Abstract
Large sulfur bacteria of the genus Achromatium are exceptional among Bacteria and Archaea as they can accumulate high amounts of internal calcite. Although known for more than 100 years, they remain uncultured, and only freshwater populations have been studied so far. Here we investigate a marine population of calcite-accumulating bacteria that is primarily found at the sediment surface of tide pools in a salt marsh, where high sulfide concentrations meet oversaturated oxygen concentrations during the day. Dynamic sulfur cycling by phototrophic sulfide-oxidizing and heterotrophic sulfate-reducing bacteria co-occurring in these sediments creates a highly sulfidic environment that we propose induces behavioral differences in the Achromatium population compared with reported migration patterns in a low-sulfide environment. Fluctuating intracellular calcium/sulfur ratios at different depths and times of day indicate a biochemical reaction of the salt marsh Achromatium to diurnal changes in sedimentary redox conditions. We correlate this calcite dynamic with new evidence regarding its formation/mobilization and suggest general implications as well as a possible biological function of calcite accumulation in large bacteria in the sediment environment that is governed by gradients. Finally, we propose a new taxonomic classification of the salt marsh Achromatium based on their adaptation to a significantly different habitat than their freshwater relatives, as indicated by their differential behavior as well as phylogenetic distance on 16S ribosomal RNA gene level. In future studies, whole-genome characterization and additional ecophysiological factors could further support the distinctive position of salt marsh Achromatium.
Similar content being viewed by others
Introduction
Large sulfur bacteria of the genus Achromatium constitute the sole genus in the family Achromatiaceae, and were first discovered in 1892 in a freshwater dredging pond in Germany (Schewiakoff, 1892). Their most conspicuous characteristic is the accumulation of numerous large inclusions filling the majority of the cell that were first described as calcium oxalate, giving the type species Achromatium oxaliferum its name, but which were later recognized as colloidal calcite (West and Griffiths, 1913; Bersa, 1920; Head et al., 1996). Recently, cyanobacteria (Gloeobacterales) were found to form amorphous calcium–magnesium–strontium–barium carbonate inclusions (Couradeau et al., 2012); however, the massive accumulation of CaCO3 in Achromatium is still a unique peculiarity in the microbial world (Head et al., 2000; Gray and Head, 2014). The biological function of internal calcite is under investigation, and hypotheses include its usage as buffer, a source of CO2 or a mechanism of buoyancy regulation (Gray, 2006, reviewed in Head et al., 2000).
Calcite-accumulating Achromatium spp. have been detected in freshwater and brackish sediments (West and Griffiths, 1909; Nadson, 1913; Kolkwitz, 1918; Bersa, 1926; Devide, 1952; de Boer et al., 1971; Head et al., 1996; Glöckner et al., 1999; Gray et al., 1999b). The rod-shaped cells range in size from 9 × 15 to 35 × 95 μm, and are motile by slow jerky rolling (Schewiakoff, 1892; Van Niel, 1948; Babenzien, 1991). Although it was hypothesized that they follow the oxycline (de Boer et al., 1971; Gray et al., 1997), most cells remain in the microxic/anoxic layers of the sediments, suggesting a microaerophilic or anaerobic metabolism (Babenzien and Sass, 1996; Head et al., 1996; Gray et al., 1997). Achromatium remains uncultured, and physiological studies are limited to field studies and microcosm experiments (Gray and Head, 2014). Achromatium influences the benthic sulfur cycle by storing internal sulfur from hydrogen sulfide, and by producing sulfate from stored sulfur (Gray et al., 1999a). Each of these partial reactions supports a lithotrophic energy metabolism, and the presence of both suggests a true lithotrophic lifestyle in Achromatium (West and Griffiths, 1913; Gray et al., 1997, 1999a). Achromatium can incorporate organic and inorganic carbon compounds; certain subpopulations fix bicarbonate into biomass whereas others merely deposit it as calcite (Gray et al., 1999a). A strictly lithoautotrophic metabolism remains to be verified (Gray and Head, 1999; Gray et al., 1999a; Head et al., 2000). Alternative electron acceptors to oxygen, such as nitrate and sulfur, have been suggested (Babenzien, 1991; Babenzien and Sass, 1996; Head et al., 2000; Gray et al., 2004).
Two calcite-accumulating species are currently recognized and phylogenetically represented, A. oxaliferum (Schewiakoff, 1892; Head et al., 1996) and ‘Candidatus A. minus’ (Glöckner et al., 1999). A third species, Achromatium volutans, is not phylogenetically characterized, and was described as marine sulfur-containing cells without calcite (Hinze, 1903; Nadson, 1913, 1914; Kolkwitz, 1918; Skuja, 1948; Van Niel, 1948). Cells matching this morphology were recently phylogenetically classified as members of the Beggiatoaceae (Schulz et al., 1999; Salman et al., 2011, 2013), leading to the family Achromatiaceae now exclusively containing members that deposit intracellular calcite (Gray and Head, 2014).
One of the few marine sites where calcite-accumulating Achromatium were detected is the Sippewissett Salt Marsh on Cape Cod, MA, USA (Lackey and Lackey, 1961). In this study, we continue where investigations ceased over 50 years ago, and integrate modern molecular techniques and biogeochemical analyses with a suite of specific methods on single-cell level to gain insights into the ecophysiological niche and adaptation of the Achromatium population to the salt marsh environment.
Materials and methods
Sampling
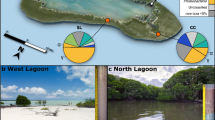
The Sippewissett Salt Marsh is located in Buzzards Bay near Falmouth (Cape Cod, MA, USA) and comprises several saltwater pools at low tide. One pool (N 41°34.548’, W 70°38.388’; Figure 1a) was sampled from June through August in 2012, 2013 and 2014 during low tide at mid-night and mid-day (details in Supplementary Information). All push cores and syringe cores were taken within an area of ca. 1 m2 in the same pool. Push cores contained 2–5 mm of supernatant pool water, and remained open.
Sampling site and characteristics of Achromatium cells. (a) Tide pool in Sippewissett salt marsh. (b) Sediment core taken in tide pool. (c) Achromatium cells concentrated in the center of petri dish (arrow) after rotation. Their white appearance is because of internal calcite and sulfur. (d) Phase contrast micrograph of Achromatium cells shows refractive inclusions. Cells differ in size, and some are dividing. (e) Differential interference contrast image: large inclusions dominate the cell interior, and small inclusions fill the interstitial space. (f) The 4',6-diamidino-2-phenylindole (DAPI) and (g) fluorescein isothiocyanate (FITC) staining and confocal imaging of native, wet cells shows the thin cytoplasm in the interstitial space between inclusions and an even distribution of DNA therein. (h) Dividing cells with a thin constriction ‘tube’ between two spherical daughter cells. (i) Dividing cells with a flat division plane between two semispherical daughter cells.
Microsensor measurements
Microsensors OX-100, H2S-100 and pH-100 were purchased from Unisense (Aarhus, Denmark), and calibrated according to the manufacturer’s instructions. Using the software Sensor Trace Suite and a computer-controlled micromanipulator (Unisense), downcore profiles were measured starting at the surface of the overlying water that was not agitated to simulate near-in situ conditions. Total dissolved sulfide concentrations were calculated from sulfide and pH profiles using equation Stot= [H2S] × (1+K1/[H3O+]), with pK1=6.554 (Millero et al., 1988).
Three daytime and nighttime cores were sampled on six different dates. Each chemical gradient was profiled at least three times per core to calculate averages. Daytime cores were profiled immediately after return from the field, illuminated at 275 μmol photons m−2 s−1. Night time cores were kept dark for ca. 6 h before profiled in the dark.
Cell counts
Three daytime and nighttime syringe cores were taken on six different days with cutoff 60 ml syringes. Sediment was collected in 2 mm intervals, corresponding to 1 ml sample volume. Samples were filtered through a 20 μm pore mesh, and Achromatium cells in the flow through were concentrated in a petri dish by rotation (Figure 1c). Cells were counted using the stereomicroscope Discovery V20 (Zeiss, Oberkochen, Germany).
Staining and fluorescence microscopy
For 4',6-diamidino-2-phenylindole (DAPI) and fluorescein isothiocyanate (FITC) staining, cells were fixed in 2% formaldehyde, washed and incubated for 1 h at 22 °C with DAPI (1 μg ml−1) and FITC (0.1 mg ml−1). For staining with Calcium Orange-5N AM and Calcium Green-1 (Molecular Probes, Eugene, OR, USA), unfixed washed cells were incubated in 2 μM dye and 0.04% Pluronic for 1 h at 22 °C, and washed twice. All stained cells were mounted in a drop of marsh water on a microscope slide surrounded by tape to avoid crushing the cells with the cover slip, and viewed with a LSM 710 laser scanning microscope (Zeiss) using the 350, 488, 541 or 561 and 506 nm laser, respectively.
Raman spectroscopy
Cells were washed in MilliQ water (EMD Millipore, Darmstadt, Germany) and mounted as described above. Individual cells were analyzed using a Horiba Jobin-Yvon (Horiba Scientific, Edison, NJ, USA) LabRAM HR confocal Raman spectrometer with a 17 mW, 633 nm He-Ne laser to analyze the molecular composition of inclusions inside Achromatium. The system features a hard coupled Olympus (Center Valley, PA, USA) BX41 microscope, a 800 mm focal length astigmatic flat-field spectrograph and a multichannel air-cooled (−70 °C) CCD detector with a 1024 × 256 pixels front illuminated chip. Individual spectra were recorded using a 100 × objective, a 600 grooves per mm grating and a 300 μm confocal hole diameter for 5 s and 3 accumulations between 100 and 1200 cm−1. The background was subtracted with the LabSpec 5 software (Horiba Scientific) and spectra were compared with the RRUFF Project’s database for reference using the Crystal Sleuth (Laetsch and Downs, 2006) software application.
Scanning electron microscope/energy-dispersive X-ray spectrometer
Washed cells were placed on specimen stubs in a drop of MilliQ water, dried at 37 °C, rinsed with MilliQ and dried again. Cells were imaged with a Hitachi TM3030 (Clarksburg, MD, USA) scanning electron microscope. Elemental analysis of a selected area was performed with an integrated energy-dispersive X-ray spectrometer (Bruker Quantax 70, Harvard, MA, USA) with 15 keV beam energy, and a silicon drift detector with a 30 mm2 area. Elements were quantified with the software Bruker Quantax 70.
The beam penetration is ca. 5 μm into the outer shell of the Achromatium cell (details in Supplementary Information). Net Ca and S signals per image thus give a relative indication of the amount of S versus Ca within the ~5 μm-thick penetration depth. The sum of net amounts of Ca and S were normalized to 100%; average percentages of the individual compounds and s.d. values were calculated across all images from the same sediment depth and time of day. The calcium/sulfur (Ca/S) ratios were determined from these values.
16S rRNA gene analysis
A single, washed Achromatium cell was subjected to whole-genome multiple displacement amplification using the illustra Genomi Phi Amplification kit (GE Healthcare Life Sciences, Pittsburgh, PA, USA). The 16S ribosomal RNA (rRNA) gene of the amplicon was retrieved using 1 μ M final concentration of universal primers 27F and 1492R (Lane, 1991), 1 × GoTaqFlexi buffer (Promega, Madison, WI, USA), 2.5 mM MgCl2, 0.2 mM of each dNTP and 0.25 U GoTaq DNA polymerase. The PCR protocol was: 94 °C for 2 min; 25 cycles of 94 °C for 30 s, 50 °C for 30 s, 72 °C for 1 min; and 72 °C for 5 min. The PCR product was cloned using the TopoTA cloning kit following the manufacturer’s instructions (Life Technologies, Grand Island, NY, USA), sequenced via Sanger sequencing and submitted to the EMBL/EBI/DDBJ databases under accession number HG934343.
For bulk community analysis, the upper 2 cm of the Achromatium-rich sediment core were filtered through a 20 μm mesh before 4 ml of settled particles were pelleted via centrifugation and transferred to bead tubes of the PowerSoil DNA Isolation kit (MoBio, Carlsbad, CA, USA). In parallel, ~200 Achromatium cells were washed and used for DNA extraction with a standard phenol/chloroform extraction protocol (Sambrook and Russel, 2006). The amplification reaction of 16S rRNA genes contained 10 to 50 ng DNA of sediment community or Achromatium cell mixture, 1 × Phusion HF MasterMix (New England Biolabs, Ipswich, MA, USA), 8% dimethyl sulfoxide, 25 μM universal reverse primer 907R and 6.25 μM barcoded universal forward primer 515F. The PCR protocol was: 98 °C for 1 min; 10 cycles of 98 °C for 5 s, 68 °C for 10 s, 72 °C for 7 s; 12 cycles of 98 °C for 5 s, 58 °C for 10 s, 72 °C for 7 s; 10 cycles of 98 °C for 5 s and 72 °C for 21 s. PCR products were sequenced using 454/Roche’s pyrosequencing platform (Branford, CT, USA), analyzed with QIIME (Caporaso et al., 2010) where taxonomic classification used the Ribosomal Database Project as reference (Cole et al., 2005), and submitted to the EMBL/EBI/DDBJ databases under the Bio Project ID PRJNA251325.
Tree reconstruction
The 16S rRNA gene sequences between nucleotide positions 252–1487 (Escherichia coli numbering) of the Achromatiaceae and related Gammaproteobacteria were used for maximum likelihood tree reconstruction with the Silva database release 102 (Pruesse et al., 2007; Quast et al., 2013) and the Arb software package (Ludwig et al., 2004). Bootstraps analysis included 1000 runs. Partial sequences from pyrosequencing were added to the tree with parsimony criteria without allowing changes in the tree topology. Branches with <50% confidence were manually revised by creating multifurcations.
Catalyzed reporter deposition/fluorescence in situ hybridization
Specific probe design was done in Arb (Ludwig et al., 2004) and tested for specificity using RDP and EMBL databases. The horseradish peroxidase-conjugated probe Achr819, 5'-CCTAACAGCTAGTCGACA-3', was purchased from Biomers (Ulm, Germany). Washed Achromatium cells were fixed in 2% formaldehyde, dried on poly-L-lysine-coated glass slides (Thermo Scientific, Waltham, MA, USA) and sprayed with 0.4% agarose. Hybridization was performed following published guidelines (Pernthaler et al., 2002) using horseradish peroxidase-conjugated probes EUB338 (Amann et al., 1990) and GAM42a with unconjugated competitor BET42a (Manz et al., 1992) as positive, and conjugated NON338 (Wallner et al., 1993) as negative control. Probe Achr819 was used at 100 ng μl−1 with helpers (5'-CCCAGGCGGTCGACTTA-3', and 5'-CGCCACTAARCCCTTAR-3'; Fuchs et al., 2000). Hybridization occurred at up to 25% formamide.
Results
Description of the site
The Sippewissett Salt Marsh is influenced by the tides and contains several pools at low tide surrounded by dense growth of cordgrass (Spartina alterniflora) (Figure 1a). At the bottom of the pools, a brown, liquid suspension of decomposing phytodetritus (ca. 8 cm deep) was lying on top of sandy ground (Figure 1b). The phytodetritus layer contained high densities of conspicuous large pink aggregates composed of purple sulfur bacteria and sulfate-reducing bacteria (Seitz et al., 1993; Wilbanks et al., 2014). The overlying water measured ∼1–2 cm during low tide and ca. 50 cm during high tide. The salinity fluctuated between 33 PSU at night and 38 PSU during the day in both the overlying water and sediment.
Geochemistry
The distribution of oxygen, hydrogen sulfide and pH varied among the cores investigated at the same times of day, but showed common trends (Figure 2). In day cores, photosynthesis created oversaturated oxygen concentrations in the overlying water with peak concentrations close to the sediment surface (Figure 2a). When profiled outdoors in full sunlight, oversaturation could reach ~1000 μM (data not shown). Within the first 2 mm of the sediment, the high oxygen concentrations declined steeply toward the detection limit. Oxygen concentrations in the overlying water of night cores gradually decreased toward the sediment surface, and met the detection limit at the sediment/water interface.
Microsensor profiles of dissolved oxygen (a), sulfide (b) and pH (c) measured in phytodetritus sediment cores obtained from a tidal pond in Sippewissett salt marsh. Cell counts (d) were determined from cells in parallel cores from the same location; intracellular Ca/S ratios (e) were determined in cells retrieved from cores after microprofiling. Open symbols represent data collected during the day and filled symbols data from the night. Microsensor data points represent the average values of three profiles measured in a given core. Error bars in (d) denote s.d. of values across cores at same time points.
Concentrations of total sulfide in the sediment fluctuated drastically between 0.5 and 2.5 mM (Figure 2b). During the night, the sulfide gradients commonly reached the sediment surface, whereas in day cores sulfide was depleted at 2, 3 and 5 mm sediment depth.
The pH of night cores dropped from ca. 7.5 in the overlying water to ca. 7.0–7.3 in the sediment (Figure 2c). During the day, the profiles fluctuated considerably, with the core showing highest photosynthetic activity (core 3) having highest pH values in the overlying water (pH <8.8) resulting from CO2 consumption during photosynthesis (Revsbech et al., 1983; Des Marais et al., 1989). In the sediment, pH decreased to pH 6–7 as typical for anaerobic carbonate-buffered sediments (Soetaert et al., 2007).
The marsh-dwelling Achromatium
In the upper 3 cm of the tide pool sediment we observed large white bacteria that were rod-shaped or spherical (Figure 1d). The average size was 20 × 26 μm (n=135), with the smallest cell measuring 10 × 17 μm and the largest 27 × 38 μm. Division stages included cells with a thin constriction ‘tube’ between two spherical cells (Figures 1d and h), and cells with a flat division plane between two semispherical cells (Figure 1i and Supplementary Figure S1A).
Some cells seemed to have an external sheath with attached rod-shaped (Supplementary Figure S1B) or filamentous (Supplementary Figure S1C) bacteria that were previously interpreted as flagella for cellular movement (de Boer et al., 1971; La Rivière and Schmid, 1989). However, as was noticed before (Head et al., 1996) and here, most cells lacked this coating and/or putative epibionts while still exhibiting a rolling motility, leaving the function of the coating open to interpretation (Head et al., 1996, 2000).
Most marsh-dwelling Achromatium cells occurred within the first centimeter of the phytodetritus sediment (Figure 2d); cell numbers were as high as 1023 cells per ml sediment. During the day, the cells were concentrated within the upper 4 mm, whereas during the night they were more dispersed within the surficial 1.25 cm of the sediment and the sediment/water interface (Figure 2d).
Cell inclusions
Light microscopy showed that cells were filled with large globules (4–5 μm in diameter), occupying most of the cell interior, and with small granules (1–1.5 μm) in the interstitial space (Figures 1d and e). Confocal Raman spectroscopy revealed that the large inclusions consisted of calcite (CaCO3), and that native sulfur (S8) filled the interstitial space (Figure 3a). The large calcite granules disintegrated when cells were exposed to acidified seawater (pH ⩽5), leaving small inclusions in most cases (Figure 3d.2). Raman spectroscopy on acid-treated cells gave in some but not all cases the specific signal for S8 sulfur (data not shown). When treated with 80% ethanol or pure methanol the small inclusions disintegrated (Figure 3d.3). Staining with the low-affinity dye Calcium Orange-5N, with a detection range between 1 and 50 mM free Ca2+ (Kd=20 μM) (Zhao et al., 1996; Naraghi, 1997), revealed a third type of inclusion. These small, Ca2+-enriched bodies were also situated in the interstitial space between the large calcite inclusions (Figure 3e), and still stained positive in acid-treated cells (data not shown), suggesting that they are not calcite. Applying the high-affinity dye Calcium Green-1, which detects free Ca2+ between 0.04 and 40 μM (Kd=190 nM) (Eberhard and Erne, 1991; Lakowicz et al., 1992; Sanders et al., 1994), also stained the small Ca2+-enriched bodies, but in addition stained the periphery of the large calcite inclusions (Figure 3f). Fluorescein isothiocyanate stained the cytoplasm as thin layers between the granules (Figure 1g and Supplementary Figure S1A). A single, large internal vacuole, which is typical for large sulfur bacteria of the sister family Beggiatoaceae (Maier and Gallardo, 1984; Nelson et al., 1989; Salman et al., 2013), was never observed. DAPI-staining revealed an even distribution of DNA across the cytoplasm (Figure 1f) that formed numerous nucleoid-like aggregates in dried cells (Figure 3g.2).
Properties of marsh Achromatium cells. (a) Representative Raman analysis of intracellular molecules; comparison of measured spectra with standards from the RRUFF reference database (a.1); distribution map of signals inside single cell (a.2). (b, c) Scanning electron microscope/energy-dispersive X-ray spectrometer (SEM/EDS) elemental analysis of a representative cell from sediment depth 1 cm (b) and from 3 cm (c), showing the SEM image (b.1 and c.1) and the localization of calcium (b.2 and c.2) and sulfur (b.3 and c.3). (d) Differential dissolution of intracellular inclusions (here not showing the same cell); native cell (d.1), cell treated with mild acid (d.2) and treatment with methanol after acid step (d.3). (e) Calcium Orange-5N signal (e.1), transmitted light image (e.2). (f) Calcium Green-1 signal (f.1), transmitted light image (f.2). Catalyzed reporter deposition/fluorescence in situ hybridization (CARD/FISH) analysis; hybridization with probe Achr819 (g.1), counterstaining with 4',6-diamidino-2-phenylindole (DAPI; g.2). Scales: (a–c) 5 μm, (d–g) 10 μm.
Scanning electron microscope imaging and light microscopy of Achromatium cells revealed a decrease in visible large calcite granules with depth. Cells from the sediment surface were usually tightly filled (Figure 3b.1), whereas cells at 3–4 cm depth usually contained only small inclusions (Figure 3c.1). This trend was also observed in freshwater Achromatium populations (Lauterborn, 1915; Babenzien, 1991). Energy-dispersive X-ray spectrometer elemental analysis of individual cells showed a successive increase in sulfur deposition with depth (Figures 2e, 3b and c and Supplementary Table S1). This trend was particularly pronounced in cells collected during the day where measurable calcium dropped from 99% to 60%, and sulfur increased from 0.7% to 40% with depth (Supplementary Table S1). At night, cells had a generally lower Ca/S ratio but showed the same downcore trend (Figure 2e); we detected 88% Ca versus 12% S at 1 cm and 2% Ca versus 98% S at 3 cm.
As previously described, extended exposure in oxic sea water or on a microscope slide caused all types of inclusions to fade, which was attributed to sulfur oxidation to sulfate with resulting acidification of the medium and dissolution of the calcite inclusions (West and Griffiths, 1913; Starr and Skerman, 1965).
Phylogenetic identification
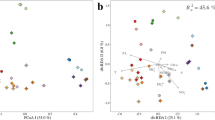
The full-length 16S rRNA gene sequence from a single, calcite-accumulating cell revealed its affiliation to the Achromatium genus (91.2–93.8% sequence identity), and was most similar to cluster A (Figure 4a), one of the two previously assigned Achromatium clusters (Glöckner et al., 1999; Gray et al., 1999b). The new sequence lacked helix 38 in the V6 region (Figure 4b and Supplementary Figure S2), which is a designated feature of cluster A Achromatium phylotypes (Glöckner et al., 1999; Gray et al., 1999b).
Phylogenetic tree of the Achromatiaceae and related Gammaproteobacteria based on nearly full-length 16S rRNA gene sequences (a). Helix 38 is missing in the V6 region of Sippewissett Achromatium (positions 1024 to 1036, E. coli numbering) (b), as in other phylotypes affiliated with cluster A (Glöckner et al., 1999; Gray et al., 1999b).
Pyrosequencing of partial 16S rRNA genes from a mix of ca. 200 hand-picked Achromatium cells and associated bacteria yielded 8666 reads, of which 151 were 99.5–100% identical and affiliated with the Achromatiaceae. They were 95% identical to the full-length marsh Achromatium sequence. Together, these sequences formed a distinct monophyletic cluster of salt marsh Achromatium (Figure 4a).
In situ hybridization with a probe designed specifically for the salt marsh Achromatium branch confirmed the presence of the marsh Achromatium rRNA in 47±8% of calcite-accumulating cells collected from the Sippewissett Salt Marsh (n=632; Figure 3g). Hybridizations with the general bacterial probe EUB338 (n=310) and with the nonsense probe Non338 (n=306) stained 85±13% and 0% of all calcite-accumulating cells, respectively.
Dominant phylotypes in the sequencing result of washed Achromatium cells were Alteromonadales (16.4%), Oceanospirillales (12.9%), Flavobacteria (11.1%), Vibrionales (3.88%), and Desulfovibrionales (0.6%) (Supplementary Table S2). The abundances of these phylotypes in the sediment libraries were much lower (between 2.2% and 0.2%, Supplementary Table S2), and may represent putative epibionts of Achromatium.
Community analysis
Partial 16S rRNA gene sequencing of bulk DNA from the sediment-associated community in the upper 2 cm revealed a dominance of bacteria associated with sulfur cycling in marine environments (Supplementary Table S2). Both sediment layers were dominated by Proteobacteria (54.7% and 43.3%), most significantly Gammaproteobacteria (32.3% and 20.5%), including anoxygenic phototrophs of the Chromatiaceae, and Deltaproteobacteria (12.4% and 16.4%), including strictly anaerobic sulfate reducers of the Desulfobacteraceae.
Discussion
Identification of Achromatium
The population of calcite-accumulating sulfur bacteria in the Sippewissett Salt Marsh phylogenetically affiliates with the genus Achromatium, forming a novel monophyletic group within the family Achromatiaceae (Figure 4). These marsh-dwelling Achromatium share typical morphological characteristics with freshwater Achromatium, including cell size, storage compounds, motility and sheathing (Schewiakoff, 1892; Van Niel, 1948; de Boer et al., 1971; Head et al., 1996; Glöckner et al., 1999; Gray and Head, 1999; Head et al., 2000). Consequently, the ability to accumulate calcite is not restricted to freshwater or brackish Achromatiaceae. Additional data from this family could lead to a reclassification of the species A. oxaliferum in the future as this species designation currently extends to both clusters A and B (Gray and Head, 2014), whereas cluster B also contains the candidate species ‘Achromatium minus’ (Glöckner et al., 1999; Gray et al., 1999b). Based on the level of 16S rRNA gene divergence of marsh Achromatium to all other members of the family (>6% 16S rRNA gene sequence difference), and their stable adaptation to a substantially different habitat, we propose a new candidatus designation for the marsh-dwelling Achromatium 16 S rRNA genes retrieved here, and suggest ‘Candidatus Achromatium palustre’ (Figure 4; according to Murray and Schleifer, 1994). However, only 47% of collected Achromatium cells hybridized with the probe designed specifically for this cluster, whereas nearly all tested cells stained with the general bacterial probe (Figure 3g). We conclude that approximately half of the Achromatium population in Sippewissett is composed of the new candidate group, and the remainder of the cells belongs to one or several other Achromatium phylotype(s).
Previous studies showed that natural communities of freshwater Achromatium contain morphologically and genetically distinct subpopulations (7–9% 16S rRNA gene sequence divergence) that can be linked to their position in the sediment and their specific adaptation to location-specific redox conditions (Glöckner et al., 1999; Gray et al., 1999a, b, 2004). Sippewissett Achromatium 16S rRNA gene sequences differ by up to 4.8%; however, specific niche differentiation of putative subpopulations along salinity or redox gradients require further investigation.
The marsh habitat and ecophysiological implications for Achromatium
In some aspects, the environmental niche of Sippewissett Achromatium resembles settings described for freshwater populations; that is, opposing gradients of sulfide and oxygen, oversaturation of oxygen during the day at the shallow sediment/water interface and possible movement of Achromatium between aerobic and anaerobic zones (Babenzien, 1991; Babenzien and Sass, 1996; Gray et al., 1997, 1999b). In other aspects, the salt marsh creates a substantially different habitat. High ambient seawater sulfate concentrations of ~28 mM fuel extensive sulfate reduction, leading to high sulfide concentrations in the sediments, that is, several millimoles per liter in Sippewissett (Figure 2b). The very active and dynamic sulfur cycling in Sippewissett is reflected by overall high total sulfide concentrations and high core-to-core fluctuations of total sulfide (Figure 2b), as well as by the high abundance of sulfur-cycling microorganisms (Wilbanks et al., 2014, Supplementary Table S2). This sulfide pool provides a substantial reservoir of electron donor for the sulfide-oxidizing community, including pink consortia (Seitz et al., 1993; Wilbanks et al., 2014) and Achromatium, while also being toxic at certain concentrations. Sippewissett Achromatium take part in the sedimentary sulfur cycle by storing zerovalent sulfur in the form of S8 sulfur (Figure 3a) that is most likely produced during the oxidation of sulfide in anoxic regions of the sediment, as the amount of intracellular sulfur increases with depth (Figures 2e and 3c). Closer to the surface, and especially when oxygen is produced here during the day, Ca/S ratios per cell increase, with cells being nearly depleted in sulfur in highly oxic surface layers (Figures 2e and 3b). Similar findings were reported for freshwater Achromatium (Gray et al., 1999a). These results indicate a physiological link between calcite globules and oxygen. Furthermore, salt marsh Achromatium may have a high oxygen tolerance, as most cells were situated in layers that were significantly elevated in oxygen, and free of sulfide (Figure 2). Complete oxidation of sulfur to sulfate in Achromatium may thus occur in fully oxic layers in the salt marsh, and in hypoxic layers in freshwater (Gray et al., 1997).
In anaerobic regions of the sediment, nitrate can be potentially used as electron acceptor by freshwater Achromatium (Gray et al., 1997, 2004; Gray and Head, 1999). Respiration of intracellular sulfur was also proposed, but not tested (Head et al., 2000), and was described as an anaerobic survival strategy for two Beggiatoa strains (Nelson and Castenholz, 1981; Schwedt et al., 2011). A third potential reservoir of electron acceptor in anaerobic salt marsh sediments could be solid-phase and porewater zerovalent sulfur (ca. 500 μM S0, Zopfi et al., 2004).
During daytime/illumination, freshwater Achromatium migrate downward into the sediment column; they are expected to follow the retreating sulfide horizon and/or to avoid elevated oxygen concentrations (Gray et al., 1997; Head et al., 2000). Population migration dynamics in marsh Achromatium, however, show persistence in the top 4 mm during the day, suggesting oxygen tolerance or even an aerobic metabolism. Here, increasing oxygen penetration and a retreating sulfide horizon during the day could provide a break from high sulfide exposure at night, and present opportunity to aerobically oxidize intracellular sulfur to sulfate. Consistent with this interpretation, cells are drastically depleted in sulfur toward the surface during the day, whereas cells sampled at night contain significantly more sulfur (Figure 2e). The physiological balancing act between the supply of the electron donor sulfide and lethal levels in the marsh sediments might also be reflected in the relatively low cell densities of Achromatium in Sippewissett (102–103 cells per ml) as compared with freshwater populations with 103–105 cells per ml (Head et al., 1996, Gray et al., 1997). The observation that Achromatium populations dynamically store and release intracellular sulfur and calcite as well as have the ability to migrate dramatically in space within a few hours (Gray et al., 1997, 1999a) can now be extended to salt marsh populations.
The formation of calcite
Transmission electron microscopy images of thin sections of calcite granules in Achromatium show an electron-dense central nucleation point, a laminated structure of granules and a possible membrane surrounding each granule (Head et al., 2000). Internal calcite is suggested to be colloidal instead of crystalline (West and Griffiths, 1913), and X-ray diffraction data (Head et al., 1995) imply that deposition is under strict biological control (Gray, 2006). Labeling experiments demonstrate that Achromatium accumulate new intracellular calcite within a few hours (Head et al., 1996; Gray et al., 1999a). Altogether, calcite formation is a rapid and dynamic biological process (Gray, 2006).
In this study, specific staining of free Ca2+ provides evidence that apart from large calcite and small sulfur compartments, Achromatium may contain a third, calcium-rich compartment (Figures 3e and f). Cells from greater sediment depths show no visual evidence of large calcite inclusions, but yield calcium signals in energy-dispersive X-ray spectrometer, possibly resulting from the novel, small-scale calcium inclusions reported here (Figure 3c.2 and Supplementary Table S1). We hypothesize that these provide reservoirs of Ca2+ ions to allow controlled precipitation of calcite. If they resemble early developmental stages of the large calcite-containing granules, they may be surrounded by a membrane (de Boer et al., 1971; Head et al., 2000) that could enable a directed transport of CO32−/H+/OH− to specifically localize calcite formation or dissolution as a consequence of pH change. It was suggested that Achromatium has active Ca2+ transport mechanisms (Head et al., 2000), but energetically expensive transmembrane transport could be minimized by maintaining an intracellular Ca2+ reservoir. Staining free Ca2+ at the outermost rim of large calcite inclusions (Figure 3f) implies that calcite is forming and dissolving at the periphery of inclusions, as suggested before (Head et al., 2000; Gray, 2006).
Previous theories about the biological function of intracellular calcite in Achromatium were based on biogeochemical characteristics of freshwater habitats. We address these hypotheses in the Supplementary Information, but elaborate here the hypothesis we rank as most plausible, that is, calcite is an effective buffer and can act against locally fluctuating pH conditions during sulfide/sulfur oxidation (modified from La Rivière and Schmid, 1992). This hypothesis may be valid for freshwater and marine Achromatium, and takes into account the extraordinary cell biology of large bacteria. Other large sulfur-oxidizing bacteria (for example, Thiomargarita) contain intracellular vacuoles to increase surface-to-volume ratios and overcome diffusion limitation (Schulz and Jørgensen, 2001). In these bacteria, local intracellular alkalinity/acidification during sulfide/sulfur oxidation may be avoided by discharging solutes efficiently from the <2 μm thin cytoplasm (based on Schulz and Jørgensen, 2001). Protons generated in the cytoplasm could also be pumped into the vacuole for use in antiporter-mediated nitrate accumulation (Mussmann et al., 2007). Accordingly, limited H+/OH− exchange in large, unvacuolated Achromatium cells would lead to drastic pH changes in the cytoplasm during sulfide/sulfur oxidation, but strongly buffering storage compounds such as calcite would provide an effective physiological response. We propose that calcite accumulation is directly coupled to the oxidation of sulfide to elemental sulfur, whereas calcite dissolution is a consequence of sulfur oxidation to sulfate (Supplementary Figure S3). The protons to be consumed in the first half reaction (Supplementary Equations S1–S3) could be supplied by bicarbonate with the consequence of calcite precipitation from CO32− and Ca2+ (Supplementary Equation S4 and Supplementary Figure S3, bottom). In the second half reaction, protons are released (Supplementary Equations S5–S7) that dissolve calcite, and carbonate can buffer the acidity-forming bicarbonate (Supplementary Equation S8 and Supplementary Figure S3, top). This implies that Achromatium require calcite only when producing protons from sulfur oxidation, which is the case during the day in oxygenated layers of the sediment, and could thus explain why there is such a drastic increase in visible large CaCO3 inclusions in cells collected at the surface during the day (Figures 2e and 3b).
Measurements of total Ca in freshwater sediments showed a positive correlation with Achromatium cell numbers (Head et al., 2000). If freshwater Achromatium also contain the Ca2+-rich reservoirs reported here, these intracellular Ca measurements would include CaCO3 and Ca2+. In this light, a given cell may always contain more or less the same amount of calcium that oscillates between the solid-state CaCO3 form and the dissolved Ca2+ ion. The sulfur content, however, may change drastically when the cell migrates between different redox conditions, where it may, for example, perform pH-neutral sulfur respiration. In conclusion, calcite may have similar dynamics and/or may serve similar functions in freshwater and marine Achromatium populations.
Conclusion
The salt marsh habitat creates a substantially different geochemical realm for Achromatium—highly sulfidic sediment is opposed by oxygen supersaturation at the surface during the day, providing a temporary sulfide-free zone for putative sulfide-stressed Achromatium. Salt marsh Achromatium persist at this oxic layer, in contrast to freshwater Achromatium. Calcite in these large, diffusion-limited bacteria could function as internal buffer to regulate the surplus or deficit of H+ and OH− in dynamic habitats such as sediments. Calcite precipitation may be initiated in intracellular compartments that contain free Ca2+ ions. Future studies on adaptation and intracellular calcite dynamics in uncultured marine and freshwater Achromatium populations may include analyses and comparisons of metagenomes, -proteomes and -transcriptomes.
Accession codes
References
Amann R, Binder BJ, Olson RJ, Chrisholm SW, Devereux R, Stahl DA . (1990). Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl Environ Microbiol 56: 1919–1925.
Babenzien HD . (1991). Achromatium oxaliferum and its ecological niche. Zentralbl Mikrobiol 146: 41–49.
Babenzien HD, Sass H . (1996). The sediment-water interface - habitat of the unusual bacterium Achromatium oxaliferum. Arch Hydrobiol Spec Issues Adv Limnol 48: 247–251.
Bersa E . (1920) Über das Vorkommen von kohlensaurem Kalk in einer Gruppe von Schwefelbakterien. Sitzungsbericht Akademie der Wissenschaften, mathematisch-naturwissenschaftliche Klasse, I Abteilung: Wien.
Bersa E . (1926). Neue kalkführende Schwefelbakterien. Planta 24: 373–379.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al. (2010). Qiime allows analysis of high-throughput community sequencing data. Nat Methods 7: 335–336.
Cole JR, Chai B, Farris RJ, Wang Q, Kulam SA, McGarrell DM et al. (2005). The Ribosomal Database Project (RDP-II): sequences and tools for high-throughput rRNA analysis. Nucl Acids Res 33: D294–D296.
Couradeau E, Benzerara K, Gerard E, Moreira D, Bernard S, Brown GE Jr et al. (2012). An early-branching microbialite cyanobacterium forms intracellular carbonates. Science 336: 459–462.
deBoer WE, La Riviere JWM, Schmidt K . (1971). Some properties of Achromatium oxaliferum. Antonie van Leeuwenhoek 37: 553–563.
Des Marais DJ, Cohen Y, Nguyen H, Cheatham M, Cheatham T, Munoz E . (1989). Carbon isotopic trends in the hypersaline ponds and microbial mats at Guerrero Negro, Baja California Sur, Mexico: implications for precambrian stromatolites. In: Cohen Y, Rosenberg E (eds), Microbial Mats: Physiological Ecology of Benthic Microbial Communities. American Society for Microbiology: Washington DC, pp 191–203.
Devide Z . (1952). Zwei neue farblose Schwefelbakterien: Thiogloea ruttneri n. gen., n. sp. und Thiogloea ragusina n. sp. Hydrobiologia 14: 446–455.
Eberhard M, Erne P . (1991). Calcium binding to fluorescent indicators: calcium green, calcium orange and calcium crimson. Biochem Biophys Res Commun 180: 209–215.
Fuchs BM, Glockner FO, Wulf J, Amann R . (2000). Unlabeled helper oligonucletides increase the accessibility to 16S rRNA of fluorescently labeled oligonucleotide probes. Appl Environ Microbiol 66: 3603–3607.
Glöckner FO, Babenzien HD, Wulf J, Amann R . (1999). Phylogeny and diversity of Achromatium oxaliferum. Syst Appl Microbiol 22: 28–38.
Gray ND, Pickup RW, Jones JG, Head IM . (1997). Ecophysiological evidence that Achromatium oxaliferum is responsible for the oxidation of reduced sulfur species to sulfate in a freshwater sediment. Appl Environ Microbiol 63: 1905–1910.
Gray ND, Head IM . (1999). New insights on old bacteria: diversity and function of morphologically conspicuous sulfur bacteria in aquatic systems. Hydrobiologia 401: 97–112.
Gray ND, Howarth R, Pickup RW, Jones JG, Head IM . (1999a). Substrate uptake by uncultured bacteria from the genus Achromatium determined by microautoradiography. Appl Environ Microbiol 65: 5100–5106.
Gray ND, Howarth R, Rowan A, Pickup RW, Jones JG, Head IM . (1999b). Natural communities of Achromatium oxaliferum comprise genetically, morphologically, and ecologically distinct subpopulaitons. Appl Environ Microbiol 65: 5089–5099.
Gray ND, Comaskey D, Miskin IP, Pickup RW, Suzuki K, Head IM . (2004). Adaptation of sympatric Achromatium spp. to different redox conditions as a mechanism for coexistence of functionally similar sulphur bacteria. Environ Microbiol 6: 669–677.
Gray ND . (2006). The unique role of intrcellular calcification in the genus Achromatium. In: Shively JM (ed), Inclusions in Prokaryotes. Springer: Berlin, Heidelberg, pp 299–309.
Gray ND, Head IM . (2014). The family Achromatiaceae. In: Rosenberg E, Delong EF, Lory S, Stackebrandt E, Thompson FL (eds), The Prokaryotes: Gammaproteobacteria. Springer: Berlin-Heidelberg, pp 1–14.
Head IM, Gray ND, Pickup RW, Jones JG . (1995). The biological role of Achromatium oxaliferum. In: Grimalt JO, Dorronsoro C (eds), Geochemistry: Developments and Applications to Energy, Climate, Environment and Human History. AIGOA: Donostia-San Sebastian, Spain, pp 895–898.
Head IM, Gray ND, Clarke KJ, Pickup RW, Jones JG . (1996). The phylogenetic position and ultrastructure of the uncultured bacterium Achromatium oxaliferum. Microbiol 142: 2341–2354.
Head IM, Gray ND, Howarth R, Pickup RW, Clarke KJ, Jones JG . (2000). Achromatium oxaliferum - understanding the unmistakable. In: Schink B (ed), Advances in Microbial Ecology. Kluwer Academic/Plenum Publishers: New York, pp 1–40.
Hinze G . (1903). Thiophysa volutans, ein neues Schwefelbakterium. Ber Deutsch Bot Ges 21: 309–316.
Kolkwitz R . (1918). Über die Schwefelbakterienflora des Solgrabens von Arten. Berichte Deutsch Botanischen Gesellsch 36: 218–224.
La Rivière JVM, Schmid K . (1989). The genus Achromatium. Staley JT, Bryant MP, Pfennig N, Holt JG (eds), Bergey's Manual of Systematic Bacteriology. Williams and Wilkins: Baltimore, pp 2131–2133.
La Rivière JWM, Schmid K . (1992). Morphologically conspicuous sulfur-oxidising Eubacteria. In: Balows A, Trüper HG, Dworkin M, Harder W, Schleifer KH (eds), The Prokaryotes. Springer-Verlag: New York, pp 3934–3947.
Lackey JB, Lackey EW . (1961). The habitat and description of a new genus of sulphur bacterium. J Gen Appl Microbiol 26: 29–39.
Laetsch T, Downs RT . (2006) Software for identification and refinement of cell parameters from powder diffraction data of minerals using the RRUFF Project and American Mineralogist Crystal Structure Databases. 19th General Meeting of the International Mineralogical Association, Kobe: Japan.
Lakowicz JR, Szmacinski H, HJohnson ML . (1992). Calcium imaging using fluorescence lifetimes and long-wavelength probes. J Fluoresc 2: 47–62.
Lane DJ . (1991). 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds), Nucleic Acid Techniques in Bacterial Systematics. Wiley: Chichester, UK, pp 115–175.
Lauterborn H . (1915). Die sapropelische Lebewelt. Ein Beitrag zur Biologie des Faulschlamms natürlicher Gewässer. Verhandl Naturhistor Mediz Ver Heidelberg 13: 395–481.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al. (2004). ARB: a software environment for sequence data. Nucl Acids Res 32: 1363–1371.
Maier S, Gallardo VA . (1984). Thioploca araucae sp. nov. and Thioploca chileae sp. nov. Int J Syst Bacteriol 34: 414–418.
Manz W, Amann R, Ludwig W, Wagner M, Schleifer KH . (1992). Phylogenetic oligodeoxynucleotide probes for the major subclasses of proteobacteria: problems and solutions. Syst Appl Microbiol 15: 593–600.
Millero FJ, Plese T, Fernandez M . (1988). The dissociation of hydrogen sulfide in seawater. Limnol Oceanogr 33: 269–274.
Murray RGE, Schleifer K-H . (1994). Taxonomic notes - a proposal for recording the properties of putative taxa of prokaryotes. Int J Syst Bacteriol 44: 174–176.
Mussmann M, Hu FZ, Richter M, de Beer D, Preisler A, Jorgensen BB et al. (2007). Insights into the genome of large sulfur bacteria revealed by analysis of single filaments. PLoS Biol 5: 1923–1937.
Nadson GA . (1913). Über Schwefelmikroorganismen des Hapsaler Meerbusens. Bulletin du Jardin Impériale Botanique de St-Pétersbourg 13: 106–112.
Nadson GA . (1914). Über die Schwefelbakterien: Thiophysa und Thiosphaerella. Z Mikrobiol 1: 52–72.
Naraghi M . (1997). T-jump study of calcium binding kinetics of calcium chelators. Cell Calcium 22: 255–268.
Nelson DC, Castenholz RW . (1981). Use of reduced sulfur compounds by Beggiatoa sp. J Bacteriol 147: 140–154.
Nelson DC, Wirsen CO, Jannasch HW . (1989). Characterization of large, autotrophic Beggiatoa spp. abundant at hydrothermal vents of the Guaymas Basin. Appl Environ Microbiol 55: 2909–2917.
Pernthaler A, Pernthaler J, Amann R . (2002). Fluorescence in situ hybridization and catalyzed reporter deposition for the identification of marine bacteria. Appl Environ Microbiol 68: 3094–3101.
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig WG, Peplies J et al. (2007). SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucl Acids Res 35: 7188–7196.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P et al. (2013). The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41: D590–D596.
Revsbech NP, Jorgensen BB, Blackburn TH . (1983). Microelectrode studies of the photosynthesis and O2, H2S, and pH profiles of a microbial mat. Limnol Oceanogr 28: 1062–1074.
Salman V, Amann R, Girnth A-C, Polerecky L, Bailey JV, Høgslund S et al. (2011). A single-cell sequencing approach to the classification of large, vacuolated sulfur bacteria. Syst Appl Microbiol 34: 243–259.
Salman V, Bailey JV, Teske A . (2013). Phylogenetic and morphologic complexity of giant sulphur bacteria. Antonie Van Leeuwenhoek 104: 169–186.
Sambrook J, Russel DW . (2006). Purification of nucleic acids by extraction with phenol:chloroform. CSH Protocls. Nr. 1, pdb-prot4455.
Sanders R, Gerritsen HC, Draaijer A, Houpt PM, Levine YK . (1994). Fluorescence lifetime imaging of free calcium in single cells. Bioimaging 2: 131–138.
Schewiakoff W . (1892) Über einen neuen bacterienähnlichen Organismus des Süsswassers. Habilitation thesis, University Heidelberg: Heidelberg.
Schulz HN, Brinkhoff T, Ferdelman TG, Marine MH, Teske A, Jørgensen BB . (1999). Dense populations of a giant sulfur bacterium in Namibian shelf sediments. Science 284: 493–495.
Schulz HN, Jørgensen BB . (2001). Big bacteria. Annu Rev Microbiol 55: 105–137.
Schwedt A, Kreutzmann AC, Polerecky L, Schulz-Vogt HN . (2011). Sulfur respiration in a marine chemolithoautotrophic Beggiatoa strain. Front Microbiol 2: 276.
Seitz AP, Nielsen TH, Overmann J . (1993). Physiology of purple sulfur bacteria forming macroscopic aggregates in Great Sippewissett salt marsh, Massachusetts. FEMS Microbiol Ecol 12: 225–236.
Skuja H . (1948) Taxonomie des Phytoplanktons einiger seen in Uppland, Schweden. Symbolae Botanicae Upsaliensis. A.B. Lundequistska Bokhandeln: Uppsala, pp 1–399.
Soetaert K, Hofmann AF, Middelburg JJ, Meysman FJR, Greenwood J . (2007). The effect of biogeichemical processes on pH. Mar Chem 105: 30–51.
Starr MP, Skerman VBD . (1965). Bacterial diversity: the natural history of selected morphologically unusual bacteria. Annu Rev Microbiol 19: 407–454.
Van Niel CB . (1948). Family A. Achromatiaceae Massart. In: Breed RS, Murray EGD, Hitchens AP (eds), Bergey's Manual of Determinative Bacteriology, 6 edn, The Williams and Wilkins Company: Baltimore, pp 997–999.
Wallner G, Amann R, Beisker W . (1993). Optimizing fluorescent in situ hybridization with rRNA-targeted oligonucleotide probes for flow cytometric identification of microorganisms. Cytometry 14: 136–143.
West GS, Griffiths BM . (1909). Hillhousia mirabilis, a giant sulphur bacterium. Proc R Soc Lond B 81: 398–405.
West GS, Griffiths BM . (1913). The lime-sulphur bacteria of the genus Hillhousia. Ann Bot 27: 83–91.
Wilbanks EG, Jaekel U, Salman V, Humphrey PT, Eisen JA, Facciotti MT et al. (2014). A sulfurous symbiosis: microscale sulfur cycling in the pink berry consortia of the Sippewissett salt marsh. Environ Microbiol 16: 3398–3415.
Zhao M, Hollingworth S, Baylor SM . (1996). Properties of tri- and tetra-carboxylate Ca2+ indicators in frog skeletal muscle fibres. Biophys J 70: 896–916.
Zopfi J, Ferdelman TG, Fossing H . (2004). Distribution and fate of sulfur intermediates - sulfite, tetrathionate, thiosulfate, and elemental sulfur - in marine sediments. Amend JP, Edwards KJ, Lyons TW (eds), Sulfur Biogeochemistry - Past and Present. Geological Society of America, Special Paper 379: Boulder, CO, pp 97–116.
Acknowledgements
Much of this work was performed during several MBL Microbial Diversity summer courses. We warmly thank the course directors, Dan Buckley, Steve Zinder, Dianne Newman and Jared Leadbetter, as well as all course associates for the outstanding opportunity and support. Research at MBL was funded by the Howard Hughes Medical Foundation, the Simons Foundation, the Gordon and Betty Moore Foundation (2493), the National Science Foundation (DEB-0917499), the US Department of Energy (DE-FG02-10ER13361), the NASA Astrobiology Institute and the Beckman Foundation. In addition, we thank the Borisy lab at MBL, Ginny Edgcomb at WHOI and Becky Williams and Kit Umbach at Cornell University for lab space and technical support. Finally, we thank the editor and reviewers for their contribution to greatly improve this manuscript. VS is supported by the Deutsche Forschungsgemeinschaft (Sa 2505/1-1), TB by the ERC advanced Grant PARASOL (No. 322551), and TY received financial assistance at the MBL from the Horace W Stunkard Scholarship Fund.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Salman, V., Yang, T., Berben, T. et al. Calcite-accumulating large sulfur bacteria of the genus Achromatium in Sippewissett Salt Marsh. ISME J 9, 2503–2514 (2015). https://doi.org/10.1038/ismej.2015.62
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.62
This article is cited by
-
Correlative SIP-FISH-Raman-SEM-NanoSIMS links identity, morphology, biochemistry, and physiology of environmental microbes
ISME Communications (2022)
-
The diversity of molecular mechanisms of carbonate biomineralization by bacteria
Discover Materials (2021)
-
Intracellular amorphous Ca-carbonate and magnetite biomineralization by a magnetotactic bacterium affiliated to the Alphaproteobacteria
The ISME Journal (2021)
-
Microbial community composition and dolomite formation in the hypersaline microbial mats of the Khor Al-Adaid sabkhas, Qatar
Extremophiles (2019)
-
Intracellular calcite and sulfur dynamics of Achromatium cells observed in a lab-based enrichment and aerobic incubation experiment
Antonie van Leeuwenhoek (2019)