Abstract
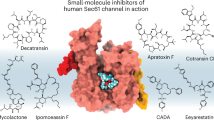
The rapid rise of antibiotic-resistant bacteria is one of the major concerns in modern medicine. Therefore, to treat bacterial infections, there is an urgent need for new antibacterials–preferably directed against alternative bacterial targets. One such potential target is the preprotein translocation motor SecA. SecA is a peripheral membrane ATPase and a key component of the Sec secretion pathway, the major route for bacterial protein export across or into the cytoplasmic membrane. As SecA is essential for bacterial viability, ubiquitous and highly conserved in bacteria, but not present in eukaryotic cells, it represents an attractive antibacterial target. Using an in silico approach, we have defined several potentially druggable and conserved pockets on the surface of SecA. We show that three of these potentially druggable sites are important for SecA function. A starting collection of ~500 000 commercially available small-molecules was virtually screened against a predicted druggable pocket in the preprotein-binding domain of Escherichia coli SecA using a multi-step virtual ligand screening protocol. The 1040 top-scoring molecules were tested in vitro for inhibition of the translocation ATPase activity of E. coli SecA. Five inhibitors of the translocation ATPase, and not of basal or membrane ATPase, were identified with IC50 values <65 μm. The most potent inhibitor showed an IC50 of 24 μm. The antimicrobial activity was determined for the five most potent SecA inhibitors. Two compounds were found to possess weak antibacterial activity (IC50 ~198 μm) against E. coli, whereas some compounds showed moderate antibacterial activity (IC50 ~100 μm) against Staphylococcus aureus.
Similar content being viewed by others
Introduction
The fight against bacterial infections is a major concern in modern medicine due to the rapid rise of antibiotic-resistant bacteria.1 The problem of antibiotic resistance expanded since the 1980s with an increase in the number of infections due to methicillin-resistant Staphylococcus aureus, vancomycin-resistant Enterococcus and fluoroquinolone-resistant Pseudomonas aeruginosa.2 The development of drug resistance is a natural phenomenon as bacteria have a short life cycle and the ability to adapt quickly to changes in the environment.1 The phenomenon is further accelerated by the inappropriate use of antibiotics. Currently, >70% of all pathogenic bacteria are resistant to one or more commercially available antibiotics and pose a severe threat to the general health and welfare.3
As resistance has developed against all the currently available antibiotics, there is an urgent need for new antibacterial drugs. Preferably these should have a novel mode of action to avoid cross-resistance with existing antibacterial agents. Nevertheless, the antibiotic research activities of the pharmaceutical industry are decreasing and the number of newly approved antibiotics dwindled sharply by 2010.4
A potential target for the development of novel antibiotics is the preprotein translocation motor ATPase SecA.5, 6, 7 SecA is an attractive antibacterial target as it is essential for bacterial viability, ubiquitous and highly conserved in bacteria, and absent in eukaryotes. Moreover, SecA meets also the other criteria that are considered to be desirable for a drug target:7 its function and structure are well understood; it exists in a membrane-bound state that might be more easily accessible to drugs; it is stable in vitro and amenable to high-throughput screening; and several interaction partners have been identified, providing multiple options for potential targeting sites.
SecA is one of the key components of the Sec pathway, the major route for bacterial proteins that are translocated across or into the cytoplasmic membrane. SecA converts the chemical energy of ATP to mechanical work of ‘pushing’ preproteins through a channel in the bacterial cytoplasmic membrane, formed by the integral membrane proteins SecY, SecE and SecG. These Sec-dependent preproteins contain N-terminal signal peptides and are translocated in a non-native state.8, 9 After translocation of the preprotein into the cytoplasmic membrane, the periplasm (Gram− bacteria) or the extracellular milieu (Gram+ bacteria), the signal peptide is cleaved off and the exported protein folds into its native state.8
The enzymatic activity of SecA can be dissected biochemically in three distinct states depending on its role during protein translocation. When SecA is circulating in the cytoplasm, it is not involved in the translocation process. In this cytoplasmic state, the ATPase activity of SecA is suppressed by intramolecular mechanisms (basal ATPase activity). However, the basal ATPase activity is slightly stimulated upon binding to negatively charged phospholipids and SecYEG (membrane ATPase). The binding of a single translocation-competent preprotein to membrane-bound dimeric SecA10 results in maximal stimulation of the ATPase activity (translocation ATPase).11
SecA contains four different structural domains as following: the nucleotide-binding domain (NBD), the intramolecular regulator of ATPase 2 (IRA2), the preprotein-binding domain (PBD) and the C-domain (Figure 1a). The NBD together with IRA2 form the so-called DEAD (Asp-Glu-Ala-Asp) motor that is structurally and functionally related to the ATPase domain of DEAD helicases.12 The PBD and the C-domain, which are absent from other helicases, confer SecA with the ability to handle preproteins and regulate its enzymatic activity.13
Cylinder representation of ecSecA structures showing the various conformational states. (a) The structure of ecSecA consists of four domains as following: the nucleotide-binding domain (NBD; dark blue), the intramolecular regulator of ATPase 2 (IRA2; cyan), the preprotein-binding domain (PBD; magenta) and the C-domain (green). Here a homology model of ecSecA is shown (the DEAD motor and C-domain are based on bsSecA, PDB ID: 1M6N; PBD is based on ecSecA, PDB ID: 2VDA). (b) Homology model of ecSecA (same as a) with a ‘closed’ DEAD motor. (c) ecSecA NMR structure (PDB ID: 2VDA) with an ‘open’ DEAD motor. (d) Homology model of ecSecA (based on Thermotoga maritima SecA, PDB ID: 3DIN) in a closed PBD conformation. (e) Homology model of ecSecA (modeled like a) in an open PBD conformation. (f) Homology model of ecSecA (based on bsSecA, PDB ID: 1M6N) in a wide-open PBD conformation.
SecA has been visualized by X-ray crystallography and NMR in various conformations. Two different states of the SecA DEAD motor have been observed: a ‘closed’14 (Figure 1b) and an ‘open’15 conformation (Figure 1c). SecA has also been visualized in a ‘closed’16 (Figure 1d), an ‘open’15, 17 (Figure 1e) and a ‘wide-open’ conformation14 (Figure 1f), depending on the position of the PBD that swivels ~70° around a slender stem that spouts out of the NBD (Sardis et al., submitted).
Several different strategies have been employed to identify novel SecA inhibitors, including in vitro, in vivo and in silico approaches. In vitro screening has led to the identification of the first natural inhibitor of SecA, the equisetin derivative CJ-21 058, which inhibits the translocation ATPase activity of Escherichia coli SecA (ecSecA; IC50 of 15 μg ml−1) and showed antibacterial activity against S. aureus and Enterococcus faecalis (MIC of 5 μg ml−1).18
An in vitro high-throughput screening of~27 000 small-molecules against ecSecA(W775A), an elevated basal ATPase activity derivative,19 identified several inhibitors of the ATPase activities of ecSecA (IC50 values of 50–150 μm).20, 21 In addition, a small-scale in vitro screen of synthesized thiazolo[4,5-d]pyrimidine derivatives has led to the identification of several inhibitors, including one with an IC50 of 135 μm against the translocation ATPase activity of ecSecA.22
The fluorescein analogs rose bengal and erythrosin B (IC50 of 0.9 and 10 μm respectively; inhibiting ecSecA translocation ATPase), were identified from an in vitro screen using a truncated ecSecA variant with elevated ATPase activity. Rose bengal inhibited Bacillus subtilis growth with a MIC value of 3.1 μm.23
In vivo approaches have resulted in the discovery of pannomycin,24 a natural product that is structurally similar to the equisetin derivative CJ-21 058. This secondary metabolite from the fungus Geomyces pannorum, which was identified using a two-plate differential sensitivity antisense assay, shows very weak antibacterial activity (mm range). In another study, an in vivo high-throughput screening using a SecA–LacZ reporter fusion was used to identify Sec-dependent translocation inhibitors.25 However, all identified molecules disrupt the bacterial membrane.
The availability of high-resolution structures of various SecAs has enabled in silico structure-based virtual ligand screening (SB-VLS) approaches to discover SecA inhibitors. The first SecA inhibitors discovered via SB-VLS were identified by screening of 60 000 compounds from the MayBridge database against the ATPase cleft of the helicase motor of ecSecA (PDB ID: 2FSG). This screen and subsequent optimization has resulted in the discovery of inhibitors of the ecSecA basal ATPase with IC50 values of ~20–60 μm.26, 27
Another VLS screen directed against the ATPase cleft of a SecA homology model identified inhibitors of Candidatus Liberibacter asiaticus SecA (claSecA), a Gram− citrus pathogen.28 In total 5016 small-molecules from the ChemBridge and Specs databases were screened and a potent compound with an IC50 value of 2.5 μm was identified. Also, an optimized homology model of claSecA was used to screen 20 000 compounds from the ZINC database against the ATPase site.29 Five basal ATPase inhibitors were identified (IC50 values, 0.25–0.92 μm), which showed antimicrobial activity against Agrobacterium tumefaciens, with MBC values ranging from 128 to 256 μg ml−1.
Here we present a SB-VLS approach to identify SecA inhibitors. We first determined the presence of potential druggable pockets on the surface of 11 available SecA structures of various bacterial species. In each SecA structure, 6–15 potential druggable pockets were identified of which six are highly conserved. Three of them were shown to be important for in vivo function of ecSecA and saSecA1 and one of them was tested for its druggability. VLS was performed against the signal peptide-binding region of ecSecA that was identified using NMR (PDB ID: 2VDA). After docking and scoring of the compounds, the best 1040 molecules were tested in vitro for inhibition of the translocation ATPase activity of ecSecA. Five small-molecules were identified that specifically inhibited the translocation ATPase activity of ecSecA with IC50 values ⩽65 μm. The most potent, specific inhibitor showed an IC50 of 24 μm. The compounds inhibited growth of Gram− and Gram+ bacteria with moderate to high IC50 values of 50–200 μm.
Materials and Methods
Computational procedures
Ten SecA crystal/NMR structures from the RCSB Protein Data Bank30 and one SecA homology model from SWISS-MODEL31 were investigated using ICM (Molsoft, San Diego, CA, USA). These 11 structures are described in Supplementary Table S1. The presence of binding pockets on the surface of SecA was predicted using the icmPocketFinder utility (standard settings).32 Potential druggable pockets were defined based on their druggability index (for example, appropriate size and chemical nature to bind drug-like small-molecules).
Comparison of the different potential druggable pockets was performed by looking at conservation of corresponding amino acids defining the pocket in the various SecA structures.
Structure-based virtual ligand screening
To discover novel SecA inhibitors, we initiated a SB-VLS of a selected pocket (Site III, Figure 2) on the surface of ecSecA, which we hypothesized to be druggable. The SB-VLS was performed according to an optimized consensus screening protocol.33, 34 In short, as a starting point for the selection of new SecA ligands, we used the in silico small-molecules collection from the Express Pick ChemBridge database (www.chembridge.com), version January 2010, of which each in silico-selected compound can be purchased, and the solution structure of the SecA-signal peptide complex (PDB ID: 2VDA), which was retrieved from the RCSB Protein Data Bank, PDB. The collection of compounds was first filtered using the ADME/Tox open-source FAF-Drugs2 program35 to remove non-drug-like molecules: compounds with 1 Lipinski violation or with reactive groups were, thus, removed at this step. Next, the OpenEye OMEGA conformer generation software36 was used to generate three-dimensional multiconformer structures for each of the small-molecules and to add hydrogen atoms and Gasteiger partial charges.36 We generated eight different normal mode perturbed starting structures of the SecA structure with ElNemo (http://www.elnemo.org) after manual removal of the bound peptide from the starting structure. Docking then proceeded via the FRED rigid body docking program37 to dock the pregenerated multiconformer library on the target structures. After scoring of all SecA-compound complexes, for each target, the top 40 000 compounds were subjected to flexible docking and scoring by the Surflex program.38 A consensus score for the different targets was then calculated by averaging of all docking score per compound and a final scoring list was produced.
Potential druggable pockets on the surface of SecA. Ribbon structures are shown for ecSecA (a) and saSecA1 (b), and the potential druggable pockets are shown as colored densities. Conserved pockets between ecSecA and saSecA1 are shown in the same colors. The three sites in ecSecA and saSecA1 that were selected for further functional characterization are indicated. Site I is located at the interface of the NBD and IRA2; Site II is located at the interface of the PBD and the C-domain; and Site III is located in the PBD. IRA2, intramolecular regulator of ATPase 2; NBD, nucleotide-binding domain; PBD, preprotein-binding domain.
2D similarity search
An in silico two-dimensional (2D) similarity search was performed against the ChemBridge database to identify compounds with better inhibitory activity than the identified hit molecules (parent molecules). To this end, we used the online 2D similarity search engine from ChemBridge (www.hit2lead.com), using standard parameters.
Bacterial strains and culture conditions
Different bacterial strains were used for this work (Supplementary Table S3). The cells were grown in different media including Luria Bertani (LB; 1% tryptone; 0.5% yeast extract; 85 mm NaCl), NZY+ medium (1% casein hydrolysate; 0.5% yeast extract; 85 mm NaCl; 12.5 mm MgCl2; 12.5 mm MgSO4; 0.4% glucose; pH 7.5), HS medium (1 × S-base (0.2% (NH4)2SO4; 1.4% K2HPO4; 0.6% KH2PO4; 0.1% sodium citrate; 1 mm MgSO4); 0.5% glucose; 0.005% l-tryptophan; 0.02% casein hydrolysate; 0.5% yeast extract; 0.8% arginine; 0.04% histidine), LS medium (1 × S-base; 0.5% glucose; 0.0005% L-tryptophan; 0.01% casein hydrolysate; 0.1% yeast extract; 2.5 mm MgCl2; 0.5 mm CaCl2) and tryptic soy broth (Difco, Franklin Lakes, NJ, USA) or on LB agar plates (Supplementary Table S3). When required antibiotics were added to the media at following concentrations: ampicillin, 100 μg ml−1, and chloramphenicol, 25 μg ml−1. The growth temperature of the cells is indicated (Supplementary Table S3).
Genetic manipulations and protein expression
Cloning, mutagenesis and purification of the Sec translocase components is described in the Supplementary Information.
In vivo genetic complementation assays
The in vivo functionality of mutated ecSecA was investigated using the E. coli BL21.19 secAts strain.39 Cells transformed with the pET5 vector or pET5 carrying the wild-type or mutant E. coli secA gene were grown overnight in LB with ampicillin at 30 °C. The cultures were diluted in LB with ampicillin (OD600 nm=0.01) and further incubated at 30 °C until OD600 nm=0.5. Next, 10 μl of a 10-fold serial dilution (100–10−2) of these cultures was spotted on two LB/ampicillin agar plates. One plate was incubated overnight at the permissive temperature of 30 °C, whereas the second plate was incubated overnight at the non-permissive temperature of 42 °C.
The in vivo effect of mutations in saSecA1 was investigated using the B. subtilis NIG1152 secAts strain40 that can be fully complemented by S. aureus secA1. This strain was transformed with the pHT01 vector or pHT01 carrying the wild-type or mutant S. aureus secA1 gene using the method of Anagnostopoulos and Spizizen41 (MoBiTec, Goettingen, Germany, following instructions of the manufacturer). The cells were grown overnight in LB with chloramphenicol at 30 °C. The cultures were diluted in LB with chloramphenicol (OD600 nm=0.01) and incubated at 30 °C until OD600 nm=0.5. Next, 10 μl of a 10-fold serial dilution (100–10−4) of the cultures was spotted on two LB/chloramphenicol agar plates. One plate was incubated overnight at 30 °C, whereas the second plate was incubated at 42 °C.
In vitro screening of compounds selected by SB-VLS
The 1040 top-scoring molecules were ordered (www.chembridge.com), dissolved in 100% dimethyl sulfoxide (DMSO) and tested in vitro for inhibition of the translocation ATPase activity of ecSecA using a SecA ATPase activity assay, which is a malachite green colorimetric assay that measures the amount of inorganic phosphate formed during ATP hydrolysis.42, 43 The in vitro translocation system needed for this assay was formed by the purified proteins proPhoA(Cys-), ecSecA and inner membrane vesicle (IMV)-embedded ecSecYEG. Compounds (200 μm; 2% final DMSO concentration) were incubated for 10 min at 37 °C with 1 μg ecSecA, 2 μl IMVs (0.1 mg ml−1 total protein) and 0.5 μg proPhoA(Cys-)43, 44 in 50 mm Tris-HCl pH 8.0, 50 mm KCl, 5 mm MgCl2, 0.4 mg ml−1 bovine serum albumin (buffer B). The membrane ATPase activity (1 μg ecSecA and 2 μl IMVs in buffer B) and translocation ATPase activity of ecSecA in the presence of DMSO (2% final concentration) were used as positive and negative controls, respectively (eight wells of each control reaction per plate). ATPase reactions were then started by adding ATP to a final concentration of 1 mm (in a total reaction volume of 50 μl). After 30-min incubation at 37 °C, ATPase reactions were stopped by adding 150 μl Malachite Green reagent (Sigma, St Louis, MO, USA) to the reaction mixture. After 20-min incubation at room temperature, absorbance was measured at 660 nm.
The ecSecA translocation ATPase activity was analyzed by comparing the measured absorbance value per compound well with the plate-averaged negative and positive control wells using the following relationship:

where A660 is the well-specific absorbance value, and μneg and μpos are the plate-averaged negative and positive control values, respectively.
Compounds showing >80% inhibition of the translocation ATPase activity at the tested concentration (200 μm) were defined as hits.
The hit molecules were further studied by titrating them (0–200 μm) in the basal, membrane and translocation ATPase activity assays. The basal and membrane ATP hydrolysis was measured as described above, except that basal ATPase activity was measured in the absence of IMV-embedded ecSecYEG and proPhoA(Cys-), whereas the membrane ATPase activity was measured in the absence of proPhoA(Cys-). Data were expressed as the percentage of the rate of ATP hydrolysis in the absence of inhibitor (see equation above).
Normalized data were plotted versus the logarithm of the inhibitor concentration and IC50 values for inhibition of the SecA translocation, membrane and basal ATPase activity were estimated by fitting the data by nonlinear regression analysis to a sigmoidal inhibition curve with variable slope using GraphPad Prism (La Jolla, CA, USA).
Antimicrobial activity test
The antimicrobial activity of compounds against various bacterial strains was tested using the broth microdilution method. An overnight culture of the tested species was 200 times diluted in LB medium and incubated at 37 °C until the OD600 nm reached 0.3. Next, the cultures were diluted to an OD600 nm of <0.01. Twenty μl of this culture was added to each well of a 96-well microtiter plate containing various compound concentrations ranging from 0 μm to 200 μm (final DMSO concentration of 2.5%), corresponding to ~1 × 104 cells per well in a final volume of 200 μl. The microtiter plates were statically incubated at 37 °C for 20 h. After incubation, the OD600 nm of the cultures was measured and the data were normalized against the control (0 μm compound; 2.5% DMSO). The IC50 values (50% inhibitory concentration) were defined as the concentration of compound that reduced bacterial growth by 50%.
Cytotoxicity assay
The cytotoxicity for human T-lymphoblast (CEM) cells was determined for compounds P97-A9, P87-A4, P91-E9, 17D9, 16F6, P88-B11 and P88-E11. To each well of a 96-well microtiter plate, ~6 × 105 cells were added together with a given amount of the compounds (fivefold serial dilutions ranging from 100 μm to 0 μm). The cells were cultured for 96 h at 37 °C in a humidified CO2-controlled atmosphere. Next, the cells were counted in a Coulter Counter (Coulter Electronics Ltd, Harpenden Herts, UK). IC50 values were determined.
Results and Discussion
Prediction of druggable sites in SecA
To investigate the presence of potential druggable sites on the surface of SecA, an in silico analysis was performed on 11 different SecA structures, using the icmPocketFinder utility as is included in the Molsoft ICM-Pro package45 (Supplementary Table S1). This tool can detect the presence of pockets in proteins and predict their druggability based on physicochemical properties and the size distribution of known drugs.32 Six to 15 potential druggable pockets of varying size, accessibility and location were identified in each SecA structure (Figure 2; those of ecSecA and saSecA1 are shown).
It was not unexpected that various pockets could be identified on the SecA surface as SecA is a multi-domain protein with dynamic domain motions.46 During these conformational changes, potential druggable sites can be revealed more prominently or their exposure to solvent can be altered. The multiple interaction partners of SecA, such as nucleotides, signal peptides and preprotein mature domains,17, 44, 47, 48 may occupy such pockets.
Conservation of druggable pockets
To determine which of the predicted druggable pockets are conserved, a structural comparison across the available SecA structures was performed (Supplementary Figure S1A). Pockets that are situated at the same location in SecA were considered to be conserved. Only two pockets were conserved in all SecAs (Supplementary Figure S1B; pockets 1 and 2), four other pockets were conserved in 10 out of the 11 structures (Supplementary Figure S1B; pocket 3–6) and six were less conserved. Interestingly, most of the predicted pockets were found to be located at interdomain interfaces (Supplementary Figure S1A). As the pockets do not exactly overlap, more general composite sites were defined that comprise one or more conserved pockets.
As >50% of the SecA residues are highly conserved, surface pockets would also be expected to be conserved. The conservation of the surface exposed residues was defined using the ConSurf Server (http://consurf.tau.ac.il/; PDB ID: 2VDA, default settings; data not shown) and is overall consistent with the location of the conserved pockets. One of the pockets, which is present in all investigated SecA structures (Supplementary Figure S1A; pocket 1), is located at the highly conserved nucleotide-binding site, which is also present in other proteins of the DEAD helicase family and is involved in binding the nucleotide.
Functional importance of three selected druggable sites
The functional importance of three conserved druggable sites (Figure 2; Sites I, II and III) on the molecular surface of SecA was examined in vivo. On the basis of the literature data, each of these sites is important for ecSecA function. Whereas Site I lies at the NBD–IRA2 interface and has a role in the binding/hydrolysis of ATP by ecSecA,13, 39 Site II is located at the PBD/C-domain interface and might be important for the regulation of the ATPase activity and for the transmission of conformational changes between the DEAD motor and the specificity domains.9, 19 Site III encompasses the signal peptide-binding site, which is located in the PBD and is, therefore, expected to control preprotein interactions.17, 49
To investigate whether the three sites are important for the function of SecA, various single or multiple amino-acid substitutions were introduced in each of the three predicted druggable sites of both ecSecA and saSecA1 (Supplementary Figure S2A). The effect of the introduced mutations on SecA function was then investigated using an in vivo genetic complementation assay using two temperature-sensitive strains (viable at 30 °C but not at 42 °C), E. coli BL21.19 secAts39 and B. subtilis NIG1152 secAts.40 The functionality of the mutated ecSecAs and saSecA1s was evaluated based on their ability to restore the growth of the secAts strains at 42 °C. Many of the mutated residues in Sites I, II and III compromised the complementation ability of the respective secA genes at 42 °C for both ecSecA and saSecA1 (Supplementary Figure S2B), indicating that each of the three potential druggable sites are essential for SecA function. We focused hereafter on ecSecA alone.
Three of the potential druggable sites (Sites I, II and III), which are conserved among the SecAs of Gram− and Gram+ bacteria, are functionally important. This raises the expectation that any identified SecA inhibitor targeting one of these sites may act as a broad-spectrum antibiotic.
In vitro functionality of mutated ecSecA
The functional importance of the identified sites was further investigated and directly correlated with specific enzymatic activity defects in vitro using mutant derivative ecSecA proteins (Figure 3a). Two derivatives of ecSecA were used to test the functional importance of the predicted druggable sites; ecSecA (L274A/T275A/E276A) that bears a triple mutation in Site II and ecSecA (Y326A)50 that bears a point mutation in Site III. Both mutants fail to stimulate their ATPase activities when inverted IMVs and preprotein are added (Figure 3a, lanes 6 and 9, respectively).
Identified SecA inhibitors. (a) kcat values (turnover number; pmoles Pi per pmol SecA per min) of basal (B), membrane (M) and translocation (T) ATPase activities of WT ecSecA, and ecSecA(L274A/T275A/E276A) and ecSecA(Y326A) PBD mutants. (b) Titration curve of parent molecule P87-A4 and its daughter molecule 17D9. The compounds were titrated (0–200 μm) in the ecSecA basal (Δ), membrane (○) and translocation (♦) ATPase activity assays. Normalized ATPase activity data were plotted versus the logarithm of the compound concentration. (c) Structural information of parent molecule P87-A4 and its daughter molecule 17D9. (d) IC50 values for inhibition of the ecSecA translocation ATPase activity determined both for parent and their corresponding daughter molecules. Only daughter molecules with better inhibition than the parent molecules are shown. PBD, preprotein-binding domain; WT, wild type.
These results demonstrate the functional importance of Sites II and III in the activation of the translocase.
Structure-based VLS
Site III, corresponding to the signal peptide-binding cleft,17 was subsequently tested for its druggability using SB-VLS to screen ~500 000 commercially available small-molecules (ChemBridge compound collection). Site III, one of the most attractive druggable sites with respect to its size and physicochemical properties, has not been targeted with inhibitors previously and may reveal protein–protein interaction inhibitors that are specific to SecA.
The VLS was performed according to an adapted hierarchical consensus screening protocol.33, 34 First, the chemical library was filtered based on the physicochemical properties (ADME/tox profile) of the molecules. Second, the remaining compounds were docked into the predicted druggable site using both rigid body and flexible docking approaches. Docked molecules were scored and ranked using standard scoring functions, which select small-molecules that are most likely to bind the targeted ecSecA-binding site. The top 2000 molecules were analyzed interactively after flexible docking and scoring and 1040 of these were identified as most likely binders.
In vitro screening of top-ranked compounds
The 1040 top-scoring molecules selected via VLS from the initial collection of compounds (Express Pick ChemBridge database at www.chembridge.com, version January 2010) were purchased and tested in vitro for inhibition of the translocation ATPase activity of ecSecA using an established in vitro assay.11, 42, 43 Hit molecules were defined as those that inhibit the translocation ATPase activity by >80% at a final concentration of 200 μm. Using this threshold, 12 hit molecules were identified (Supplementary Table S2, indicated as ‘parent molecules’). This hit ratio of 1.15% falls within the anticipated range and is comparable to previous studies, where we applied SB-VLS only with success rates of 1.251 and 6.4%.52 It should be noted, however, that hit rates are very much dependent on the type of assay used for prioritizing the compounds and thresholds are usually subjective. It is generally accepted, however, that random screens typically range from 0.1 to 0.5% at best.53 Two of the identified hit molecules inhibited not only the translocation ATPase activity but also the basal and membrane ATPase activities (Supplementary Figure S3A) at low concentrations (IC50 values ~3 μm). Moreover, they were toxic to eukaryotic cells (Supplementary Figure S3B and C) and were not studied further.
Ten of the hit compounds inhibited the translocation ATPase activity of ecSecA (Supplementary Figure S4A; Figure 4b, only P87-A4 is shown), without having an effect on the basal and membrane ATPase activity. These compounds were further studied by titrating them (0–200 μm) in the basal, membrane and translocation ATPase activity assay (Supplementary Figure S4A). Normalized ATPase activity data, shown as a percentage of residual ATPase activity, were plotted versus the logarithm of the inhibitor concentration, and IC50 values for inhibition of the SecA translocation ATPase activity were estimated by nonlinear regression analysis. The IC50 values of the 10 tested compounds range between 46 and 134 μm (Figure 3d).
Antibacterial activity of SecA inhibitor compounds. The antibacterial activity of compounds P97-A9, P87-A4, P91-E9, 17D9 and 16F6 was tested against Gram− (E. coli TG1 and BL21, and P. aeruginosa; left) and Gram+ bacteria (S. aureus and B. subtilis; right). The relative growth of the bacteria (OD600 nm normalized to the 2.5% DMSO control) was plotted against the compound concentration. IC50 values for growth inhibition of the bacteria are indicated if relevant. The gray dashed line in the left upper panel is shown as indicative for the growth of the bacteria in the absence of any inhibitor. DMSO, dimethyl sulfoxide.
The identification of SecA inhibitors using the SB-VLS approach showed that this approach is successful in the rational screening for molecules that inhibit SecA. This is further corroborated by previous SB-VLS studies that targeted the ATPase cleft of the helicase motor of both ecSecA and claSecA,26, 28, 29 whereas here we focus on the signal peptide site of ecSecA. Focus on the preprotein cleft may have higher chances of identifying SecA-specific inhibitors, whereas inhibitors targeting the ATPase cleft may have more broad effects as similar clefts are also present in other ATPases.
In vitro screening of daughter molecules
To identify inhibitors with improved properties, we sought to identify structural analogs of the 10 hit compounds. An in silico 2D similarity search against the ChemBridge compound collection was performed employing a simple similarity search at www.hit2lead.com and a Tanimoto similarity cut-off of 0.8. In total, 289 structural analogs were identified and 144 diverse compounds were selected after visual inspection for further analysis in the in vitro translocation ATPase assay.
Of the 20 hit molecules that were identified, 16 compounds were found to inhibit the translocation ATPase activity of ecSecA (IC50 values between 24 and 187 μm; Supplementary Figure S4B), without affecting the basal and membrane ATPase activities. Two daughter molecules inhibited the ATPase activity of ecSecA to a similar or higher extent than their parent molecules (for example, compound P87-A4 compared with 17D9; Figures 3b–d), whereas other daughter molecules showed less inhibition than their parent compound (Supplementary Figure S4C; Supplementary Table S2).
An in silico 2D similarity search can be a very useful tool to identify SecA inhibitors with improved properties, although not all of the identified structural analogs showed better IC50 values than their parent molecules. As the most potent SecA inhibitor has a modest IC50 value of 24 μm, future iterations of the screening of structural analogs will be necessary to identify SecA inhibitors that are functional at lower concentrations.
Antimicrobial activity of the compounds
Next, the in vivo antimicrobial activity against both Gram− and Gram+ bacteria was determined for parent molecules P97-A9, P87-A4 and P91-E9, and their daughter molecules 17D9 and 16F6.
P97-A9 inhibits the growth of the Gram− bacterium P. aeruginosa with an IC50 of 31.8 μm and the Gram+ bacteria S. aureus and B. subtilis with an IC50 of 21.3 and 34.3 μm, respectively (Figure 4). P97-A9 had no effect on the growth of E. coli. At concentrations >50 μm, reduced solubility of P97-A9 in the bacterial growth medium was observed.
Parent molecule P87-A4 displays weak antibacterial activity against E. coli and S. aureus (inhibition by ~35 and ~25% at ~200 μm; Figure 4) but has no effect against P. aeruginosa and B. subtilis.
17D9, the daughter molecule of P87-A4, showed a more pronounced inhibitory effect on the tested bacteria compared with its parent molecule. The growth of E. coli BL21 cells was inhibited with an IC50 value of 198 μm, whereas the growth of E. coli TG1 cells was inhibited by >40% at a compound concentration of 200 μm. At this concentration, S. aureus and B. subtilis growth was inhibited by 40% and 30%, respectively, whereas only a small inhibitory effect was seen on P. aeruginosa (Figure 4).
Parent molecule P91-E9 and 16F6, the daughter molecule of P91-E9, both inhibited the growth of S. aureus (IC50=90–100 μm) but showed weak or no inhibition in the other bacteria tested (Figure 4).
All the compounds had a weak to moderate inhibitory effect on the growth of Gram+ and/or Gram− bacteria. The fact that antibacterial activity against S. aureus is observed for all compounds might be explained by the absence of an outer membrane, although the same inhibitory effect on the Gram+ B. subtilis growth was not observed for all compounds. For P87-A4 and its daughter molecule 17D9, we observed an inhibitory effect against the Gram− bacterium E. coli, indicating that these molecules permeate the outer membrane. To further verify whether SecA is indeed the molecular target in the cases in which growth inhibition was observed, we attempted overexpression experiments in which SecA was synthesized at higher levels due to a tetracycline-regulated promoter. However, these efforts were not successful as even low level overexpression of the secA gene led to growth inhibition in the absence of any inhibitor.10
Cytotoxicity testing of the compounds
The five compounds were also tested on the yeast C. albicans and human T lymphoblasts (CEM cells), which do not possess a SecA homolog. Some of the compounds inhibited the growth of C. albicans (IC50 values >25 μm; Supplementary Figure S4A and B) or the human lymphoblasts (IC50 values >8 μm; Supplementary Figure S4C). This indicates that the compounds may not only have an inhibitory effect on SecA, but also affect other functions or structures on other cell types. The observation that daughter molecule 17D9 is four times less toxic to CEM cells than its parent molecule P87-A4 makes it likely that structural derivatives with reduced toxicity and improved other drug-like properties may be identified in future compound optimization.
Conclusions
An in silico analysis of all available SecA structures revealed the presence of various potential druggable pockets on its surface. Comparison of these pockets showed that six of them are highly conserved among different bacterial species, including both Gram− and Gram+ bacteria. Three selected conserved pockets were found to be very important for the function of both ecSecA and saSecA1, suggesting that they might represent promising targets for the development of SecA inhibitors. Hence, SB-VLS was performed against the signal peptide-binding region of ecSecA to identify small-molecules that block the SecA/preprotein interaction and specifically inhibit Sec-dependent protein translocation. This rational approach, including a second round of ligand-based selection through 2D similarity search, led to the identification of five novel SecA inhibitory molecules with an IC50 value ⩽65 μm for inhibition of the translocation ATPase activity of ecSecA of which the most potent compound had an IC50 value of 24 μm. This compound may be a good starting point for further optimization to design more potent SecA inhibitors. The five selected compounds showed weak antimicrobial effects against Gram+ and/or Gram− bacteria. Besides, they had inhibitory effects on eukaryotic cells. Further optimization of the compounds in terms of potency and selectivity towards SecA will be required.
References
Rodriguez-Rojas, A., Rodriguez-Beltran, J., Couce, A. & Blazquez, J. Antibiotics and antibiotic resistance: a bitter fight against evolution. Int. J. Med. Microbiol. 303, 293–297 (2013).
Nordmann, P., Naas, T., Fortineau, N. & Poirel, L. Superbugs in the coming new decade; multidrug resistance and prospects for treatment of Staphylococcus aureus, Enterococcus spp. and Pseudomonas aeruginosa in 2010. Curr. Opin. Microbiol. 10, 436–440 (2007).
Berdy, J. Thoughts and facts about antibiotics: where we are now and where we are heading. J. Antibiot. (Tokyo) 65, 441 (2012).
Cooper, M. A. & Shlaes, D. Fix the antibiotics pipeline. Nature 472, 32 (2011).
Stephens, C. & Shapiro, L. Bacterial protein secretion—a target for new antibiotics? Chem. Biol. 4, 637–641 (1997).
Economou, A. Sec, drugs and rock'n'roll: antibiotic targeting of bacterial protein translocation. Expert Opin. Ther. Targets 5, 141–153 (2001).
Rao, C. V. S., De Waelheyns, E., Economou, A. & Anne, J. Antibiotic targeting of the bacterial secretory pathway. Biochim. Biophys. Acta 1843, 1762–1783 (2014).
Kusters, I. & Driessen, A. J. SecA, a remarkable nanomachine. Cell. Mol. Life Sci. 68, 2053–2066 (2011).
Chatzi, K. E., Sardis, M. F., Economou, A. & Karamanou, S. SecA-mediated targeting and translocation of secretory proteins. Biochim. Biophys. Acta 1843, 1466–1474 (2014).
Gouridis, G. et al. Quaternary dynamics of the SecA motor drive translocase catalysis. Mol. Cell 52, 655–666 (2013).
Lill, R., Dowhan, W. & Wickner, W. The ATPase activity of SecA is regulated by acidic phospholipids, SecY, and the leader and mature domains of precursor proteins. Cell 60, 271–280 (1990).
Caruthers, J. M. & McKay, D. B. Helicase structure and mechanism. Curr. Opin. Struct. Biol. 12, 123–133 (2002).
Sianidis, G. et al. Cross-talk between catalytic and regulatory elements in a DEAD motor domain is essential for SecA function. EMBO J. 20, 961–970 (2001).
Hunt, J. F. et al. Nucleotide control of interdomain interactions in the conformational reaction cycle of SecA. Science 297, 2018–2026 (2002).
Papanikolau, Y. et al. Structure of dimeric SecA, the Escherichia coli preprotein translocase motor. J. Mol. Biol. 366, 1545–1557 (2007).
Zimmer, J., Nam, Y. & Rapoport, T. A. Structure of a complex of the ATPase SecA and the protein-translocation channel. Nature 455, 936–943 (2008).
Gelis, I. et al. Structural basis for signal-sequence recognition by the translocase motor SecA as determined by NMR. Cell 131, 756–769 (2007).
Sugie, Y. et al. CJ-21,058, a new SecA inhibitor isolated from a fungus. J. Antibiot. (Tokyo) 55, 25–29 (2002).
Vrontou, E., Karamanou, S., Baud, C., Sianidis, G. & Economou, A. Global co-ordination of protein translocation by the SecA IRA1 switch. J. Biol. Chem. 279, 22490–22497 (2004).
Segers, K., Klaassen, H., Economou, A., Chaltin, P. & Anne, J. Development of a high-throughput screening assay for the discovery of small-molecule SecA inhibitors. Anal. Biochem. 413, 90–96 (2011).
Segers, K. & Anne, J. Traffic jam at the bacterial sec translocase: targeting the SecA nanomotor by small-molecule inhibitors. Chem. Biol. 18, 685–698 (2011).
Jang, M. Y., De Jonghe, S., Segers, K., Anne, J. & Herdewijn, P. Synthesis of novel 5-amino-thiazolo[4,5-d]pyrimidines as E. coli and S. aureus SecA inhibitors. Bioorg. Med. Chem. 19, 702–714 (2011).
Huang, Y. J. et al. Fluorescein analogues inhibit SecA ATPase: the first sub-micromolar inhibitor of bacterial protein translocation. ChemMedChem 7, 571–577 (2012).
Parish, C. A. et al. Antisense-guided isolation and structure elucidation of pannomycin, a substituted cis-decalin from Geomyces pannorum. J. Nat. Prod 72, 59–62 (2009).
Alksne, L. E. et al. Identification and analysis of bacterial protein secretion inhibitors utilizing a SecA-LacZ reporter fusion system. Antimicrob. Agents Chemother. 44, 1418–1427 (2000).
Li, M., Huang, Y. J., Tai, P. C. & Wang, B. Discovery of the first SecA inhibitors using structure-based virtual screening. Biochem. Biophys. Res. Commun. 368, 839–845 (2008).
Chen, W. et al. The first low microM SecA inhibitors. Bioorg. Med. Chem. 18, 1617–1625 (2010).
Akula, N., Zheng, H., Han, F. Q. & Wang, N. Discovery of novel SecA inhibitors of Candidatus Liberibacter asiaticus by structure based design. Bioorg. Med. Chem. Lett. 21, 4183–4188 (2011).
Akula, N., Trivedi, P., Han, F. Q. & Wang, N. Identification of small molecule inhibitors against SecA of Candidatus Liberibacter asiaticus by structure based design. Eur. J. Med. Chem. 54, 919–924 (2012).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Schwede, T., Kopp, J., Guex, N. & Peitsch, M. C. SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res. 31, 3381–3385 (2003).
An, J., Totrov, M. & Abagyan, R. Pocketome via comprehensive identification and classification of ligand binding envelopes. Mol. Cell. Proteomics 4, 752–761 (2005).
Miteva, M. A., Lee, W. H., Montes, M. O. & Villoutreix, B. O. Fast structure-based virtual ligand screening combining FRED, DOCK, and Surflex. J. Med. Chem. 48, 6012–6022 (2005).
Du, J., Bleylevens, I. W., Bitorina, A. V., Wichapong, K. & Nicolaes, G. A. Optimization of compound ranking for structure-based virtual ligand screening using an established FRED-Surflex consensus approach. Chem. Biol. Drug Des. 83, 37–51 (2014).
Lagorce, D., Sperandio, O., Galons, H., Miteva, M. A. & Villoutreix, B. O. FAF-Drugs2: free ADME/tox filtering tool to assist drug discovery and chemical biology projects. BMC Bioinformatics 9, 396 (2008).
Hawkins, P. C., Skillman, A. G., Warren, G. L., Ellingson, B. A. & Stahl, M. T. Conformer generation with OMEGA: algorithm and validation using high quality structures from the Protein Databank and Cambridge Structural Database. J. Chem. Inf. Model 50, 572–584 (2010).
McGann, M. FRED pose prediction and virtual screening accuracy. J. Chem. Inf. Model. 51, 578–596 (2011).
Jain, A. N. Surflex: fully automatic flexible molecular docking using a molecular similarity-based search engine. J. Med. Chem. 46, 499–511 (2003).
Mitchell, C. & Oliver, D. Two distinct ATP-binding domains are needed to promote protein export by Escherichia coli SecA ATPase. Mol. Microbiol. 10, 483–497 (1993).
Klein, M., Meens, J. & Freudl, R. Functional characterization of the Staphylococcus carnosus SecA protein in Escherichia coli and Bacillus subtilis secA mutant strains. FEMS Microbiol. Lett. 131, 271–277 (1995).
Anagnostopoulos, C. & Spizizen, J. Requirements for Transformation in Bacillus subtilis. J. Bacteriol. 81, 741–746 (1961).
Karamanou, S. et al. A molecular switch in SecA protein couples ATP hydrolysis to protein translocation. Mol. Microbiol. 34, 1133–1145 (1999).
Gouridis, G., Karamanou, S., Koukaki, M. & Economou, A. In vitro assays to analyze translocation of the model secretory preprotein alkaline phosphatase. Methods Mol. Biol. 619, 157–172 (2010).
Gouridis, G., Karamanou, S., Gelis, I., Kalodimos, C. G. & Economou, A. Signal peptides are allosteric activators of the protein translocase. Nature 462, 363–367 (2009).
Abagyan, R., Totrov, M. & Kuznetsov, D. ICM: a new method for protein modeling and design: applications to docking and structure prediction from the distorted native conformation. J. Comp. Chem. 15, 488–506 (1994).
Sardis, M. F. & Economou, A. SecA: a tale of two protomers. Mol. Microbiol. 76, 1070–1081 (2010).
Lill, R. et al. SecA protein hydrolyzes ATP and is an essential component of the protein translocation ATPase of Escherichia coli. EMBO J. 8, 961–966 (1989).
Cunningham, K. et al. SecA protein, a peripheral protein of the Escherichia coli plasma membrane, is essential for the functional binding and translocation of proOmpA. EMBO J. 8, 955–959 (1989).
Auclair, S. M. et al. Mapping of the signal peptide-binding domain of Escherichia coli SecA using Forster resonance energy transfer. Biochemistry 49, 782–792 (2010).
Kourtz, L. & Oliver, D. Tyr-326 plays a critical role in controlling SecA-preprotein interaction. Mol. Microbiol. 37, 1342–1356 (2000).
Segers, K. et al. Design of protein membrane interaction inhibitors by virtual ligand screening, proof of concept with the C2 domain of factor V. Proc. Natl Acad. Sci. USA 104, 12697–12702 (2007).
Chatzigeorgiou, A. et al. Blocking CD40-TRAF6 signaling is a therapeutic target in obesity-associated insulin resistance. Proc. Natl Acad. Sci. USA 111, 2686–2691 (2014).
Alvarez, J. & Shoichet, B. Virtual Screening in Drug Discovery, CRC Press, (2005).
Acknowledgements
We are grateful to T. D’huys for his contribution to the cloning of the saSecA1 Site I mutants; M. Kulharia for technical assistance during the initial phase of the virtual screening process; L. Van Berckelaer, J. Balzarini and D. Schols for cytotoxicity tests; and P. Chaltin and H. Klaassen for useful discussions. This work was supported by grants SBO IWT 50146; FWO (Krediet aan Navorsers) 1500912N and ESCMID Grant 2012; the Cardiovascular Research Institute Maastricht (to GAFN); the Transnational University Limburg (to GAFN) and KUL-Spa (Onderzoekstoelagen 2013; Bijzonder Onderzoeksfonds; KU Leuven; to AE); RiMembR (Vlaanderen Onderzoeksprojecten; #G0C6814N; FWO; to AE); an Excellence grant (#1473 to AE; Greek General Secretariat of Research).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The Journal of Antibiotics website
Supplementary information
Rights and permissions
About this article
Cite this article
De Waelheyns, E., Segers, K., Sardis, M. et al. Identification of small-molecule inhibitors against SecA by structure-based virtual ligand screening. J Antibiot 68, 666–673 (2015). https://doi.org/10.1038/ja.2015.53
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2015.53
This article is cited by
-
Inhibitors of protein translocation across membranes of the secretory pathway: novel antimicrobial and anticancer agents
Cellular and Molecular Life Sciences (2018)