Abstract
Autosomal dominant nocturnal frontal lobe epilepsy is a familial partial epilepsy syndrome and the first human idiopathic epilepsy known to be related to specific gene defects. Clinically available molecular genetic testing reveals mutations in three genes, CHRNA4, CHRNB2 and CHRNA2. Mutations in CHRNA4 have been found in families from different countries; the Ser280Phe in an Australian, Spanish, Norwegian and Scottish families, and the Ser284Leu in a Japanese, Korean, Polish and Lebanese families. Clear evidence for founder effect was not reported among them, including a haplotype study carried out on the Australian and Norwegian families. Japanese and Koreans, because of their geographical closeness and historical interactions, show greater genetic similarities than do the populations of other countries where the mutation is found. Haplotype analysis in the two previously reported families showed, however, independent occurrence of the Ser284Leu mutation. The affected nucleotide was highly conserved and associated with a CpG hypermutable site, while other CHRNA4 mutations were not in mutation hot spots. Association with a CpG site accounts for independent occurrence of the Ser284Leu mutation.
Similar content being viewed by others
Introduction
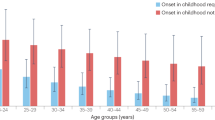
Autosomal dominant nocturnal frontal lobe epilepsy (ADNFLE; MIM 118504) is a familial partial epilepsy syndrome characterised by clusters of brief frontal lobe motor seizures during drowsiness or sleep.1, 2 Seizures—often misdiagnosed as other nocturnal motor activities such as parasomnia or night terror1, 2, 3, 4—vary from simple arousals to hyperkinetic activity with tonic or dystonic features. Onset usually occurs during the first two decades (mean age 10 years), but later onset has also been reported.5
ADNFLE is the first human idiopathic epilepsy known to be related to specific gene defects.6 Clinically available molecular genetic testing reveals mutations in three genes encoding the α4, β2 and α2 subunits of the neuronal nicotinic acetylcholine receptor (CHRNA4, CHRNB2 and CHRNA2, respectively).7, 8, 9 Overall mutations are found in less than 20% of individuals with ADNFLE/NFLE phenotypes, suggesting their genetic heterogeneity.10 Approximately 10–20% of patients have a positive family history and fewer than 5% a negative one.11
Among the four mutations in CHRNA4 (Ser280Phe, Ser284Leu, Leu291dup and Thr293Ile), Ser280Phe and Ser284Leu have been identified in several unrelated families (mutation names may be different from those of previous papers; we use NP_000735.1): the Ser280Phe in an Australian, Spanish, Norwegian and Scottish families,12, 13, 14, 15 and the Ser284Leu in a Japanese, Korean, Polish and Lebanese families.16, 17, 18, 19 It has been assumed that founder effect is not relevant to ADNFLE, because most of the families studied come from different countries,20 and a previous haplotype study of the Australian and Norwegian family14 showed no genetic connection.
Japanese and Koreans, because of their geographical closeness and historical interactions, show greater genetic similarities than do the populations of other countries where the mutation is found; indeed, previous studies between Japanese and Koreans have shown a closer genetic similarities than that between other east Asians.21, 22 Founder effects between the two countries have also been reported in several autosomal recessive diseases,23, 24 however, not in autosomal dominant ones. For ADNFLE patients, however, propagating their pathological genes is easier than it is for individuals with other more visible epilepsy syndromes. They have no seizures during the daytime, show normal neurological examinations and have a normal life expectancy and reproductive capacity. In addition, family members or the affected individual himself may believe that his symptoms are not an indication of illness and thus may not seek medical attention (or, knowing himself to be ill, he may deliberately refuse to reveal his condition to others).25 To determine whether founder effect is present in ADNFLE, we compare haplotype structures in the two previously reported Japanese and Korean families.
Materials and methods
We reviewed the medical histories and electroencephalogram findings of a three-generational family from Japan and one from Korea (pedigrees in Figure 1). Samples were collected from the two families and DNA was extracted from whole blood using standard protocols. A genetic analysis was carried out for seven members of the Japanese and eight of the Korean family. Among them, four of the Japanese had ADNFLE, as did eight of the Koreans. Four single nucleotide polymorphisms (SNPs) around the CHRNA4 were selected from a database of Japanese single nucleotide polymorphisms (JSNPS: http://snp.ims.u-tokyo.ac.jp/index_ja.html). To identify SNP-based haplotypes, four SNPs including rs 6089899, rs2093107, rs4809538 and rs4603829 (Figure 2) were typed by fluorescent sequencing on an automated DNA sequencer. Sequencing was carried out in both directions using four primer pairs. For comparison, the genotype frequencies of HapMap-JPT are listed in Table 1a.
We next examined the evolutionary conservation and phenotypic effect of Ser280Phe and Ser284Leu, which have shown recurrent occurrence in CHRNA4. The evolutionary conservation was estimated using phyloP score, available from UCSC Genome Browser (UCSC Genome Bioinformatics: http://genome.ucsc.edu/). The phenotypic effect of mutant protein was predicted using SIFT score (http://sift.jcvi.org/). In the phyloP, the conserved nucleotides are assigned positive scores, and the fast-evolving ones negative scores; the SIFT score ranges from 0 to 1. Deleterious amino-acid substitutions are assigned the scores of <0.05; tolerable substitutions are ⩾0.05.
We also checked hypermutable sites in CHRNA4. In the hypermutable sites analysis, we focused on CpG hypermutability. CpG is known as one of the major causes of codon substitution in mammalian genes, and is used to refer to cytosine followed by guanine in the Watson–Crick pair of a cytosine and guanine. The coding sequence of CHRNA4 was analysed and the adjacent nucleotides were taken into account to distinguish CpG hypermutations from non-CpG transitions.
Results
As shown in the pedigrees (Figure 1), there were five affected members and an obligated carrier in the Japanese family and nine affected members in the Korean one. Mutation in the two pedigrees showed an autosomal dominant mode with incomplete penetrance. All the affected individuals had a seizure semiology consistent with frontal lobe epilepsy and carried the Ser284Leu mutation in the CHRNA4. Both families had similar clinical manifestations, and their electroencephalogram findings were consistent with those of ADNFLE. They had brief motor seizures during sleep and no auras were reported. They shared features such as drug resistance and mental retardation, which are uncommon findings in ADNFLE. Still, SNP-based haplotypes were different in each family (Table 1b).
Ser280Phe and Ser284Leu showed higher phyloP scores (5.88 and 5.80, respectively) than the other nucleotides in CHRNA4 (mean, 0.08). SIFT scores were lesser than 0.05 (0.00, each). CpG dinucleotide sites are shown in Figure 2. Nine CpG dinucleotide sites were detected in the coding sequence of CHRNA4, three of them in the second transmembrane domain (M2). Among the four CHRNA4 mutations, only Ser284Leu mutation is associated with a CpG hypermutable site.
Discussion
All CHRNA4 mutations identified so far in ADNFLE have been located only in the M2 region, which has one of the most important functional roles in the neuronal nicotinic acetylcholine receptor . This receptor has a pentameric structure comprised of various combinations of alpha and beta subunits encoded by CHRNA4 and CHRNB2. The overall tertiary structure of nicotinic acetylcholine receptor subunits is similar; each subunit comprises four segments (M1 to M4) of the transmembrane domain and the N and C termini on the extracellular side of the membrane. M2 lines the central pore of the receptor and determines the ion selectivity of the receptor.26
The results obtained here do not support the hypothesis that the Ser284Leu mutations originated with a common founder. Association with a CpG site accounts for the independent occurrence of the Ser284Leu mutation. The other three mutations, however, are not associated with hypermutable sites such as homonucleotide runs, direct and inverted repeats and CpG dinucleotide sites.27 Thus, the repetitive occurrence of Ser280Phe is not easily understood.
To explain it, we need to take into account ADNFLE development. One possible explanation is that ADNFLE is caused by mutations on a few functionally important sites within the M2.14 Together with S280F, in vitro expression studies indicate that all CHRNA4 mutations increase receptor sensitivity to acethylcholine, suggesting gain of function.28, 29 In our results, both S280F and S284L showed high phyloP scores and low SIFT scores, indicating that the affected nucleotides are highly conserved and that their amino-acid substitutions will be deleterious.
Another explanation is that rare mutations have strong phenotypic effects in complex disorders.30 A recent report uncovered significant excess of rare variants in neurological disorders, providing a ‘rare allele-major effect model’, and suggesting that the rare variants or de novo mutations in neurologically expressed genes are more likely to accumulate.
This is the first report comparing the haplotype structure of Japanese and Korean ADNFLE families. Further functional studies will be required to ascertain the ADNFLE's pathophysiology.
References
Scheffer, I. E., Bhatia, K. P., Lopes-Cendes, I., Fish, D. R., Marsden, C. D., Andermann, F. et al. Autosomal dominant frontal epilepsy misdiagnosed as sleep disorder. Lancet 343, 515–517 (1994).
Scheffer, I. E., Bhatia, K. P., Lopes-Cendes, I., Fish, D. R., Marsden, C. D., Andermann, E. et al. Autosomal dominant nocturnal frontal lobe epilepsy. A distinctive clinical disorder. Brain 118 (Pt 1), 61–73 (1995).
Oldani, A., Zucconi, M., Ferini-Strambi, L., Bizzozero, D. & Smirne, S. Autosomal dominant nocturnal frontal lobe epilepsy: electroclinical picture. Epilepsia. 37, 964–976 (1996).
Hayman, M., Scheffer, I. E., Chinvarun, Y., Berlangieri, S. U. & Berkovic, S. F. Autosomal dominant nocturnal frontal lobe epilepsy: demonstration of focal frontal onset and intrafamilial variation. Neurology 49, 969–975 (1997).
Steinlein, O. K. Neuronal nicotinic receptors in human epilepsy. Eur. J. Pharmacol. 393, 243–247 (2000).
Steinlein, O. K., Mulley, J. C., Propping, P., Wallace, R. H., Phillips, H. A., Sutherland, G. R. et al. A missense mutation in the neuronal nicotinic acetylcholine receptor alpha 4 subunit is associated with autosomal dominant nocturnal frontal lobe epilepsy. Nat. Genet. 11, 201–203 (1995).
Steinlein, O., Sander, T., Stoodt, J., Kretz, R., Janz, D. & Propping, P. Possible association of a silent polymorphism in the neuronal nicotinic acetylcholine receptor subunit alpha4 with common idiopathic generalized epilepsies. Am. J. Med. Genet. 74, 445–449 (1997).
De Fusco, M., Becchetti, A., Patrignani, A., Annesi, G., Gambardella, A., Quattrone, A. et al. The nicotinic receptor beta 2 subunit is mutant in nocturnal frontal lobe epilepsy. Nat. Genet. 26, 275–276 (2000).
Aridon, P., Marini, C., Di Resta, C., Brilli, E., De Fusco, M., Politi, F. et al. Increased sensitivity of the neuronal nicotinic receptor alpha 2 subunit causes familial epilepsy with nocturnal wandering and ictal fear. Am. J. Hum. Genet. 79, 342–350 (2006).
Ottman, R., Hirose, S., Jain, S., Lerche, H., Lopes-Cendes, I., Noebels, J. L. et al. Genetic testing in the epilepsies—report of the ILAE Genetics Commission. Epilepsia. 51, 655–670 (2010).
Hirose, S. & Kurahashi, H. Autosomal dominant nocturnal frontal lobe epilepsy (April 2010) GeneReviews at GeneTests: Medical Genetics Information Resource [database online]. Copyright, University of Washington, Seattle, 1997–2010. Available at http://www.genetests.org.
Phillips, H. A., Scheffer, I. E., Berkovic, S. F., Hollway, G. E., Sutherland, G. R., Mulley, J. C. et al. Localization of a gene for autosomal dominant nocturnal frontal lobe epilepsy to chromosome 20q 13.2. Nat. Genet. 10, 117–118 (1995).
Saenz, A., Galan, J., Caloustian, C., Lorenzo, F., Marquez, C., Rodriguez, N. et al. Autosomal dominant nocturnal frontal lobe epilepsy in a Spanish family with a Ser252Phe mutation in the CHRNA4 gene. Arch. Neurol. 56, 1004–1009 (1999).
Steinlein, O. K., Stoodt, J., Mulley, J., Berkovic, S., Scheffer, I. E. & Brodtkorb, E. Independent occurrence of the CHRNA4 Ser248Phe mutation in a Norwegian family with nocturnal frontal lobe epilepsy. Epilepsia. 41, 529–535 (2000).
McLellan, A., Phillips, H. A., Rittey, C., Kirkpatrick, M., Mulley, J. C., Goudie, D. et al. Phenotypic comparison of two Scottish families with mutations in different genes causing autosomal dominant nocturnal frontal lobe epilepsy. Epilepsia. 44, 613–617 (2003).
Hirose, S., Iwata, H., Akiyoshi, H., Kobayashi, K., Ito, M., Wada, K. et al. A novel mutation of CHRNA4 responsible for autosomal dominant nocturnal frontal lobe epilepsy. Neurology 53, 1749–1753 (1999).
Cho, Y. W., Motamedi, G. K., Laufenberg, I., Sohn, S. I., Lim, J. G., Lee, H. et al. A Korean kindred with autosomal dominant nocturnal frontal lobe epilepsy and mental retardation. Arch. Neurol. 60, 1625–1632 (2003).
Phillips, H. A., Marini, C., Scheffer, I. E., Sutherland, G. R., Mulley, J. C. & Berkovic, S. F. A de novo mutation in sporadic nocturnal frontal lobe epilepsy. Ann. Neurol. 48, 264–267 (2000).
Rozycka, A., Skorupska, E., Kostyrko, A. & Trzeciak, W. H. Evidence for S284L mutation of the CHRNA4 in a white family with autosomal dominant nocturnal frontal lobe epilepsy. Epilepsia. 44, 1113–1117 (2003).
Combi, R., Dalpra, L., Tenchini, M. L. & Ferini-Strambi, L. Autosomal dominant nocturnal frontal lobe epilepsy—a critical overview. J. Neurol. 251, 923–934 (2004).
Saha, N. & Tay, J. S. Origin of the Koreans: a population genetic study. Am. J. Phys. Anthropol. 88, 27–36 (1992).
Kim, J. J., Verdu, P., Pakstis, A. J., Speed, W. C., Kidd, J. R. & Kidd, K. K. Use of autosomal loci for clustering individuals and populations of East Asian origin. Hum. Genet. 117, 511–519 (2005).
Inagaki, K., Suzuki, T., Ito, S., Suzuki, N., Fukai, K., Horiuchi, T. et al. OCA4: evidence for a founder effect for the p.D157N mutation of the MATP gene in Japanese and Korean. Pigment Cell. Res. 18, 385–388 (2005).
Khajoee, V., Ihara, K., Kira, R., Takemoto, M., Torisu, H., Sakai, Y. et al. Founder effect of the C9 R95X mutation in Orientals. Hum. Genet. 112, 244–248 (2003).
Thomas, P., Picard, F., Hirsch, E., Chatel, M. & Marescaux, C. Autosomal dominant nocturnal frontal lobe epilepsy. Rev. Neurol. (Paris) 154, 228–235 (1998).
Unwin, N. Acetylcholine receptor channel imaged in the open state. Nature 373, 37–43 (1995).
Rogozin, I. B. & Pavlov, Y. I. Theoretical analysis of mutation hotspots and their DNA sequence context specificity. Mutat. Res. 544, 65–85 (2003).
Moulard, B., Picard, F., le Hellard, S., Agulhon, C., Weiland, S., Favre, I. et al. Ion channel variation causes epilepsies. Brain Res. Rev. 36, 275–284 (2001).
Bertrand, D., Picard, F., Le Hellard, S., Weiland, S., Favre, I., Phillips, H. et al. How mutations in the nAChRs can cause ADNFLE epilepsy. Epilepsia. 43 (Suppl 5), 112–122 (2002).
Myers, R. A., Casals, F., Gauthier, J., Hamdan, F. F., Keebler, J., Boyko, A. R. et al. A population genetic approach to mapping neurological disorder genes using deep resequencing. PLoS. Genet. 7, e1001318 (2011).
Acknowledgements
We thank the members of the family for their cooperation in this study and Akiyo Hamachi and Minako Yonetani for technical assistance. This work was supported in part by Grant-in-Aid for Scientific Research (A) 21249062; Japan Society for the Promotion of Science(JSPS); ‘High-Tech Research Center’ Project for Private Universities—matching fund subsidy from the Ministry of Education, Culture, Sports, Science and Technology; 2006–2010—‘The Research Center for the Molecular Pathomechanisms of Epilepsy, Fukuoka University’; research Grants (21B-5) for Nervous and Mental Disorder from the Ministry of Health, Labor and Welfare; Health and Labor Science Research Grant (21210301); KB220001 from the Ministry of Health, Labor and Welfare; Adaptable and Seamless Technology Transfer Program through target-driven R&D (A-STEP) exploratory research; Japan Science and Technology Agency (JSP); International Research Fund for Subsidy of Kyushu University School of Medicine Alumini; The Japan Epilepsy Research Foundation (H22-005); and research Grant from Keimyung University, Korea.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Hwang, SK., Makita, Y., Kurahashi, H. et al. Autosomal dominant nocturnal frontal lobe epilepsy: a genotypic comparative study of Japanese and Korean families carrying the CHRNA4 Ser284Leu mutation. J Hum Genet 56, 609–612 (2011). https://doi.org/10.1038/jhg.2011.69
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2011.69
Keywords
This article is cited by
-
A review of the diverse genetic disorders in the Lebanese population: highlighting the urgency for community genetic services
Journal of Community Genetics (2015)
-
The Molecular Biology of Genetic-Based Epilepsies
Molecular Neurobiology (2014)
-
A multi-faceted approach to understanding male infertility: gene mutations, molecular defects and assisted reproductive techniques (ART)
Journal of Assisted Reproduction and Genetics (2014)