC carcinogenesis would help improve diagnosis and therapy of the disease.
We screened 665 miRNAs, from lung-SCC and paired normal epithelial specimens, using stem-loop RT–PCR and identified 21 miRNAs downregulated in cancer tissues. To measure the effectiveness of our miRNA expression signature, we compared our data with the lists of downregulated miRNAs from past reports. Several genes (miR-30a-3p, miR-126, miR-574-3p and miR-320) were concordantly identified in our study and that of a recent report that analyzed miRNA profiles of lung-SCC.23 Our data suggest that downregulated miRNAs can function as tumor suppressor genes and contribute to lung-SCC oncogenesis. Future experiments analyzing the function of miRNA regulation of gene expression in lung-SCC are likely to improve the understanding of mechanisms underlying this deadly disease.
In our expression signature, miR-133a is the most downregulated miRNA and expression studies revealed a reduction in miR-133a levels in many types of cancers.14, 18, 24, 25, 26, 27 Very interestingly, our recent reports of head and neck squamous cell carcinoma (SCC) and esophageal SCC showed that miR-133a is also significantly reduced in cancer tissues compared with normal epithelium.14, 17 The functional significance of miR-133a was investigated using head and neck SCC, esophageal SCC and bladder cell lines, and our data showed that restoration of miR-133a expression inhibited cancer cell proliferation, invasion and migration, and directly regulated actin-related genes such as FSCN1, LASP1 and TAGLN2.13, 14, 18, 28 The identification of miR-133a regulation of cancer pathways could provide new insights into potential mechanisms of human carcinogenesis.
In this study, we focused on the functional significance of miR-133a, because the role of this miRNA in lung-SCC is not well understood. First, we confirmed the reduction of miR-133a expression in clinical specimens. Furthermore, we show that restoration of miR-133a in cancer cells significantly inhibited cell proliferation, suggesting that miR-133a functions as a tumor suppressor in lung-SCC as well as other cancers. To investigate the messenger RNA–miRNA networks in lung-SCC, we adopted a method of genome-wide gene expression analysis in PC10 and H157 cells, using miR-133a transfectants to identify oncogenic targets. We selected four candidate target genes, transgelin 2 (TAGLN2), actin-related protein2/3 complex, subunit 5, 16kDa (ARPC5), LAG1 homolog, ceramide synthase 2 (LASS2) and glutathione S-transferase pi 1 (GSTP1), from our expression profile. Two genes (ARPC5 and GSTP1) are upregulated in cancer tissues and were analyzed in detail.
This is the first report that ARPC5 is regulated by miR-133a in lung-SCC cells. Actin is rapidly formed in response to specific cellular signals that converge on the Arp2/3 complex to regulate assembly.29 The human Arp2/3 complex is composed of seven subunits including ARPC5, the smallest subunit.30 We found that increased expression of ARPC5 gene in lung-SCC correlated with reduced levels of miR-133a. A TargetScan database search revealed that ARPC5 is a candidate target of miR-133a. ARPC5 mRNA and protein expression is reduced in miR-133a transfectant cell lines, indicating that miR-133a regulates ARPC5 levels in lung-SCC cells. Silencing of ARPC5 reduced cancer cell proliferation in PC10 cells, suggesting that ARPC5 might contribute to lung-SCC cancer development. Thus, ARPC5 functions as an oncogene and may represent a new diagnostic marker and therapeutic target for the treatment of lung-SCC. In addition, we previously performed genome-wide gene expression analysis using miR-133a transfectants derived from head and neck SCC and bladder cancer cell lines. According to the expression signatures, ARPC5 is a candidate of miR-133a targets in cancer cells. This result suggests that miR-133a–ARPC5 pathway is universally important to cancer cell. These microarray data are registered in Gene Expression Omnibus database, accession numbers GSE 19717, 20028 and 26032.
GSTP1, a member of the GST enzyme superfamily, catalyzes the conjugation of electrophiles to glutathione in the process of detoxification.31 GSTP1 protein has several critical roles in both normal and neoplastic cells, including phase II xenobiotic metabolism, stress responses, signaling and apoptosis. Over-expression of GSTP1 has been observed in many types of cancer, including lung cancer.32, 33, 34 More recently, we demonstrated that miR-133a functions as a tumor suppressor and directly regulates GSTP1 in head and neck SCC16 and bladder cancer.35 This analysis suggests that a similar regulation occurs in lung-SCC and may indicate that regulation of GSTP1 by the tumor suppressor miR-133a is a common feature of human SCC.
In conclusion, the reduction of miR-133a was a frequent event in lung-SCC cancer cells. miR-133a may function as a tumor suppressor and it regulates ARPC5 and GSTP1. Both of these genes were upregulated in lung-SCC specimens. The miR-133a-regulated novel cancer pathways could provide new insights into molecular mechanisms in lung-SCC and might contribute to the development of new therapeutic strategy for the disease.
References
Chansky, K., Sculier, J. P., Crowley, J. J., Giroux, D., Van Meerbeeck, J., Goldstraw, P. et al. The International Association for the Study of Lung Cancer Staging Project: prognostic factors and pathologic TNM stage in surgically managed non-small cell lung cancer. J. Thorac. Oncol. 4, 792–801 (2009).
Sawabata, N., Asamura, H., Goya, T., Mori, M., Nakanishi, Y., Eguchi, K. et al. Japanese Lung Cancer Registry Study: first prospective enrollment of a large number of surgical and nonsurgical cases in 2002. J. Thorac. Oncol. 5, 1369–1375 (2010).
Schiller, J. H., Harrington, D., Belani, C. P., Langer, C., Sandler, A., Krook, J. et al. Comparison of four chemotherapy regimens for advanced non-small-cell lung cancer. N. Engl. J. Med. 346, 92–98 (2002).
Sandler, A., Gray, R., Perry, M. C., Brahmer, J., Schiller, J. H., Dowlati, A. et al. Paclitaxel-carboplatin alone or with bevacizumab for non-small-cell lung cancer. N. Engl. J. Med. 355, 2542–2550 (2006).
Lemmon, M. A. & Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 141, 1117–1134 (2010).
Bublil, E. M. & Yarden, Y. The EGF receptor family: spearheading a merger of signaling and therapeutics. Curr. Opin. Cell Biol. 19, 124–134 (2007).
Bronte, G., Rizzo, S., La Paglia, L., Adamo, V., Siragusa, S., Ficorella, C. et al. Driver mutations and differential sensitivity to targeted therapies: a new approach to the treatment of lung adenocarcinoma. Cancer Treat. Rev. 36 (Suppl 3), S21–9 (2010).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Filipowicz, W., Bhattacharyya, S. N. & Sonenberg, N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat. Rev. Genet. 9, 102–114 (2008).
Rodenhuis, S. & Slebos, R. J. The ras oncogenes in human lung cancer. Am. Rev. Respir. Dis. 142, S27–30 (1990).
Calin, G. A. & Croce, C. M. MicroRNA signatures in human cancers. Nat. Rev. Cancer. 6, 857–866 (2006).
Johnson, S. M., Grosshans, H., Shingara, J., Byrom, M., Jarvis, R., Cheng, A. et al. RAS is regulated by the let-7 microRNA family. Cell 120, 635–647 (2005).
Chiyomaru, T., Enokida, H., Tatarano, S., Kawahara, K., Uchida, Y., Nishiyama, K. et al. miR-145 and miR-133a function as tumour suppressors and directly regulate FSCN1 expression in bladder cancer. Br. J. Cancer 102, 883–891 (2010).
Kano, M., Seki, N., Kikkawa, N., Fujimura, L., Hoshino, I., Akutsu, Y. et al. miR-145, miR-133a and miR-133b: tumor suppressive miRNAs target FSCN1 in esophageal squamous cell carcinoma. Int. J. Cancer 127, 2804–2814 (2010).
Kikkawa, N., Hanazawa, T., Fujimura, L., Nohata, N., Suzuki, H., Chazono, H. et al. miR-489 is a tumour-suppressive miRNA target PTPN11 in hypopharyngeal squamous cell carcinoma (HSCC). Br. J. Cancer 103, 877–884 (2010).
Mutallip, M., Nohata, N., Hanazawa, T., Kikkawa, N., Horiguchi, S., Fujimura, L. et al. Glutathione S-transferase P1 (GSTP1) suppresses cell apoptosis and its regulation by miR-133alpha in head and neck squamous cell carcinoma (HNSCC). Int. J. Mol. Med. 27, 345–352 (2011).
Nohata, N., Sone, Y., Hanazawa, T., Fuse, M., Kikkawa, N., Yoshino, H. et al. miR-1 as a tumor suppressive microRNA targeting TAGLN2 in head and neck squamous cell carcinoma. Oncotarget 2, 29–44 (2011).
Yoshino, H., Chiyomaru, T., Enokida, H., Kawakami, K., Tatarano, S., Nishiyama, K. et al. The tumour-suppressive function of miR-1 and miR-133a targeting TAGLN2 in bladder cancer. Br. J. Cancer 104, 808–818 (2011).
Sugimoto, T., Seki, N., Shimizu, S., Kikkawa, N., Tsukada, J., Shimada, H. et al. The galanin signaling cascade is a candidate pathway regulating oncogenesis in human squamous cell carcinoma. Genes Chromosomes Cancer 48, 132–142 (2009).
Almeida, M. I., Reis, R. M. & Calin, G. A. MicroRNA history: discovery, recent applications, and next frontiers. Mutat. Res. 717, 1–8 (2011).
Lin, P. Y., Yu, S. L. & Yang, P. C. MicroRNA in lung cancer. Br. J. Cancer 103, 1144–1148 (2010).
Du, L. & Pertsemlidis, A. microRNAs and lung cancer: tumors and 22-mers. Cancer Metastasis Rev. 29, 109–122 (2010).
Yang, Y., Li, X., Yang, Q., Wang, X., Zhou, Y., Jiang, T. et al. The role of microRNA in human lung squamous cell carcinoma. Cancer Genet. Cytogenet. 200, 127–133 (2010).
Arndt, G. M., Dossey, L., Cullen, L. M., Lai, A., Druker, R., Eisbacher, M. et al. Characterization of global microRNA expression reveals oncogenic potential of miR-145 in metastatic colorectal cancer. BMC Cancer 9, 374 (2009).
Childs, G., Fazzari, M., Kung, G., Kawachi, N., Brandwein-Gensler, M., McLemore, M. et al. Low-level expression of microRNAs let-7d and miR-205 are prognostic markers of head and neck squamous cell carcinoma. Am. J. Pathol. 174, 736–745 (2009).
Ichimi, T., Enokida, H., Okuno, Y., Kunimoto, R., Chiyomaru, T., Kawamoto, K. et al. Identification of novel microRNA targets based on microRNA signatures in bladder cancer. Int. J. Cancer 125, 345–352 (2009).
Wong, T. S., Liu, X. B., Wong, B. Y., Ng, R. W., Yuen, A. P. & Wei, W. I. Mature miR-184 as potential oncogenic microRNA of squamous cell carcinoma of tongue. Clin. Cancer Res. 14, 2588–2592 (2008).
Chiyomaru, T., Enokida, H., Kawakami, K., Tatarano, S., Uchida, Y., Kawahara, K. et al. Functional role of LASP1 in cell viability and its regulation by microRNAs in bladder cancer. Urol. Oncol. [Epub ahead of print] PMID: 20843712 (2010).
Nurnberg, A., Kitzing, T. & Grosse, R. Nucleating actin for invasion. Nat. Rev. Cancer. 11, 177–187 (2011).
Millard, T. H., Behrendt, B., Launay, S., Futterer, K. & Machesky, L. M. Identification and characterisation of a novel human isoform of Arp2/3 complex subunit p16-ARC/ARPC5. Cell Motil. Cytoskeleton 54, 81–90 (2003).
Sheehan, D., Meade, G., Foley, V. M. & Dowd, C. A. Structure, function and evolution of glutathione transferases: implications for classification of non-mammalian members of an ancient enzyme superfamily. Biochem. J. 360, 1–16 (2001).
Sweeney, C., Nazar-Stewart, V., Stapleton, P. L., Eaton, D. L. & Vaughan, T. L. Glutathione S-transferase M1, T1, and P1 polymorphisms and survival among lung cancer patients. Cancer Epidemiol. Biomarkers Prev. 12, 527–533 (2003).
Ranganathan, S. & Tew, K. D. Immunohistochemical localization of glutathione S-transferases alpha, mu, and pi in normal tissue and carcinomas from human colon. Carcinogenesis 12, 2383–2387 (1991).
Miyanishi, K., Takayama, T., Ohi, M., Hayashi, T., Nobuoka, A., Nakajima, T. et al. Glutathione S-transferase-pi overexpression is closely associated with K-ras mutation during human colon carcinogenesis. Gastroenterology 121, 865–874 (2001).
Uchida, Y., Chiyomaru, T., Enokida, H., Kawakami, K., Tatarano, S., Kawahara, K. et al. MiR-133a induces apoptosis through direct regulation of GSTP1 in bladder cancer cell lines. Urol. Oncol. [Epub ahead of print] PMID: 21396852 (2011).
Acknowledgements
This study was supported by JSPS KAKENHI (C), 22591559 and was also supported in part by Global COE Program (Global Center for Education and Research in Immune System Regulation and Treatment), MEXT, Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Moriya, Y., Nohata, N., Kinoshita, T. et al. Tumor suppressive microRNA-133a regulates novel molecular networks in lung squamous cell carcinoma. J Hum Genet 57, 38–45 (2012). https://doi.org/10.1038/jhg.2011.126
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2011.126
Keywords
This article is cited by
-
CPEB2 enhances cell growth and angiogenesis by upregulating ARPC5 mRNA stability in multiple myeloma
Journal of Orthopaedic Surgery and Research (2023)
-
ARPC5 is transcriptionally activated by KLF4, and promotes cell migration and invasion in prostate cancer via up-regulating ADAM17
Apoptosis (2023)
-
RETRACTED ARTICLE: microRNA-133a exerts tumor suppressive role in oral squamous cell carcinoma through the Notch signaling pathway via downregulation of CTBP2
Cancer Gene Therapy (2022)
-
Alternative splicing of ceramide synthase 2 alters levels of specific ceramides and modulates cancer cell proliferation and migration in Luminal B breast cancer subtype
Cell Death & Disease (2021)
-
Constructing and validating a diagnostic nomogram for multiple sclerosis via bioinformatic analysis
3 Biotech (2021)