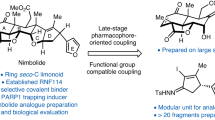
Abstract
GEX1A/herboxidiene (1) is a natural product isolated from Streptomyces sp. and has been reported to target the pre-mRNA splicing process. Although 1 was shown to have antitumor activity in vivo, weight loss was observed in mice when 1 was consecutively administered. We assumed that the carboxylic acid moiety was one of the causes of this toxicity. In this study, a series of amide, carbamate and urea analogues of 1 were synthesized and their antiproliferative activity was evaluated in vitro. The synthesis of urea analogues featured Curtius rearrangement following amine treatment with the one-pot procedure from 1. Furthermore, a structure–activity relationship study of the urea analogues revealed that the pharmacologically preferable basic side chains were acceptable and that compound 9g was equipotent to parent 1. These basic urea analogues would be promising leads for the development of novel antitumor agents.
Similar content being viewed by others
Introduction
Microorganisms are known to biosynthesize substances that are valuable for mankind. These natural products can be useful as drugs by themselves or sometimes as tools for the discovery of novel drug targets.1 Inhibitors of the pre-mRNA splicing process, which attracts attention as antitumor drug target, are one of the latter representatives. The pre-mRNA splicing inhibitors that originated from microorganisms are classified in three chemotypes. GEX1A (1, also known as herboxidiene2, 3) and its analogues isolated from a culture broth of Streptomyces sp. are the first reported inhibitors of this class.4 These polyketides have been shown to induce cell cycle arrest at G1 and G2/M phases in human cells5, 6 and to inhibit the pre-mRNA splicing process by binding to spliceosome-associated protein (SAP) 155, a subunit of SF3b, in the splicesome.7 Similarly, the second chemotype, FR901464 (2), and its analogues, spliceostatin A (3) and meayamycin (4), are known to interact with SAP 130 or SAP 155 to exert potent cytotoxicity.8, 9, 10, 11, 12, 13 The last molecules in this class are pladienolides (5, 6), which also binds to SAP 130, and its carbamate analogue E7107 (7) has been evaluated in clinical trials for the treatment of various types of cancers.14, 15, 16, 17, 18, 19 Thus, the inhibition of the pre-mRNA splicing process by the binding of these natural products to the SF3b subunit of the spliceosome has attracted considerable attention as a novel antitumor target.
Several synthetic studies of 1 have been performed to understand the structure–activity relationship and its improved analogues have been reported.20, 21, 22, 23, 24, 25, 26, 27, 28 In comparison with 7, however, there was room for the further modification of carboxylic acid, which was a characteristic moiety of 1. Moreover, the consecutive injection of 1 has been reported to induce weight loss in mice4 and we assumed that the carboxylic moiety would be related to this toxicity. We herein report the synthesis of deacidified analogues of 1 (Figure 1) and the evaluation of their antiproliferative activity against several cancer cell lines.
Results
The synthesis of GEX1A analogues was outlined in Scheme 1. Amides 8a and 8b were each synthesized with good yields using 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide with the corresponding amine. Urea 9a and 9b were easily obtained by Curtius rearrangement using diphenylphosphoryl azide,29 followed by treatment of the appropriate amines without the protection of the hydroxyl group at 18-position. In the process of synthesizing reverse amides 11a and 11b, the hydrolysis of an isocyanate group that came from Curtius rearrangement did not proceed well under acidic or basic conditions. Therefore, we decided to synthesize them via an imidazolyl urea intermediate 10, which was found to be hydrolyzed immediately under basic condition. Then, the addition of 4-(4,6-dimethoxy-1,3,5-triazin-2-yl)-4-methylmorpholinium chloride30 together with a corresponding carboxylic acid in phosphate buffer into the reactant of 10 afforded the desired reverse amides in one pot. As for carbamates 13a and 13b, the esterification of the carboxyl moiety and the protection of the secondary hydroxyl group with tert-butyldimethylsilyl were first carried out. Compound 12 was treated with LiAlH4 to afford alcohol and then converted into activated carbonate by treatment with p-nitrophenyl chloroformate and NaH. The introduction of the carbamate moiety was performed in the presence of triethylamine. Finally, the target compounds 13a and 13b were obtained after a deprotection step with tetra-n-butylammonium fluoride.
We first evaluated the antiproliferative activity of the compounds with neutral and basic side chains against HeLa S3 cells, which were also used in a previous study.28 In our initial analogue set, the potent activity of the urea linker compounds (9a, 9b) was fortunately found, while the activity decreased in the corresponding amides (8a, 8b), carbamates (13a, 13b) and reverse amides (11a, 11b) (Table 1). In general, acidic compounds tend to accumulate in the liver and this sometimes causes toxic side effects. In addition, solid tumors often form a specific microenvironment that is characterized by pH that is lower than physiological levels. This implies that the basic analogues of 1 would be more promising as antitumor therapeutics. Thus, we further evaluated the urea analogues with a basic side chain (such as 9b).
The synthesis of a series of urea analogues was easily accomplished by one-step Curtius rearrangement from the natural product. The antiproliferative activity of the urea analogues against HeLa S3 cells is shown in Table 2. We first evaluated the size of the linker ring from 5 to 7 (9c–e). The replacement of a piperazine ring with homopiperazine or pyrrolidine showed little impact on the in vitro potency (GI50=25–32 nmol l−1). Thus, the piperazinyl moiety was selected as a linker and we attempted to make a minor modification on its external nitrogen position to improve the in vitro potency of synthetic urea analogues. As a result of this modification, all of the substituents were tolerated and exhibited GI50 with the same potency as parent 1. The introduction of both a hydrogen bond donor (9f) and acceptors (9g, h) at the head resulted in a twofold increase in potency in comparison with 9c (GI50=9.0–16 nmol l−1). In particular, 9g that has a 1-cyanoethyl substituent exhibited the same potency as parent 1. The next point of interest was the tolerability of the bulky substituent at the terminus. The analogues with 2-morpholinoethyl (9i) or N-methylpiperidinyl (9j) moieties maintained high potency (GI50=12 nmol l−1). In addition, a hetero aromatic ring (9k) exhibited good tolerability (GI50=17 nmol l−1).
We next evaluated the antiproliferative activity of the selected urea analogues against four other cancer cell lines (Table 3). The GI50 values of the synthetic analogues exerted potent antiproliferative activity in the low nanomolar range against SK-MEL-2 (malignant melanoma), A549 (lung adenocarcinoma) and EBC-1 (lung squamous cell carcinoma). Although PC-3 (prostate adenocarcinoma) showed ~10-fold resistance to all of the compounds, no difference was observed in the order of the GI50 value among the urea analogues.
In conclusion, the deacidified modification of 1 with pharmacologically preferable basic side chains was acceptable and 9g was equipotent to parent 1. The findings suggest that these basic urea analogues will be promising leads for further evaluation.
Discussion
In the present study, we prepared and evaluated the antiproliferative activity of a series of carboxylic acid modified analogues of GEX1A (1), a natural product known to have an antitumor effect. Recently, 1 has been revealed to inhibit the pre-mRNA splicing process by targeting SAP 155, a subunit of SF3b in the splicesome.7 Two other structure-related natural products, FR901464 and pladienolide, have also been reported as the pre-mRNA splicing inhibitor and their analogues were examined as potential antitumor drugs. In particular, E7107, a two-ring extension analogue with a carbamate linker of pladienolide B, has entered clinical trials.14, 15, 16, 17, 18, 19 More recently, H3 Biomedicine (Cambridge, MA, USA) announced that the Food and Drug Administration accepted an IND application to begin clinical trials for a new pladienolide-based SF3B1 modulator H3B-8800 in 2016.31 This clearly indicates that the SF3b complex is a pharmacologically relevant protein and that the splicing factor is attractive target for antitumor drug discovery.
Although several semisynthetic or total synthetic studies for analogues of 1 have been carried out to improve their activity,20, 21, 22, 23, 24, 25, 26, 27, 28 few efforts have been made to modify the carboxylic acid moiety that is unique to 1 among the splicing inhibitor groups. In addition, acidic compounds generally tend to accumulate in the liver, but not in tumor microenvironment. This biodistribution preference might cause hepatotoxicity or a weak antitumor effect in vivo. We therefore focused on the synthesis of deacidified analogues of 1 by modifying the carboxylic acid. In this study, we evaluated the antiproliferative activities against cancer cell lines in vitro because the splicing inhibitory potency of the analogues of 1 has been shown to be well correlated with their antiproliferative activity.28
The modification of 1 consisted of two steps: (1) linker screening and (2) side-chain optimization (Figure 1). In the first step, the urea linker was selected because of their remaining potency (Table 1). For the consecutive examination of the side chain, the urea analogues with a variety of basic side chains were easily synthesized from 1 through the one-pot and two-step procedures using the corresponding amines in the second step (Table 2).
As a consequence of the in vitro antiproliferative assay, all of the basic urea analogues, especially 9g, showed potent activity against various human cancer cell lines. We expect that these basic analogues will possess the improved biodistribution preference and PK parameters. Thus, the further evaluation of the in vivo efficacy will be reported in the future.
Materials and methods
Unless otherwise indicated, all of the reagents and solvents were purchased from commercial sources and were used without further purification. 1 used in this study was isolated from a culture extract of a Streptomyces sp. GEX1 strain in our laboratory.4 Moisture- or oxygen-sensitive reactions were carried out under an argon or nitrogen gas atmosphere. Silica gel column chromatography was performed using Purif-α2 (SHOKO Scientific, Yokohama, Japan) with prepacked cartridge columns. The 1H-NMR spectra were measured using a JNM-AL300 Instrument (JEOL, Tokyo, Japan) at 300 MHz in CDCl3. The chemical shifts are expressed in p.p.m. (δ) relative to an internal standard, tetramethylsilane (p.p.m.=0.00). The mass spectra were measured in ESI mode using an LC-MS system (Nihon Waters KK, Tokyo, Japan).

Reagents and conditions. (a) amines, EDCI·HCl, 1-hydroxybenzotriazole, DMF, rt; (b) DPPA, triethylamine, toluene, 110 °C and then the addition of amines, 0 °C; (c) DPPA, triethylamine, toluene, 110 °C and then the addition of imidazole, 0 °C; (d) NaOHaq., THF, rt and then the addition of carboxylic acids, DMT-MM, phosphate buffer, MeOH, rt; (e) Me2SO4, K2CO3, N,N-dimethylacetamide, 90 °C; (f) tert-butyldimethylsilyl trifluoromethanesulfonate, 2,6-lutidine, CH2Cl2, 0 °C; (g) LiAlH4, THF, 0 °C; (h) p-NO2C6H4OC(O)Cl, NaH, THF, 0 °C; (i) amines, triethylamine, THF, rt; (j) TBAF, THF, 60 °C. DMT-MM, 4-(4,6-dimethoxy-1,3,5-triazin-2-yl)-4-methylmorpholinium chloride; DPPA, diphenylphosphoryl azide; EDCI, 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide; TBAF, tetra-n-butylammonium fluoride.
Synthesis of urea analogues
To the suspension of 1 (14 mg, 32 μmol) in toluene (5.0 ml), triethylamine (13 μl, 94 μmol) and diphenylphosphoryl azide (21 μl, 94 μmol) were added in a dropwise manner at room temperature. This mixture was stirred at 110 °C for 3 h, and then cooled to 0 °C. Morpholine (50 μl, 0.58 mmol) was added to the mixture and stirred at 0 °C for 30 min. The reaction was quenched by the addition of a saturated NH4Cl aqueous solution, and the reactant was extracted with CH2Cl2, washed with brine and dehydrated with MgSO4. After filtration, the filtrate was evaporated under reduced pressure. The residue was purified by silica gel column chromatography to yield 15 mg (89%) of 9a.
Urea analogues9b, 9f, 9g, 9h and 9i were prepared similarly to 9a with 1-cyclohexylpiperazine, N-(2-hydroxyethyl)piperazine, 3-(piperazin-1-yl)propanenitrile, 1-(2-dimethylaminoethyl)piperazine and 1-[2-(morpholin-4-yl)ethyl]piperazine instead of morpholine, with a total yield of 77%, 56%, 54%, 60% and 63%, respectively.
Urea analogue 9a
1H-NMR δ: 6.25 (dd, J=15.0, 10.9 Hz, 1H), 5.90 (d, J=10.9 Hz, 1H), 5.47 (dd, J=15.0, 8.8 Hz, 1H), 4.90 (t, J=5.1 Hz, 1H), 3.91–3.79 (m, 1H), 3.72–3.65 (m, 4H), 3.57–3.40 (m, 5H), 3.36–3.27 (m, 5H), 3.14–3.01 (m, 1H), 2.97 (t, J=5.5 Hz, 1H), 2.58–2.38 (m, 2H), 1.93–1.81 (m, 2H), 1.72 (d, J=1.3 Hz, 3H), 1.65–1.46 (m, 3H), 1.38–1.13 (m, 9H), 1.06 (d, J=6.6 Hz, 3H), 0.87 (d, J=6.9 Hz, 3H), 0.67 (d, J=6.6 Hz, 3H). LC-MS (ESI): m/z 523 (M+H)+.
Urea analogue 9b
1H-NMR δ: 6.25 (dd, J=15.0, 10.9 Hz, 1H), 5.90 (d, J=10.9 Hz, 1H), 5.47 (dd, J=15.0, 8.8 Hz, 1H), 4.93–4.87 (m, 1H), 3.91–3.79 (m, 1H), 3.59–3.28 (m, 10H), 3.12–3.03 (m, 1H), 2.97 (t, J=5.3 Hz, 1H), 2.59–2.51 (m, 5H), 2.49–2.37 (m, 1H), 2.32–2.22 (m, 1H), 1.94–1.76 (m, 6H), 1.72 (d, J=1.0 Hz, 3H), 1.67–1.46 (m, 4H), 1.37–1.11 (m, 14H), 1.06 (d, J=6.6 Hz, 3H), 0.87 (d, J=6.9 Hz, 3H), 0.67 (d, J=6.6 Hz, 3H). LC-MS (ESI): m/z 604 (M+H)+.
Urea analogue 9f
1H-NMR δ: 6.25 (dd, J=15.0, 10.9 Hz, 1H), 5.90 (d, J=10.9 Hz, 1H), 5.47 (dd, J=15.0, 8.7 Hz, 1H,), 4.91 (t, J=4.9 Hz, 1H), 3.90–3.79 (m, 1H), 3.67–3.60 (m, 2H), 3.54 (s, 3H), 3.51–3.34 (m, 6H), 3.30 (d, J=9.9 Hz, 1H), 3.13–3.01 (m, 1H), 2.98 (t, J=5.3 Hz, 1H), 2.60–2.35 (m, 8H), 1.90–1.82 (m, 2H), 1.72 (d, J=0.7 Hz, 3H), 1.65–1.45 (m, 3H), 1.39–1.21 (m, 3H), 1.29 (s, 3H), 1.18 (d, J=6.3 Hz, 3H), 1.06 (d, J=6.9 Hz, 3H), 0.87 (d, J=6.9 Hz, 3H), 0.67 (d, J=6.6 Hz, 3H). LC-MS (ESI): m/z 566 (M+H)+.
Urea analogue 9g
1H-NMR δ: 6.25 (dd, J=15.0, 10.8 Hz, 1H), 5.90 (d, J=10.8 Hz, 1H), 5.47 (dd, J=15.0, 8.8 Hz, 1H), 4.88 (br s, 1H), 3.92–3.78 (m, 1H), 3.58–3.25 (m, 6H), 3.54 (s, 3H), 3.30 (d, J=9.5 Hz, 1H), 3.14–3.02 (m, 1H), 3.00–2.94 (m, 1H), 2.75–2.65 (m, 2H), 2.60–2.37 (m, 8H), 1.96–1.80 (m, 2H), 1.72 (s, 3H), 1.65–1.44 (m, 3H), 1.39–1.14 (m, 3H), 1.29 (s, 3H), 1.18 (d, J=6.2 Hz, 3H), 1.11–1.01 (m, 3H), 0.93–0.81 (m, 3H), 0.73–0.60 (m, 3H). LC-MS (ESI): m/z 575 (M+H)+.
Urea analogue 9h
1H-NMR δ: 6.25 (dd, J=15.0, 10.8 Hz, 1H), 5.90 (d, J=10.8 Hz, 1H), 5.47 (dd, J=15.0, 8.8 Hz, 1H), 4.88 (br s, 1H), 3.91–3.79 (m, 1H), 3.59–3.26 (m, 6H), 3.54 (s, 3H), 3.30 (d, J=9.9 Hz, 1H), 3.15–3.01 (m, 1H), 2.97 (t, J=5.5 Hz, 1H), 2.56 (d, J=9.5 Hz, 1H), 2.53–2.39 (m, 9H), 2.28 (s, 6H), 1.97–1.79 (m, 2H), 1.72 (d, J=0.7 Hz, 3H), 1.64–1.45 (m, 3H), 1.33–1.15 (m, 3H), 1.29 (s, 3H), 1.18 (d, J=6.6 Hz, 3H), 1.06 (d, J=6.6 Hz, 3H), 0.87 (d, J=7.0 Hz, 3H), 0.66 (d, J=6.6 Hz, 3H). LC-MS (ESI): m/z 593 (M+H)+.
Urea analogue 9i
1H-NMR δ: 6.25 (dd, J=15.0, 10.6 Hz, 1H), 5.90 (d, J=10.6 Hz, 1H), 5.47 (dd, J=15.0, 8.8 Hz, 1H), 4.88 (br s, 1H), 3.90–3.80 (m, 1H), 3.75–3.67 (m, 4H), 3.54 (s, 3H), 3.52–3.32 (m, 6H), 3.30 (d, J=9.9 Hz, 1H), 3.13–3.01 (m, 1H), 2.97 (t, J=5.3 Hz, 1H), 2.58–2.44 (m, 14H), 1.96–1.79 (m, 2H), 1.72 (d, J=0.7 Hz, 3H), 1.63–1.45 (m, 3H), 1.39–1.15 (m, 3H), 1.29 (s, 3H), 1.18 (d, J=6.6 Hz, 3H), 1.06 (d, J=6.6 Hz, 3H), 0.87 (d, J=6.6 Hz, 3H), 0.66 (d, J=6.6 Hz, 3H). LC-MS (ESI): m/z 635 (M+H)+.
The in vitro antiproliferative assay
The antiproliferative activity against human cervical adenocarcinoma HeLa S3 cells, lung adenocarcinoma A549 cells, lung squamous cell carcinoma EBC-1 cells, prostate adenocarcinoma PC-3 cells and malignant melanoma SK-MEL-2 cells was measured by WST-1 or XTT assays. Briefly, 500 cells per well in minimum essential medium containing 10% fetal bovine serum (FBS) for HeLa S3, 1000 cells per well in Ham's F-12K (Kaighn's) medium containing 10 vol% FBS for A549, 2000 cells per well in minimum essential medium Eagle’s containing 10 vol% FBS for EBC-1, 4000 cells per well in Ham's F-12K (Kaighn's) medium containing 10 vol% FBS for PC-3 and 2000 cells per well in minimum essential medium Eagle’s containing 10 vol% FBS for SK-MEL-2 cells were inoculated onto 96-microwell plates (Nalge Nunc International KK, Tokyo, Japan). The plates were incubated in a CO2 incubator at 37 °C for 24 h, and then GEX1A analogues diluents or vehicle control (culture medium supplemented with 0.5 vol% of DMSO) were added to the wells (n=3). The plates were incubated in a CO2 incubator at 37 °C for 72 h and the cell viability was measured with WST-1 or XTT assays (Roche Diagnostics, Tokyo, Japan). The OD at 450 nm in reference to 650 nm was measured by a SpectraMax 340PC384 microplate Spectrophotometer (Molecular Devices, Tokyo, Japan). The cellular proliferation over 72 h was calculated with the subtraction of the OD value at 0 h (OD0) from the OD value at 72 h (OD72). The cellular proliferation of the vehicle control was set at 100% and the GI50 value was calculated. All media and FBS were purchased from Life Technologies Japan (Tokyo, Japan).
References
Ōmura, S. A splendid gift from the earth: the origins and impact of the avermectins (Nobel Lecture). Angew. Chem. Int. Ed. 55, 10190–10209 (2016).
Isaac, B. G., Ayer, S. W., Elliott, R. C. & Stonard, R. J. Herboxidiene: a potent phytotoxic polyketide from Streptomyces sp. A7847. J. Org. Chem. 57, 7220–7226 (1992).
Miller-Wideman, M. et al. Herboxidiene, a new herbicidal substance from Streptomyces chromofuscus A7847. Taxonomy, fermentation, isolation, physico-chemical and biological properties. J. Antibiot. 45, 914–921 (1992).
Sakai, Y. et al. GEX1 compounds, novel antitumor antibiotics related to herboxidiene, produced by Streptomyces sp. I. Taxonomy, production, isolation, physicochemical properties and biological activities. J. Antibiot. 55, 855–862 (2002).
Sakai, Y. et al. GEX1 compounds, novel antitumor antibiotics related to herboxidiene, produced by Streptomyces sp. II. The effects on cell cycle progression and gene expression. J. Antibiot. 55, 863–872 (2002).
Horiguchi, T., Shirasaki, M. & Tanida, S. TAN-1609 (Herboxidiene): a microbial polyketide which blocks the cell-cycle at G2 phase in human and murine tumor cells. Takeda Kenkyusyo Ho 55, 149–159 (1996).
Hasegawa, M. et al. Identification of SAP155 as the target of GEX1A (Herboxidiene), an antitumor natural product. ACS Chem. Biol. 6, 229–233 (2011).
Nakajima, H. et al. New antitumor substances, FR901463, FR901464 and FR901465. I. Taxonomy, fermentation, isolation, physico-chemical properties and biological activities. J. Antibiot. 49, 1196–1203 (1996).
Nakajima, H. et al. New antitumor substances, FR901463, FR901464 and FR901465. II. Activities against experimental tumors in mice and mechanism of action. J. Antibiot. 49, 1204–1211 (1996).
Nakajima, H. et al. New antitumor substances, FR901463, FR901464 and FR901465. III. Structures of FR901463, FR901464 and FR901465. J. Antibiot. 50, 96–99 (1997).
Motoyoshi, H. et al. Structure–activity relationship for FR901464: a versatile method for the conversion and preparation of biologically active biotinylated probes. Biosci. Biotechnol. Biochem. 68, 2178–2182 (2004).
Albert, B. J., Sivaramakrishnan, A., Naka, T., Czaicki, N. L. & Koide, K. Total syntheses, fragmentation studies, and antitumor/antiproliferative activities of FR901464 and its low picomolar analogue. J. Am. Chem. Soc. 129, 2648–2659 (2007).
Kaida, D. et al. Spliceostatin A targets SF3b and inhibits both splicing and nuclear retention of pre-mRNA. Nat. Chem. Biol. 3, 576–583 (2007).
Sakai, T. et al. Pladienolides, new substances from culture of Streptomyces platensis Mer-11107. I. Taxonomy, fermentation, isolation and screening. J. Antibiot. 57, 173–179 (2004).
Sakai, T., Asai, N., Okuda, A., Kawamura, N. & Mizui, Y. Pladienolides, new substances from culture of Streptomyces platensis Mer-11107. II. Physico-chemical properties and structure elucidation. J. Antibiot. 57, 180–187 (2004).
Mizui, Y. et al. Pladienolides, new substances from culture of Streptomyces platensis Mer-11107. III. In vitro and in vivo antitumor activities. J. Antibiot. 57, 188–196 (2004).
Asai, N. et al. Stereochemistry of pladienolide B. J. Antibiot. 60, 364–369 (2007).
Kanada, R. M. et al. Total synthesis of the potent antitumor macrolides pladienolide B and D. Angew. Chem. Int. Ed. 46, 4350–4355 (2007).
Kotake, Y. et al. Splicing factor SF3b as a target of the antitumor natural product pladienolide. Nat. Chem. Biol. 3, 570–575 (2007).
Blakemore, P. R., Kocieński, P. J., Morley, A. & Muir, K. A synthesis of herboxidiene. J. Chem. Soc, Perkin Trans. 1, 955–968 (1999).
Banwell, M., McLeod, M., Premraj, R. & Simpson, G. Total synthesis of herboxidiene, a complex polyketide from Streptomyces species A7847. Pure Appl. Chem. 72, 1631–1634 (2000).
Zhang, Y. & Panek, J. S. Total synthesis of herboxidiene/GEX 1A. Org. Lett. 9, 3141–3143 (2007).
Murray, T. J. & Forsyth, C. J. Total synthesis of GEX1A. Org. Lett. 10, 3429–3431 (2008).
Ghosh, A. K. & Li, J. A stereoselective synthesis of (+)-herboxidiene/GEX1A. Org. Lett. 13, 66–69 (2011).
Pellicena, M., Krämer, K., Romea, P. & Urpí, F. Total synthesis of (+)-herboxidiene from two chiral lactate-derived ketones. Org. Lett. 13, 5350–5353 (2011).
Premraj, R., McLeod, M. D., Simpson, G. W. & Banwell, M. G. A total synthesis of herboxidiene methyl ester. Heterocycles 85, 2949–2976 (2012).
Gao, Y., Vogt, A., Forsyth, C. J. & Koide, K. Comparison of splicing factor 3b inhibitors in human cells. ChemBioChem. 14, 49–52 (2013).
Lagisetti, C. et al. Pre-mRNA splicing-modulatory pharmacophores: the total synthesis of herboxidiene, a pladienolide-herboxidiene hybrid analog and related derivatives. ACS Chem. Biol. 9, 643–648 (2014).
Shioiri, T., Ninomiya, K. & Yamada, S. Diphenylphosphoryl azide. New convenient reagent for a modified Curtius reaction and for the peptide synthesis. J. Am. Chem. Soc. 94, 6203–6205 (1972).
Kunishima, M., Kawachi, C., Hioki, K., Terao, K. & Tani, S. Formation of carboxamides by direct condensation of carboxylic acids and amines in alcohols using a new alcohol- and water-soluble condensing agent: DMT-MM. Tetrahedron 57, 1551–1558 (2001).
H3 Bioscience Pipeline. Available at:https://www.h3biomedicine.com/science/.Accessed on 12 September 2016.
Acknowledgements
We are grateful to Drs Hayato Yabuuchi, Tetsuo Yoshida, Ryuichiro Nakai and Yutaka Kanda for their helpful support and discussions.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
We would like to dedicate this article to Professor Satoshi Ōmura in celebration of his Nobel Prize in Physiology or Medicine 2015.
Supplementary Information accompanies the paper on The Journal of Antibiotics website
Supplementary information
Rights and permissions
About this article
Cite this article
Imaizumi, T., Nakagawa, H., Hori, R. et al. The synthesis and evaluation of the antiproliferative activity of deacidified GEX1A analogues. J Antibiot 70, 675–679 (2017). https://doi.org/10.1038/ja.2016.166
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2016.166
This article is cited by
-
Molecular mechanisms by which splice modulator GEX1A inhibits leukaemia development and progression
British Journal of Cancer (2022)