Abstract
Macroscopic growths at geographically separated acid mine drainages (AMDs) exhibit distinct populations. Yet, local heterogeneities are poorly understood. To gain novel mechanistic insights into this, we used OMICs tools to profile microbial populations coexisting in a single pyrite gallery AMD (pH ∼2) in three distinct compartments: two from a stratified streamer (uppermost oxic and lowermost anoxic sediment-attached strata) and one from a submerged anoxic non-stratified mat biofilm. The communities colonising pyrite and those in the mature formations appear to be populated by the greatest diversity of bacteria and archaea (including ‘ARMAN’ (archaeal Richmond Mine acidophilic nano-organisms)-related), as compared with the known AMD, with ∼44.9% unclassified sequences. We propose that the thick polymeric matrix may provide a safety shield against the prevailing extreme condition and also a massive carbon source, enabling non-typical acidophiles to develop more easily. Only 1 of 39 species were shared, suggesting a high metabolic heterogeneity in local microenvironments, defined by the O2 concentration, spatial location and biofilm architecture. The suboxic mats, compositionally most similar to each other, are more diverse and active for S, CO2, CH4, fatty acid and lipopolysaccharide metabolism. The oxic stratum of the streamer, displaying a higher diversity of the so-called ‘ARMAN’-related Euryarchaeota, shows a higher expression level of proteins involved in signal transduction, cell growth and N, H2, Fe, aromatic amino acids, sphingolipid and peptidoglycan metabolism. Our study is the first to highlight profound taxonomic and functional shifts in single AMD formations, as well as new microbial species and the importance of H2 in acidic suboxic macroscopic growths.
Similar content being viewed by others
Introduction
Acid mine drainage (AMD) formations are common on our planet. They result from the microbially catalysed oxidation of pyrite (FeS2) and other sulphide minerals, giving rise to the production of sulphuric acid and metal-rich solutions with a low pH (e.g., Rawlings, 2002). AMD formations extend across acidic niche boundaries, in which three distinct microenvironments may coexist: water, bed sediments and macroscopic structures, including streamers, slimes, mats, snottites and drapes. Among the AMD systems that have been characterised to date at the community level, approximately 30% have been reported to contain macroscopic microbial growths or biofilms (e.g., López-Archilla et al., 2004; Tyson et al., 2004; García-Moyano et al., 2007; Macalady et al., 2007; Ziegler et al., 2009; Johnson, 2012; Jones et al., 2012).
There is increasing evidence that these acidic niche boundaries harbour a restricted set of acidophilic Bacteria and Archaea (Bond et al., 2000; Baker and Banfield, 2003; Hallberg et al., 2006; He et al., 2008; Yin et al., 2008; Baker et al., 2009) and Eukarya (Aguilera et al., 2006; Baker et al., 2009). Bacteria that are widely distributed in these acidic ecosystems include, albeit in different proportions, those belonging to the Proteobacteria, Nitrospirae, Actinobacteria, Firmicutes, Bacteroidetes and Acidobacteria phyla. The members of these taxa are extreme acidophiles displaying pH optima below 3 and optimal mesophilic growth temperatures ranging from 17 to 45 °C (Sánchez-Andrea et al., 2011). In terms of their metabolic attributes, the bacterial members of these assemblages include chemolithoautotrophs that obtain energy mainly from iron, sulphur and hydrogen metabolism as well as mixotrophs and heterotrophs. The Archaea observed in AMDs generally belong to the order Thermoplasmatales. The metabolism of the described representatives of this order is exclusively organotrophic, with the exception of members of Ferroplasmaceae, which are able to metabolise iron (Golyshina, 2011). Baker and Banfield (2003) detected many archaeal signatures, divided by the authors into numerous groups affiliated with the order Thermoplasmatales, representing a high proportion of uncultured Archaea that occur in metal-rich and low-pH habitats. Although these Archaea populate iron-rich environments, their metabolism has been speculated to most likely rely on the bacterial production of a steady supply of organic substrates, for example, sugar- and lipid-related compounds and polymers involved in biofilm formation (Ram et al., 2005; Belnap et al., 2010; Denef et al., 2010; Moreno-Paz et al., 2010; Mueller et al., 2010; Belnap et al., 2011; Mueller et al., 2011) or released by other free-living autotrophic acidophiles (Nancucheo and Johnson, 2010). Various lineages of filterable ultramicroarchaea, referred to as the archaeal Richmond Mine acidophilic nano-organisms (‘ARMAN’), have recently been detected in AMD biofilms (Sunna and Bergquist, 2003; Baker et al., 2006; Juottonen et al., 2008; Comolli et al., 2009; Ziegler et al., 2009; Baker et al., 2010; Amaral-Zettler et al., 2011; Bohorquez et al., 2012; Ziegler et al., 2013). ‘ARMAN’-related populations are believed to have a minor active role in AMD systems (yet to be defined because of the absence of cultivable members), as demonstrated by a very low contribution to the total metaproteome (0.7–1.8% of peptides), although they comprise ∼17% of the total metagenome (Baker et al., 2010).
Research directed at exploring the metabolic capacities of community members has revealed that the chemical species present in oligotrophic AMD systems favour the establishment of a microbial chemolithotropic community (Tyson et al., 2004; Ram et al., 2005; Rowe et al., 2007; Belnap et al., 2010; Amils et al., 2011; Bertin et al., 2011; González-Toril et al., 2011; Johnson, 2012). However, when macroscopic microbial growths develop, the predominant metabolism shifts from chemolithotrophy to heterotrophy (López-Archilla et al., 2004; García-Moyano et al., 2007; Macalady et al., 2007; Ziegler et al., 2009; Johnson, 2012; Jones et al., 2012). Furthermore, a number of recent studies have indicated that there is a clear correlation between the phylotype richness and the pH conditions in situ (Sánchez-Andrea et al., 2011; Kuang et al., 2013).
This study presents a comparative OMIC analysis of three streamer/mat-shaped macroscopic growths along two distinct microenvironments in a mercury AMD formation (Los Rueldos) that displays a homogeneous pH of ∼2. This study has four aims: (1) to attain a comprehensive geochemical characterisation of the environment where these growths develop, (2) to profile the total microbial community and the streamer evolution dynamics, (3) to investigate whether shifts in the microbial populations are associated with geochemical and spatial gradients and (4) to provide a detailed characterisation of the metabolic shifts responding to microbial stratification through community genomics and proteomics analyses in each of the sampled populations. To the best of our knowledge, this study represents the first comparative integrative OMIC study investigating distinct macroscopic growths, including uncommon low pH suboxic macroscopic growths, along a single AMD formation.
Materials and methods
Study site
The sampling site was located along the north-western slope of the valley of the Morgao stream (2 km northeast of the town of Mieres and 20 km southeast of Oviedo, the capital city of Asturias, NW Spain; 43°15′47′′N, 5°46′9′′W) (Supplementary Figure S1). In this area, a number of prospective galleries were originally excavated from 1960 to 1970 at different altitudes on the mountainsides in search of sulphide ore bodies mainly comprising pyrite, arsenopyrite and cinnabar. In the now-abandoned galleries and spoil heaps, herein referred to as the Los Rueldos site, sulphides were exposed to oxygen and the humid climate of the area, favouring the development of acid leachates of infiltrated water in a stable environment. The epigenetic deposit contains Hg, mainly in the form of cinnabar (HgS), accompanied by other sulphides, including pyrite (FeS2, occasionally As-rich pyrite), melnikovite (Fe3S4), sphalerite (ZnS), marcasite (FeS2 with an orthorhombic crystal structure), chalcopyrite (CuFeS2), arsenopyrite (FeAsS), galena (PbS), stibnite (Sb2S3, also known as antimonite) and realgar (As4S4) (Loredo et al., 2005). Most of these minerals are now partially weathered, and as a consequence, oxide/oxyhydroxide and sulphate minerals are also present within the site. The average annual temperature, relative humidity and rainfall are 13.8±0.6 °C (ranging from 10±0.6 to 17.1±0.6 °C), 78% and 1007.9±175.5 mm, respectively.
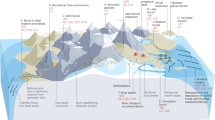
In a specific area of Los Rueldos, we identified a 70-m-long gallery that exhibits different streamer/mat biofilm morphologies (Figure 1). The gallery (1.5–2 m in height) appears to have been an excavation in the middle of the mountain slope that collapsed approximately 70 m beyond the entrance. It presents an almost non-flowing AMD with a pH of ∼2 generated by rainwater that percolates through the cave ceiling. The proliferation of some algae is observed within the first 2–3 m of the gallery. The AMD accumulates in a puddle with a depth of 30 cm at the entrance area, where the less restrictive conditions favour the presence of eukaryotes. The 70-m-long AMD bed varies in depth from 10–30 cm, in the shallower sections, to 40–70 cm in the deeper areas, corresponding to the presence of different streamer/mat morphologies. Thus, at 25 m in from the gallery entrance, at an average depth of ∼15 cm, a 4- to 5-m-long stratified streamer (referred to as the uppermost B1A and sediment-attached, lowermost B1B strata) appears, which extends throughout the entire AMD column. Approximately 10 m in from this upwelling, at a location with a depth of ∼50 cm, a different, non-stratified mat biofilm (B2) becomes visible. This biofilm displays a discrete rounded shape distributed within a 3-m-long area at a location with a considerable number of stalactites dripping onto the AMD. After another 10 m, the stratified streamer morphology appears again and extends until the collapse, which is situated in a region with a large number of stalactites and stalagmites.
Gallery scheme, showing the location of the main macroscopic growths (env1 microenvironment, where B1A and B1B develop, and env2, where B2 upwellings appear) found along the Los Rueldos AMD system. Main geochemical characteristics of the drainage are shown; metal and sulphate concentrations are represented in mg l−1. Corresponding DO profiles to each microenvironment are displayed. Photographs illustrating the general aspect of the main microbial formations within the Los Rueldos AMD are included for further clarification. The reappearance of the B1 morphology at the end of the gallery is illustrated. The discontinuous line on the top represents the gallery ceiling with the presence of stalactites.
Experimental settings and data analysis
Samples were collected in sterile 50 ml tubes in June 2010 at three sampling sites (B1A: 2 cm deep; B1B: 15 cm deep; B2: 50 cm deep) determined by the presence of each different macroscopic microbial growth morphology. The samples were kept on ice until processing (within 2 h). The full description of the Materials and methods used for (a) the environmental parameter determinations, (b) nucleic acid extraction and full-length 16S rRNA library construction and sequencing, (c) small subunit (SSU) rRNA hypervariable tag sequencing, (d) streamer formation dynamics assay (including fluorescent in situ hybridisation and X-ray photoelectron spectroscopy analyses), (e) phylogenetic analysis, (f) DNA sequencing and assembly as well as gene prediction and annotation, (g) Kyoto Encyclopaedia of Genes and Genomes (KEGG) enrichment profiling, (h) metaproteomic setup, (i) metaproteome-scale metabolic reconstruction, (j) cell counting and (k) biochemical tests are available in the Supplementary Materials and methods.
Accession numbers
The projects have been registered as umbrella BioProject at NCBI with the ID PRJNA193663 (for B1A), PRJNA193664 (for B1B) and PRJNA193665 (for B2A). This Whole Genome Shotgun project has been deposited at DDBJ/EMBL/GenBank under the accession AUZX00000000–AUZZ00000000. The GenBank accession numbers for 16S rRNA nucleotide sequences are KF225643–KF225695 (for Sanger sequences) and KF758948–KF760235 (for hypervariable tag sequences).
Results and discussion
Geochemical and physical parameters in streamer/mat macroscopic growths
Our aim was to provide a detailed characterisation of the geochemical characteristics of the uppermost oxic B1A and lowermost suboxic B1B strata and the non-stratified suboxic mat biofilm (B2) and to compare how the sampled populations differ at the levels of microbial phylotype, functional gene content and metabolic status. Both the microenvironments and corresponding streamer/mat-shaped macroscopic growths were characterised by a pH of ∼2 and an average annual temperature of ∼13 °C. They displayed a redox potential (Eh) of ∼256 mV and conductivities of 5.14 to 6.72 mS cm−1 throughout the year (Supplementary Figure S2), which are among the lowest values reported for AMD formations. They displayed metal/metalloid contents of 422 mg l−1 Fe, 129 mg l−1 Al, 5 mg l−1 As, 0.013 mg l−1 Sb, 83 ng l−1 Hg and 7.6 μg l−1 Pb (Supplementary Table S1). The multivariate statistical analysis indicated no significant geochemical differences among the three sampling sites, except that the submerged B1B (0.5±0.02 mg l−1) and B2 (0.25±0.01 mg l−1) mats at the sampling point exhibited a lower dissolved oxygen (DO) concentration compared with B1A (2.50±0.02 mg l−1), located in the air–solution interface. The DO profiles for the B1 and B2 microenvironments are displayed in Figure 1.
Microbial life in the Los Rueldos AMD is strongly stratified
Full-length 16S rRNA and SSU rRNA hypervariable tag sequencing allowed the depth variations in the most abundant microbial populations to be compared with the geochemical and spatial variations in the selected contextual data. A total of 957 full-length 16S (B1A: 418; B1B: 275 and B2: 264) and 44 469 SSU (>150 nucleotides long; B1A: 9842, B1B: 23 727 and B2: 10 900) rRNA sequences were obtained and analysed.
Bacterial stratification
The bacterial sequences were grouped into 1005 operational taxonomic units according to the SSU rRNA hypervariable tag sequencing data (97% sequence identity level), with the majority belonging to the Proteobacteria phyla (>50% of the total sequences in all samples, with a variable proportion (1.5–2.4%) of unclassified SSU rRNA sequences). The γ- (B1A: 17.8–45%; B1B: ∼87% and B2: 31–58%), β- (B1A: 28–75%; B1B: 0.2% and B2: 13–23%) and α-Proteobacteria (B1A: 0.6–1.3%; B1B: 0.03% and B2: 0.75%) classes were found, in that order. A number of sequences were also assigned to Acidobacteria (0.03–6% in all communities), Nitrospirae (B1A: 4.5–8%; B1B: 1.7–2.3% and B2: 6–11%) and Actinobacteria (B1A: 2–7%; B1B: 0.2–8% and B2: 4–18%), whereas other sequences were assigned to the candidate divisions WS6/TM7 (<0.5% in all samples) and AD3 (0.02–0.06%), Cyanobacteria (0.02% in B1A), Elusimicrobia (0.02% in B1A) and Lentisphaerae (0.01% in B1B) (Supplementary Figure S3). In addition, according to a cutoff at 90% sequence identity level, 0–8.56% (for B1A), 0–8.98% (for B1B) and 9.0–14.11% (for B2) corresponded to unclassified bacterial full-length 16S and SSU rRNA sequences (average lowest and highest proportions shown).
The sequences were further classified according to operational phylogenetic units (OPUs, equivalent to distinct taxonomic species; López-López et al., 2010) (Supplementary Table S2). The 18 identified bacterial OPUs were distributed among 13 major clusters based on clear phylogenetic positioning (Figure 2; for details see Supplementary Results and discussion). None of the OPUs were shared among the three macroscopic growths (Figure 3a), suggesting that bacterial life in the Los Rueldos AMD is strongly stratified. Notably, the OPUs belonging to B2 are widespread among all phyla (Figure 2). The Shannon–Wiener and Chao-1 indices indicated that B2 displayed higher complexity when the data were obtained from full-length 16S rRNA, and an evenness value close to 1 indicated that dominance was shared among the various taxa in this sample. Nevertheless, the diversity indices obtained through SSU rRNA hypervariable tag sequencing revealed B1B as the more diverse sample (Supplementary Table S3).
Maximum-likelihood phylogenies for the bacterial 16S rRNA full-length sequence amplicons retrieved from the constructed gene clone libraries. Scale bars represent changes per site or the ( × 100) % difference in nucleotide sequences. The main phylogenetic clusters are highlighted in grey. Bootstrap values are labelled at their corresponding nodes with circles: >85%, closed circles; >50%, open circles. The contribution of each OPU to the library is represented in the accompanying graphs. Representative sequences for each sample are differentiated by a colour code: B1A, red; B1B, blue; B2, green.
Venn diagram showing the common and distinct distribution of the number of (a) bacterial, archaeal, ‘ARMAN’-related and total OPU-based species (in this order from the left to the right), (b) potential protein-coding genes as assigned to pyrosequences, (c) proteins being expressed as found in the metaproteomes and (d) metabolic reactions associated to proteins being expressed.
Archaeal stratification
The results from SSU rRNA hypervariable tag sequencing revealed the existence of 383 archaeal operational taxonomic units (97% sequence identity level). The Shannon–Wiener and Chao-1 indices derived from the full-length 16S rRNA sequence data indicated that B2 displayed the highest archaeal complexity and that B1B exhibited higher archaeal diversity when the data were obtained through SSU rRNA hypervariable tag sequencing (Supplementary Table S3). We observed that none of the archaeal sequences were shared among the three macroscopic growths (Figure 3a) and that the B1A and B2 sequences appeared to be phylogenetically more related, even though they differ significantly in the O2 concentration (Figure 1). Most of the detected archaeal sequences belonged to the phylum Euryarchaeota, order Thermoplasmatales, although an important fraction of the full-length 16S and SSU rRNA sequences in B1A and B1B were affiliated with unclassified sequences (Figure 4). In fact, 6.15–55.7% (for B1A), 2.46–44.12% (for B1B) and 0.32–4.76% (for B2) corresponded to unclassified archaeal full-length 16S and hypervariable rRNA sequences (average lowest and highest proportions shown). Among the 14 identified OPUs, 5 units per groups of clones were related to uncultured Thermoplasmatales lacking clear, up-to-date, taxonomic information (Figure 4 and Supplementary Table S2). Cluster 1 was composed of a set of B1A and B1B sequences that the SILVA database (Yarza et al., 2008; Pruesse et al., 2012) associates with Terrestrial Miscellaneous Gp (TMEG), placing it within the order Thermoplasmatales, where OPUs 10–14 were distant from the nearest described relative, Methanomassiliicoccus luminyensis (from the recently proposed order with still unclear name, candidate order ‘Methanoplasmatales’ or ‘Methanomassiliicoccales’ ord. nov) (Paul et al., 2012; Iino et al., 2013). The rest of the sequences were associated with the genus Thermoplasma in accordance with the SILVA reference database or were identified as distantly related to Thermogymnomonas acidicola as per the Ribosomal Database Project (RDP) database. Between the two clusters containing these more diverse groups of these sequences, one was absent in B1B. As shown in Supplementary Figure S3, other groups of sequences of clones detected in the B1B and B2 samples were represented by the previously designated group E1/E2 of uncultured euryarchaea, which is now referred to as a family-level clade within the order Methanomicrobiales and family Methanoregulaceae (Sakai et al., 2012). Putative methanogens accounted for a high proportion of the microorganisms in all three communities and were the second-most numerous archaeal group (OPUs 10–14) (Figure 4).
Maximum-likelihood phylogenies for the archaeal 16S rRNA full-length sequence amplicons retrieved from the constructed gene clone libraries. Scale bars represent changes per site or the ( × 100) % difference in nucleotide sequences. The main phylogenetic clusters are highlighted in grey. Bootstrap values are indicated at the corresponding nodes with circles: >85%, closed circles; >50%, open circles. The contribution of each OPU to the library is represented in the accompanying graphs. Representative sequences for each sample are differentiated by a colour code: B1A, red; B1B, blue; B2, green.
Filterable ultramicroarchaeal (‘ARMAN’-like) stratification
Sequencing of the SSU rRNA hypervariable regions revealed the presence of 254 ‘ARMAN’-like operational taxonomic units (97% sequence identity level), and phylogenetic analysis of the retrieved ‘ARMAN’-like sequences revealed the existence of seven putative OPUs (one shared among communities; Figure 3a). There were no clear differences in the composition among the samples as all of the obtained sequences appeared to be widespread in B1A, B1B and B2. However, B1A appeared to be the community displaying the majority of ‘ARMAN’-related diversity, whereas only one type of ‘ARMAN’-related clone was detected in B2 (Figure 4 and Supplementary Figure S4 and Supplementary Table S3). The major homology of the Los Rueldos sequences was shared with sequences obtained from an acidic biofilm collected in the Harz Mountains (Germany) (Ziegler et al., 2009, 2013). Approximately 18.38–100% (for B1A), 18.46–100% (for B1B) and 19.79–100% (for B2) sequences corresponded to unclassified ‘ARMAN’-like full-length 16S and SSU rRNA sequences (average lowest and highest proportions shown). The seven OPUs could be compartmentalised into three clusters defined by homology with known ‘ARMAN’-related archaeal sequences (Figure 4 and Supplementary Figure S4 and Supplementary Table S2; for details see Supplementary Results and discussion).
Altogether, these results indicate that Los Rueldos appears to contain communities comprising greater major Bacteria and Archaea richness and diversity compared with other biofilm-containing AMD formations (Figure 5). Fluorescent in situ hybridisation (Supplementary Figure S5) and taxonomic profiling through SSU rRNA tag pyrosequencing (Supplementary Table S4 and Supplementary Figure S6) analyses indicated that the Los Rueldos clusters also displayed greater diversity at early developmental stages (for details see Supplementary Results and discussion). Los Rueldos exhibits a high proportion of uncommon Bacteria known to inhabit anoxic subsurfaces, uncultured Thermoplasmatales (including ‘ARMAN’-group Archaea) and, to a lesser extent, putative methanogenic Archaea, but not the typical genera found in described AMD formations (Bond et al., 2000; Baker and Banfield, 2003; Hallberg et al., 2006; He et al., 2008; Yin et al., 2008; Baker et al., 2009; Wilmes et al., 2009). Notably, Los Rueldos contains ∼44.95% and 11.84% of unclassified full-length 16S and hypervariable rRNA region sequences, respectively. We suggest that the distinct environmental features that characterise Los Rueldos (Supplementary Table S1) may account for these differences and for the presence of new microbial species. The Los Rueldos microenvironments were observed to share a rather small core set of microorganisms that are not highly prevalent as only 1 (or 2.6%) out of 39 OPUs, representing a fraction of the ‘ARMAN’-related sequences that remain unrelated to any known database sequences, were shared among the three communities (Figure 3a). This result indicates that the Los Rueldos AMD contains a wide set of distinct microorganisms that form not only an extensive but also a stratified ecosystem. The lowermost/submerged (suboxic) mats were most similar to each other and exhibited the highest diversity, being particularly enriched in sequences representing presumptive low-O2-adapted organisms related to the Acidithiobacillales ‘RCP1-48’ cluster, Acidobacteria, Leptospirillum ferrooxidans-like (all described Leptospirillum are known to be obligate aerobes), Acidimicrobiaceae-related Actinobacteria, Firmicutes and Lentisphaerae, associated with anoxic environments (Hedlund et al., 2011), candidate divisions WS6 and TM7, and candidate order ‘Methanoplasmatales’/or ‘Methanomassiliicoccales’. The sequences related to Rhizobium (α-Proteobacteria), ‘L. ferrodiazotrophum’, candidate division AD3, detected for first time in sandy surface soils (Zhou et al., 2003), Cyanobacteria and Elusimicrobiales, found in both oxic and anoxic environments, were mostly associated with the uppermost (oxic) mat, which also displayed a higher diversity of Archaea related to the ‘ARMAN’ group. Marked taxonomic stratification was observed at the genus level within the Los Rueldos communities. L. ferrooxidans-related bacteria, able to thrive at depths below 60 cm (Kimura et al., 2005), and the non-motile Acidithiobacillus ferrooxidans (Valdés et al., 2008) appeared within the suboxic strata of the macroscopic growths (B1B and B2), whereas Acidithiobacillus ferrivorans, motile rods (Hallberg 2010), and ‘L. ferrodiazotrophum’-related bacteria, able to perform atmospheric N2 fixation (Tyson et al., 2005), thrived in the oxic B1A layer. Other organisms, such as those belonging to Acidobacteria, common in (acidic) terrestrial locations, Firmicutes, which have been detected in AMD anaerobic sediments (e.g. Sulfobacillus spp., Sánchez-Andrea et al., 2011), most Actinobacteria (autotrophic or heterotrophic aerobes or anaerobes) and the candidate divisions WS6, abundant community members in organic-rich anaerobic environments (Dojka et al., 2000) and TM7, were absent in the oxic layer of the streamer. In addition, as no significant changes were identified in the solution parameters of pH, temperature, conductivity and redox potential within the microenvironments, the obtained results confirmed the importance of combining O2 bioavailability and spatial distance in fine-tuning the acidophilic microbial structure.
The main differences at the level of microbial diversity between the Los Rueldos macroscopic growths and similar macroscopic structures previously reported from other AMD sites. (a) Mynydd Parys mine streamer (Hallberg et al., 2006). (b) Tinto River macroscopic floating filaments (López-Archilla et al., 2004). (c) Los Rueldos mine streamer/mats (this study). (d) Iron Mountain mine acidic biofilm (Tyson et al., 2004).
Genomic shifts responding to microbial stratification
The total extracted DNA was directly pyrosequenced using a Roche GS FLX DNA sequencer, which produced a total of ∼14.4 (for B1A), ∼14.9 (for B1B) and ∼10.9 (for B2) Mbp of assembled DNA sequences. Approximately 57 815 (B1A: 22 162; B1B: 19 692 and B2: 15 961) potential protein-coding genes (cutoff of ⩾20 amino acids) were identified (Supplementary Table S5). The obtained hits were filtered based on a minimum e-value (1e−05) and an alignment length of 75% minimum, and 19% were found to be shared among the three communities (Figure 3b).
General predictive metabolic features of Los Rueldos
KEGG subsystem-based JCoast annotations were used to gain a better understanding of how these population shifts could influence their metabolic potential (Figure 6). To extend this analysis, we compared the results with the Richmond AMD (AMD.R) metagenome (Tyson et al., 2004). Common features included an over-representation of ‘Carbohydrate Metabolism’, ‘Amino Acid Metabolism’, ‘Replication and Repair’ and ‘Membrane Transport’ and an under-representation of ‘Cellular Processes’ (Figure 6). In contrast, the distribution of the subsystems ‘Glycan Biosynthesis and Metabolism’ (B1A: 2.67% rel. abundance level; B1B: 2.21%; B2: 2.19% and AMD.R: 1.23%), ‘Signal Transduction’ (B1A: 4.23%; B1B: 2.83%; B2: 3.38% and AMD.R: 1.52%), ‘Cell Growth and Death’ (B1A: 0.77%; B1B: 0.76%; B2: 0.74% and AMD.R: 0.34%) and ‘Translation’ (B1A: 6.51%; B1B: 5.45%; B2: 6.56% and AMD.R: 10.35%) appeared to be significantly different for the B1A, B1B and B2 populations compared with the AMD.R. In addition, the abundance levels of the corresponding pooled proteins assigned to the ‘Cell Motility’ subsystem were also particularly high in B1A (B1A: 3.16%; B1B: 1.37%; B2: 1.82%; AMD.R: 1.52%). The significant enrichment in ‘Cell motility’ and, to a lower extent, in ‘Signal transduction’ in B1A appears to agree with the higher protein expression levels of chemotaxis, flagellar, efflux and cell division proteins (for details, see Supplementary Results and discussion and Supplementary Table S6) and with the observed microbial stratification and presence of the highly motile At. ferrivorans (Hallberg, 2010) in the uppermost B1A layer, which is more exposed to the flow influence than the B1B and B2 strata, where flagella-absent At. ferrooxidans (Valdés et al., 2008) thrives (Figure 2). The heatmap analysis (Figure 6) also revealed that all three samples clustered separately from AMD.R, suggesting that functional differences may exist between both AMD formations, regardless of the microenvironment considered in Los Rueldos, as it was observed at the population structure level (Figure 5). Notably, the B1B sample was functionally closer to B2 than to B1A (Figure 6), even though they belong to different microenvironments. This finding may agree with the observations that both B1B and B2 hold a higher similarity in terms of microbial diversity (Figures 2 and 4) and that both conformed the suboxic parts of the microbial mats.
KEGG profiling analysis of the Los Rueldos and Richmond (AMD.R) AMD community genomes. Hierarchical clustering and heatmap colours are based on functional compositions of KEGGs as determined by the relative percentages of KEGG Orthology (KO) identifiers assigned to particular KEGG categories. The colour scale quantifies the relative percentages from one sample to the other, and goes from red (highly abundant) to black (abundant) to light green (low abundant).
Selection and evolution of key metabolic traits
We attempted to obtain insights into whether each population has selected and evolved different capabilities for the turnover of major chemical species (S, N, CO2, H2 and Fe). Although members of all three communities have the potential to contribute to the cycling of these species, a few notable observations should be mentioned (Figure 7). First, the presence of genes encoding rhodaneses, known to be activated at elevated concentrations of toxic sulphides as a defence mechanism, was 2.4-fold higher in B1B than in B1A or B2, suggesting that B1B is most likely a permanent euxinic formation compared with the other macroscopic growths. Second, nitrification/denitrification processes have minor importance in all three communities compared with the S or Fe oxidation processes. Indeed, the B1A ecosystem was the sample with a high abundance of genes for rusticyanins, sulphocyanins and cbb3-type cytochrome c oxidases (∼2- and 1.3-fold more abundant than in B1B and B2) and enzymes (e.g., TusD and DsrE) for the oxidation of sulphide (HS−), which were ∼3-fold less abundant in B1B and absent in B2. Enzymes indicative of the putative presence of sulphur or reduced inorganic sulphur compounds metabolism such as sulphide:quinone reductases, often associated with acidophilic bacteria (Quatrini et al., 2009; Liljeqvist et al., 2011; Liljeqvist et al., 2013), were also encountered in all three metagenomes, although at very similarly low abundance level. Interestingly, whereas genes encoding nitrogenase MoFe proteins for nitrogen fixation were present at similar levels in B1A and B1B but absent in B2, those for nitrate reduction were most abundant in B2 and B1B (∼2-fold more abundant than in B1A). This agrees with the presence of ‘L. ferrodiazotrophum’ in B1A and B1B biofilms; in addition, BLAST searches of the NCBI non-redundant database with the nitrogenase MoFe protein sequences showed that at least 44% of them do have as best hit a homologous protein from this bacterium (identity ranging from 37% to 94%). According to thermodynamic requirements, the respiration of both reduced S and Fe is a highly exergonic process, and in the presence of high amounts of electron donors, these respiration types could support elevated growth, yielding significant amounts of organic matter in a form of such as polysaccharides (Moreno-Paz et al., 2010). The latter compounds are well-known substrates of anaerobic fermenters and organotrophic hydrolytic microorganisms. Notably, the main end products of fermentation activity under acidic, low saline conditions are mainly short-chained fatty acids, CO2 and H2. Accordingly, the diffusion of gases into the overlaying strata of the biofilm can support the autotrophy because of a hydrogenoclastic mode of carbon fixation, generally assumed to be common in the majority of chemolithoautotrophic acidophiles. Thus, hydrogen has to be included as an electron donor, in addition to the S and Fe described above. This assumption is consistent with the elevated presence of [Ni-Fe] hydrogenases (6- and 1.4-fold more abundant than in B1B and B2) and ribulose-1,5-bisphosphate carboxylases (∼3-fold more abundant than in B1B and B2) in the B1A uppermost layer, the main activities of which involve gaining reducing equivalents from molecular hydrogen and supporting the Calvin–Benson–Bassham cycle of carbon fixation, respectively. Interestingly, a comparison between the suboxic B1B and B2 mats revealed that whereas CO2 fixation gene signatures were observed at a similar level in both microbial formations, the genes encoding hydrogenases were particularly abundant in B2 (4-fold more abundant than in B1B).
Heat map and clustering of predicted genes encoding enzymes for the sulphur, nitrogen, CO2, H2 and iron metabolism in the Los Rueldos AMD macroscopic growths. The heatmap colours represent the relative percentages of distinct presumptive reactions assigned to particular enzymes within each sample. Hierarchical clustering based on relative percentage of predicted pooled genes encoding proteins involved in specifically transformations is shown. Note: a total of 438 (or 0.76%) of the total ORF fragments (B1A: 185; B1B: 145 and B2: 108) were identified as showing close sequence similarity to genes that encode enzymes known to be involved in the turnover of S, N, CO2, H2 and Fe and, accordingly, presumable functionalities were established and accounted in this figure. Colour scale as in Figure 6.
Taken together, these results suggest higher cell densities in the uppermost oxic B1A mat, as it was particularly enriched and active in multiple exergonic processes. This observation was further supported by 4′,6-diamidino-2-phenylindole-staining-based cell counting: ∼5 × 1012± 1.36 × 1012, ∼2 × 1012±1.14 × 1012 and ∼1.5 × 1012±2.75 × 1012 cells per cm3 in the B1A, B1B and B2 structures, respectively. A global analysis of the appearance and frequencies of the recovered phylogenetic groups led us to conclude that all three studied ecosystems have many metabolic parameters in common. Notably, however, the presence of oxygen appeared to prevail as a main shaping factor in the B1A ecosystem, which differentiates it from the other two systems. On the basis of the gathered data, B1B and B2, the suboxic parts of the mats, are highly reduced ecosystems inhabited by Proteobacteria, different groups of unclassified Euryarchaeota related to the ‘ARMAN’ group and members of the newly proposed candidate order ‘Methanoplasmatales’/‘Methanomassiliicoccales’ ord. nov (Paul et al., 2012). Interestingly, the methylotrophic modification of methanogenesis, obligately relying on hydrogen uptake, has been proposed as a metabolic priority for the members of ‘Methanoplasmatales’/‘Methanomassiliicoccales’, which were observed to be particularly enriched in B2 compared with B1B. As it has also been recently demonstrated in mature ‘F. acidarmanus’ Fer1 biofilms (Baker-Austin et al., 2010), a second contributing factor to the metabolic differences observed may be related to the biofilm architecture and development, which can produce a shift in anaerobic lifestyles by affecting the diffusion of gases from the lowermost to the overlaying strata (e.g., reduced S), the access to substrates from the atmosphere (e.g., nitrogen) or the accumulation of toxic compounds (e.g., toxic sulphides). The differences observed between B1B and B2 at the level of hydrogen and sulphide metabolism underscore this hypothesis.
Shotgun metaproteome measurements and metabolic shifts
We selected a shotgun metaproteomic approach to query the active populations and to ascertain their metabolic status within the mats. A total of 3407 proteins (B1A: 1472; B1B: 970 and B2: 965) were unambiguously identified and quantified (Supplementary Table S6; for details, see Supplementary Materials and methods). Among them, 1589 were unique proteins (i.e., proteins exclusive to any/each of the three locations), more than 99% of which could be taxonomically assigned. As shown in Supplementary Figure S8, the protein expression levels revealed exponential distributions, with B1B displaying the highest range of protein expression levels (138.32–0.0046 mmol mol−1), followed by B2 (124.35–0.0044 mmol mol−1) and, to a much lower extent, B1A (33.45–0.0054 mmol mol−1). Only 206 out of 1589 (or ∼13%) uniquely expressed proteins were shared by all three communities, suggesting high divergence in the active member composition (Figure 3c). B1A and B1B, albeit in the same microenvironment, shared the least number of active proteins (286 out of 1589), whereas B1B and B2 (415) and B1A and B2 (517) were more similar in terms of the active protein content (Figure 3c). The observation that a higher number of OPUs were shared between B1B and B2 (7 OPUs) and between B1A and B2 (5) compared with B1A and B1B (only 3) (Figure 3a) may account for the observed differences. The contribution of sequences belonging to Bacteria (B1A: 89%; B1B: 81% and B2: 85% total proteins) was significantly higher than that associated with Archaea (B1A: 10%; B1B: 18% and B2: 14%), although the suboxic mats contain a slightly higher contribution of archaeal proteins (Supplementary Figure S8). Eight (B1A: 3; B1B: 3 and B2: 2) were proteins assigned to Eukaryota. Proteobacteria was the major contributor (from 64% to 69%) to the total community proteomes, followed to a much lower extent by Actinobacteria (from 8.4% to 11.7%) and Nitrospirae (from 3.4% to 5.6%) in all sampled populations. The only major differences were observed in B1B at the level of Firmicutes proteins (∼6.7%), which accounted for ∼0.3% in B2 and ∼0.1% in B1A.
By using a previously published method (Hernández et al., 2013; for details see Supplementary Materials and methods), a total of 489 predicted metabolic reactions (B1A: 201; B1B: 141 and B2: 147) associated with expressed proteins were identified. Only 65 (or ∼13%) reactions appeared to be shared by the three communities (Figure 3d). The maps displaying changes in the various reactions are shown in Supplementary Figure S9. The classification of the pathways obtained in the samples (heat map: Supplementary Figure S10) showed the same functional similarity between samples B1B and B2 as was identified by KEGG profiling (Figure 6). Supplementary Figure S10 shows several notable findings. First, the high expression level of proteins involved in propanoate and pyruvate metabolism was common to all three communities. The fact that the main end products of fermentation activity under acidic conditions are mainly short-chained fatty acids agrees with the relevant preponderance of propanoate metabolism in the Los Rueldos microenvironments. Nitrogen (3- and 6-fold), tyrosine (17.3- and 5.8-fold), sphingolipid (4.0-fold each), purine (2.0-fold each) and histidine (2.0-fold each) metabolism and peptidoglycan biosynthesis (57- and 2.5-fold) were clear indicators of the differences between the samples and allowed B1A active metabolism to be distinguished from that in B1B and B2 (pooled expression level fold changes within parentheses). The anoxic B1B and B2 mats were characterised by higher abundances and expression levels of proteins for the butanoate (2.8- and 4.2-fold), CO2 (3-fold each), CH4 (3-fold each), fructose and mannose (2-fold each) and fatty acid (2.2- and 3.9-fold) metabolism as well as lipopolysaccharide biosynthesis (11-fold each) compared with the oxic B1A mat. In addition, the metabolism of a number of compounds was most active in both B1A and B1B compared with B2 (4.0-fold), for example, amino and nucleotide sugars (4.8- and 4.4-fold), alanine, aspartate and glutamate (4-fold each) as well as nicotinate and nicotinamide (4.1- and 2.6-fold). Finally, valine, leucine and isoleucine metabolism and folate biosynthesis were presumably the most active in B1B compared with B1A (2.4-fold) and B2 (7.0-fold).
The differences at the levels of sphingolipid metabolism and peptidoglycan biosynthesis (most active in the oxic B1A mat) as well as those of fatty acid metabolism and lipopolysaccharide biosynthesis (most active in suboxic mats) may be related to differences in the population structure and metabolism of different cell membrane components. Interestingly, the results indicate that B1A and B1B populations potentially metabolise amino acids in a preferential way compared with the B2 population. A similar scenario has been suggested for communities characterised by low (but not high) pH (Belnap et al., 2011). However, the data generated herein indicate that the aromatic amino acids, histidine and tyrosine, are the best sources of carbon and energy in the uppermost population, whereas linear aromatic amino acids, such as valine, leucine and isoleucine, are barely utilised, in contrast to the lowermost populations. In addition, the metabolic analysis confirmed that none of the amino-acid pathways were particularly enriched in B2. As folate derivatives are known to provide the majority of one-carbon units to the cells and their biosynthesis was improved in B1B, we suggest that the relative contribution of the carbon pool for synthetic purposes is also stratified; a similar observation accounts for the synthesis of cofactors such as nicotinate and nicotinamide precursors, which are more produced in B1A and B1B than in B2.
Furthermore, three hydrogenases, one rusticyanin and three cbb3-type cytochrome c oxidase subunits relevant for H2 and Fe metabolism were found among the expressed proteins (Supplementary Table S6). Within the three hydrogenases (one common, one in B1A and one in B1B, with ambiguous taxonomic affiliations), the expression levels were 32- and 8.4-fold higher in B1A than in B1B and B2, respectively, which is in agreement with the genomic evidence indicating that this community was most efficient for hydrogenoclastic activity. Secondly, among the one rusticyanin (binned to phylum Proteobacteria, most likely of the Acidithiobacillales) and three cbb3-type cytochrome c oxidases (two binned to phylum Proteobacteria, most likely being members of Gallionellales, and one to Actinobacteria), three were observed at high concentrations in B1A (up to 1.1 mmol mol−1 protein), and only one (binned to Actinobacteria) was observed in B1B (1.4 mol mol−1) and B2 (0.41 mmol mol−1). These results again confirm that the oxic mat population is more active for Fe metabolism.
Conclusions
AMD environments generally host a finely tuned restricted set of acidophiles, with a phylotype richness that depends highly on the pH condition (Sánchez-Andrea et al., 2011; Kuang et al., 2013). The results of this study draw attention to the huge undiscovered and non-described pool of microorganisms populating acidic environments and having important roles in iron/carbon/hydrogen cycles, particularly in low pH suboxic macroscopic growths. In particular, by integrating biodiversity and community genomic and proteomic profiling, we determined that the Los Rueldos AMD solution was characterised by unusually low temperature and redox potential (Supplementary Table S1), which may contribute to sustaining a wide microbial population compared with similar niche boundaries. It included a high proportion of uncommon Bacteria known to inhabit anoxic subsurfaces, uncultured Euryarchaeota, including methanogens, and ‘ARMAN’-related archaea. Our study further revealed that Los Rueldos is characterised by a stratified metabolically active ecosystem. In addition, clear shifts in the carbohydrate turnover have been observed in the stratified streamer (for details, see Supplementary Results and discussion and Supplementary Figure S11). Because a homogeneous pH of ∼2 characterised all samples, the provided evidence suggests that O2 concentrations, spatial localisation and biofilm architecture are the main shaping factors, providing a much higher level of microbial and functional shifts within single AMD sites than previously thought. The body of the results presented here has enabled us to pinpoint the major active metabolic functions and has provided us with invaluable insights into the global differences of macroscopic growths at the local scale.
One of the most crucial issues in the environment studied is the origin and further development of the streamer-mat structures. Figure 8 summarises a metabolic scheme that may provide a plausible explanation. Briefly, we hypothesise that the streamer starts as a chemolithotrophy-based community (most likely relying on Fe(II) oxidation, as suggested by the SSU tag pyrosequencing taxonomic analysis conducted over early developmental stages). This community would mainly consist of members capable of exopolysaccharide (EPS) production, allowing them access to the mineral resources on the bank side. The presence of small discrete slime growths in contact with mineral rocks at the bank was observed (data not shown). Later, a heterotrophic system would evolve, in which chemolithotrophy based on both Fe(II) and reduced inorganic sulphur compounds oxidation exists but not as the main energy metabolism. Mature communities would be able to fix both inorganic nitrogen and carbon dioxide and maintain a self-contained microbial subsystem in the overall Los Rueldos metabolism. Nitrogen and carbon would enter the community through the B1A layer, where metaproteogenomic signatures have been found for both nitrogenases and RuBisCO enzymes. The chemolithotrophic growth would thus be supported by non-described Nitrospirae (e.g., ‘L. ferrodiazotrophum’), Acidithiobacillales (At. ferrivorans, ‘RCP1-48’) and β-Proteobacteria (e.g., ‘Ferrovum myxofaciens’-like) members with EPS production capabilities. The products of degradation would serve as a mainstay for the heterotrophic metabolism in B1A and would diffuse to the submerged B1B, where they could serve as the principal contributors for heterotrophic-based cell growth. Reduced inorganic sulphur compound oxidation has been demonstrated to occur through the detection of metaproteogenomic signatures in B1B, but would not represent the main metabolism within the community. The anoxic conditions favour the presence of fermentation-by products (H2 gas), which would diffuse to the overlaying stratum B1A and could be used as an additional electron donor. The implication of H2 in the B1A metabolism has been demonstrated by the presence of expressed hydrogenases. The origin of B1B remains unexplained; the stratification could appear as a consequence of the heterotrophy by-products originating in B1A, or such a stratum could possess an independent origin (its thickness suggests this assumption as being possible as well as the possibility of the origin of a EPS-based structure in an anoxic environment).
Scheme illustrating the presumably predominant metabolisms in the Los Rueldos microenvironments. Hypothetical initial steps of colonisation are indicated for the establishment of the B1 streamer as well as main metabolic transformations that give origin to auto- and heterotrophic metabolisms. AMD flow is indicated by the arrows at the right side of each.
A relevant observation in the present study is the existence of a B2 macroscropic, EPS-based submerged growth, which is uncommon in nature. It is plausible that the submerged B2 originated via a reduced inorganic sulphur compound oxidation-based metabolism and/or a heterotrophic metabolism fuelled by products originating in the B1-like formation that extends at the end of the gallery and from the point where the AMD begins to flow (Figure 8). The flow could bear ammonium, DOC (dissolved organic carbon) and Fe(III) to the B2 emplacement, where the slow AMD outflow could favour the maintenance of those components in the area. Various taxonomic groups present in B2 have been reported to be capable of EPS production (‘F. myxofaciens’-related β-Proteobacteria and Acidithiobacillales such as A. ferrooxidans) and also of supporting heterotrophic metabolism. Nevertheless, bacteria with nitrogen and carbon fixation potential (L. ferrooxidans and A. ferrooxidans) have been demonstrated to exist in B2, although the ability of such gases to access the submerged mat and reach the cell membranes remains unclear. In addition, the transformation of nitrate to nitrite has been detected but is not driven by typical nitrifiers.
Finally, the existence of such diverse communities as those existing in Los Rueldos supports the existence of a heterotrophy-dominated system. In accordance with this finding, the observation that the various identified taxa (e.g., Xanthomonadaceae) have not been reported to exist in an autotrophy-dependent manner should be highlighted.
The results presented here should be seen as those of an explorative study, as there are several limitations that prevented us from drawing definitive conclusions. First, we are aware that these results have been established with a limited coverage of gene sequences (from 10.8 to 14.9 Mbp) and proteins with detected expression level (1589). Consequently, it is unknown to what degree a larger set of data might influence the metabolic distributions within the communities examined; however, we should stress the fact that the community structure was extensively characterised by full-length 16S and SSU rRNA hypervariable tag sequencing and that good agreement between the gene content, protein expression profile and community composition was found in many of the most representative metabolisms. Second, we are aware that the results presented here have been established from a single AMD formation geochemically distinct to reported ones, and therefore this might influence the comparability within Los Rueldos communities to those found in other AMD sites around the World. Whatever the case, the results presented here suggested that the key environmental factor that appears to be shaping the communities inside the gallery is the DO concentration, very low at certain depth levels due to the absence of drainage flowing. This, together with the low temperature and low redox potential in the pool, may be decisive for the observed microbial species richness within Los Rueldos macroscopic growths, as less extremely pH and temperature values are commonly associated with increased richness and diversity in AMD systems. Low temperature AMD systems (Cae Coch, Mynydd Parys, Los Rueldos) appear to be favourable to the establishment of ‘F. myxofaciens’-related bacteria that develop predominantly in the air–solution interface and may contribute importantly to biofilm formation. In contrast to what has been described for the Cae Coch (Johnson, 2012) or Mynydd Parys (Hallberg et al., 2006), the stagnant pool from Los Rueldos would presumptively allow the establishment of a stratified and more complex microbial system. The more extreme conditions inside the Iron Mountain (Tyson et al., 2004) would restrict the genomic diversity in the existing stagnant pools or slow down the development of a stratified and mainly heterotrophic microbial growth. The stagnancy favours the creation of a system based on carbon mobilisation, where Archaea gain importance in the recycling, and also in the production of carbon compounds. The polymeric matrix where microbes appear embedded, allowing Los Rueldos streamer to be ∼15 cm thick, may acts as a safety shield against the prevailing extreme condition, the pH ∼2, and also as a massive carbon source, enabling other taxonomic groups, non-typical acidophiles, to develop more easily.
Accession codes
References
Aguilera A, Manrubia SC, Gómez F, Rodríguez N, Amils R . (2006). Eukaryotic community distribution and its relationship to water physicochemical parameters in an extreme acidic environment, Rio Tinto (southwestern Spain). Appl Environ Microbiol 72: 5325–5330.
Amaral-Zettler LA, Zettler ER, Theroux SM, Palacios C, Aguilera A, Amils R . (2011). Microbial community structure across the tree of life in the extreme Río Tinto. ISME J 5: 42–50.
Amils R, González-Toril E, Aguilera A, Rodríguez N, Fernández-Remolar D, Gómez F et al. (2011). From río tinto to Mars: the terrestrial and extraterrestrial ecology of acidophiles. Adv Appl Microbiol 77: 41–70.
Baker BJ, Banfield JF . (2003). Microbial communities in acid mine drainage. FEMS Microbiol Ecol 44: 139–152.
Baker BJ, Comolli LR, Dick GJ, Hauser LJ, Hyatt D, Dill BD et al. (2010). Enigmatic, ultrasmall, uncultivated Archaea. Proc Natl Acad Sci USA 107: 8806–8811.
Baker BJ, Tyson GW, Goosherst L, Banfield JF . (2009). Insights into the diversity of eukaryotes in acid mine drainage biofilm communities. Appl Environ Microbiol 75: 2192–2199.
Baker BJ, Tyson GW, Webb RI, Flanagan J, Hugenholtz P, Allen EE et al. (2006). Lineages of acidophilic archaea revealed by community genomic analysis. Science 314: 1933–1935.
Baker-Austin C, Potrykus J, Wexler M, Bond PL, Dopson M . (2010). Biofilm development in the extremely acidophilic archaeon 'Ferroplasma acidarmanus' Fer1. Extremophiles 14: 485–491.
Belnap CP, Pan C, Denef VJ, Samatova NF, Hettich RL, Banfield JF . (2011). Quantitative proteomic analyses of the response of acidophilic microbial communities to different pH conditions. ISME J 5: 1152–1161.
Belnap CP, Pan C, VerBerkmoes NC, Power ME, Samatova NF, Carver RL et al. (2010). Cultivation and quantitative proteomic analyses of acidophilic microbial communities. ISME J 4: 520–530.
Bertin PN, Heinrich-Salmeron A, Pelletier E, Goulhen-Chollet F, Arsène-Ploetze F, Gallien S et al. (2011). Metabolic diversity among main microorganisms inside an arsenic-rich ecosystem revealed by meta- and proteo-genomics. ISME J 5: 1735–1747.
Bohorquez LC, Delgado-Serrano L, López G, Osorio-Forero C, Klepac-Ceraj V, Kolter R et al. (2012). In-depth characterization via complementing culture-independent approaches of the microbial community in an acidic hot spring of the Colombian Andes. Microb Ecol 63: 103–115.
Bond PL, Druschel GK, Banfield JF . (2000). Comparison of acid mine drainage microbial communities in physically and geochemically distinct ecosystems. Appl Environ Microbiol 66: 4962–4971.
Bond PL, Smriga SP, Banfield JF . (2000). Phylogeny of microorganisms populating a thick, subaerial, predominantly lithotrophic biofilm at an extreme acid mine drainage site. Appl Environ Microbiol 66: 3842–3849.
Comolli LR, Baker BJ, Downing KH, Siegerist CE, Banfield JF . (2009). Three-dimensional analysis of the structure and ecology of a novel, ultra-small archaeon. ISME J 3: 159–167.
Denef VJ, Kalnejais LH, Mueller RS, Wilmes P, Baker BJ, Thomas BC et al. (2010). Proteogenomic basis for ecological divergence of closely related bacteria in natural acidophilic microbial communities. Proc Natl Acad Sci USA 107: 2383–2390.
Dojka MA, Harris JK, Pace NR . (2000). Expanding the known diversity and environmental distribution of an uncultured phylogenetic division of bacteria. Appl Environ Microbiol 66: 1617–1621.
García-Moyano A, González-Toril E, Aguilera A, Amils R . (2007). Prokaryotic community composition and ecology of floating macroscopic filaments from an extreme acidic environment, Rio Tinto (SW, Spain). Syst Appl Microbiol 30: 601–614.
Golyshina OV . (2011). Environmental, biogeographic, and biochemical patterns of archaea of the family Ferroplasmaceae. Appl Environ Microbiol 77: 5071–5078.
González-Toril E, Aguilera A, Souza-Egipsy V, López Pamo E, Sánchez España J, Amils R . (2011). Geomicrobiology of La Zarza-Perrunal acid mine effluent (Iberian Pyritic Belt, Spain). Appl Environ Microbiol 77: 2685–2694.
Hallberg KB . (2010). New perspectives in acid mine drainage microbiology. Hydrometallurgy 104: 448–453.
Hallberg KB, Coupland K, Kimura S, Johnson DB . (2006). Macroscopic streamer growths in acidic, metal-rich mine waters in north wales consist of novel and remarkably simple bacterial communities. Appl Environ Microbiol 72: 2022–2030.
He Z, Xiao S, Xie X, Hu Y . (2008). Microbial diversity in acid mineral bioleaching systems of Dongxiang copper mine and Yinshan lead-zinc mine. Extremophiles 12: 225–234.
Hedlund BP, Cho J, Derrien M, Costa KC . (2011). Phylum XXII. Lentisphaerae. Bergey’s Manual Syst Bacteriol 4: 785–788.
Hernández E, Bargiela R, Suárez Diez M, Friedrichs A, Pérez-Cobas AE, Gosalbes MJ et al. (2013). Functional consequences of microbial shifts in the human gastrointestinal tract linked to antibiotic treatment and obesity. Gut Microbes 4: 306–315.
Iino T, Tamaki H, Tamazawa S, Ueno Y, Ohkuma M, Suzuki K et al. (2013). Candidatus Methanogranum caenicola: a novel methanogen from the anaerobic digested sludge, and proposal of Methanomassiliicoccaceae fam. Nov. and Methanomassiliicoccales ord. nov., for a methanogenic lineage of the class Thermoplasmata. Microbes Environ 28: 244–250.
Johnson DB . (2012). Geomicrobiology of extremely acidic subsurface environments. FEMS Microbiol Ecol 81: 2–12.
Jones DS, Albrecht HL, Dawson KS, Schaperdoth I, Freeman KH, Pi Y et al. (2012). Community genomic analysis of an extremely acidophilic sulfur-oxidizing biofilm. ISME J 6: 158–170.
Juottonen H, Tuittila ES, Juutinen S, Fritze H, Yrjala K . (2008). Seasonality of rDNA- and rRNA-derived archaeal communities and methanogenic potential in a boreal mire. ISME J 2: 1157–1168.
Kimura H, Sugihara M, Yamamoto H, Patel BK, Kato K, Hanada S . (2005). Microbial community in a geothermal aquifer associated with the subsurface of the Great Artesian Basin, Australia. Extremophiles 9: 407–414.
Kuang JL, Huang LN, Chen LX, Hua ZS, Li SJ, Hu M et al. (2013). Contemporary environmental variation determines microbial diversity patterns in acid mine drainage. ISME J 7: 1038–1050.
Liljeqvist M, Rzhepishevska OI, Dopson M . (2013). Gene identification and substrate regulation provide insights into sulfur accumulation during bioleaching with the psychrotolerant acidophile Acidithiobacillus ferrivorans. Appl Environ Microbiol 79: 951–957.
Liljeqvist M, Valdes J, Holmes DS, Dopson M . (2011). Draft genome of the psychrotolerant acidophile Acidithiobacillus ferrivorans SS3. J Bacteriol 193: 4304–4305.
López-Archilla AI, Gerard E, Moreira D, López-García P . (2004). Macrofilamentous microbial communities in the metal-rich and acidic River Tinto, Spain. FEMS Microbiol Lett 235: 221–228.
López-López A, Yarza P, Richter M, Suárez-Suárez A, Antón J, Niemann H et al. (2010). Extremely halophilic microbial communities in anaerobic sediments from a solar saltern. Environ Microbiol Rep 2: 258–271.
Loredo J, Alvarez R, Ordonez A . (2005). Release of toxic metals and metalloids from Los Rueldos mercury mine (Asturias, Spain). Sci Total Environ 340: 247–260.
Macalady JL, Jones DS, Lyon EH . (2007). Extremely acidic, pendulous cave wall biofilms from the Frasassi cave system, Italy. Environ Microbiol 9: 1402–1414.
Moreno-Paz M, Gómez MJ, Arcas A, Parro V . (2010). Environmental transcriptome analysis reveals physiological differences between biofilm and planktonic modes of life of the iron oxidizing bacteria Leptospirillum spp. in their natural microbial community. BMC Genom 11: 404.
Mueller RS, Dill BD, Pan C, Belnap CP, Thomas BC, VerBerkmoes NC et al. (2011). Proteome changes in the initial bacterial colonist during ecological succession in an acid mine drainage biofilm community. Environ Microbiol 13: 2279–2292.
Muller J, Szklarczyk D, Julien P, Letunic I, Roth A, Kuhn M et al. (2010). eggNOG v2.0: extending the evolutionary genealogy of genes with enhanced non-supervised orthologous groups, species and functional annotations. Nucleic Acids Res 38: D190–D195.
Nancucheo I, Johnson DB . (2010). Production of glycolic acid by chemolithotrophic iron- and sulfur-oxidizing bacteria and its role in delineating and sustaining acidophilic sulfide mineral-oxidizing consortia. Appl Environ Microbiol 76: 461–467.
Paul K, Nonoh JO, Mikulski L, Brune A . (2012). ‘Methanoplasmatales,’ Thermoplasmatales-related archaea in termite guts and other environments, are the seventh order of methanogens. Appl Environ Microbiol 78: 8245–8253.
Pruesse E, Peplies J, Glockner FO . (2012). SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28: 1823–1829.
Quatrini R, Appia-Ayme C, Denis Y, Jedlicki E, Holmes DS, Bonnefoy V . (2009). Extending the models for iron and sulfur oxidation in the extreme acidophile Acidithiobacillus ferrooxidans. BMC Genom 10: 394.
Ram RJ, Verberkmoes NC, Thelen MP, Tyson GW, Baker BJ, Blake RC 2nd et al. (2005). Community proteomics of a natural microbial biofilm. Science 308: 1915–1920.
Rawlings DE . (2002). Heavy metal mining using microbes. Annu Rev Microbiol 56: 65–91.
Rowe OF, Sanchez-Espana J, Hallberg KB, Johnson DB . (2007). Microbial communities and geochemical dynamics in an extremely acidic, metal-rich stream at an abandoned sulfide mine (Huelva, Spain) underpinned by two functional primary production systems. Environ Microbiol 9: 1761–1771.
Sakai S, Ehara M, Tseng IC, Yamaguchi T, Bräuer SL, Cadillo-Quiroz H et al. (2012). Methanolinea mesophila sp. nov., a hydrogenotrophic methanogen isolated from rice field soil, and proposal of the archaeal family Methanoregulaceae fam. nov. within the order Methanomicrobiales. Int J Syst Evol Microbiol 62: 1389–1395.
Sánchez-Andrea I, Rodriguez N, Amils R, Sanz JL . (2011). Microbial diversity in anaerobic sediments at Río Tinto, a naturally acidic environment with a high heavy metal content. Appl Environ Microbiol 77: 6085–6093.
Sunna A, Bergquist PL . (2003). A gene encoding a novel extremely thermostable 1,4-beta-xylanase isolated directly from an environmental DNA sample. Extremophiles 7: 63–70.
Tyson GW, Chapman J, Hugenholtz P, Allen EE, Ram RJ, Richardson PM et al. (2004). Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature 428: 37–43.
Tyson GW, Lo I, Baker BJ, Allen EE, Hugenholtz P, Banfield JF . (2005). Genome-directed isolation of the key nitrogen fixer Leptospirillum ferrodiazotrophum sp. nov. from an acidophilic microbial community. Appl Environ Microbiol 71: 6319–6324.
Valdés J, Pedroso I, Quatrini R, Dodson RJ, Tettelin H, Blake R 2nd et al. (2008). Acidithiobacillus ferrooxidans metabolism: from genome sequence to industrial applications. BMC Genom 9: 597.
Wilmes P, Remis JP, Hwang M, Auer M, Thelen MP, Banfield JF . (2009). Natural acidophilic biofilm communities reflect distinct organismal and functional organization. ISME J 3: 266–270.
Yarza P, Richter M, Peplies J, Euzéby J, Amann R, Schleifer K-H et al. (2008). The All-Species Living Tree Project: a 16S rDNA-based phylogenetic tree of all sequenced type strains. Syst Appl Microbiol 31: 241–250.
Yin H, Cao L, Xie M, Chen Q, Qiu G, Zhou J et al. (2008). Bacterial diversity based on 16S rRNA and gyrB genes at Yinshan mine, China. Syst Appl Microbiol 31: 302–311.
Zhou J, Xia B, Huang H, Treves DS, Hauser LJ, Mural RJ et al. (2003). Bacterial phylogenetic diversity and a novel candidate division of two humid region, sandy surface soils. Soil Biol Biochem 35: 915–924.
Ziegler S, Ackermann S, Majzlan J, Gescher J . (2009). Matrix composition and community structure analysis of a novel bacterial pyrite leaching community. Environ Microbiol 11: 2329–2338.
Ziegler S, Dolch K, Geiger K, Krause S, Asskamp M, Eusterhues K . (2013). Oxygen-dependent niche formation of a pyrite-dependent acidophilic consortium built by archaea and bacteria. ISME J 7: 1725–1737.
Acknowledgements
We gratefully acknowledge the financial support provided by the Spanish Ministry of Economy and Competitiveness (Projects CSD2007-00005, BIO2010-16303 and BIO2011-25012) and the European Regional Development Fund (ERDF). Partial support by the FP7-KBBE program through the projects Microme (Grant No. 222886-2) and MAGICPAH (FP7-KBBE-2009-245226) is also gratefully acknowledged. Proteomics analysis performed in the ONJ laboratory was supported by a research grant (FNU 09-072287) and an EliteForsk award to ONJ from the Danish Council for Independent Research. CMG thanks the Spanish Ministry of Economy and Competitiveness for a PhD fellowship. We thank Professor Antonio M Gil Serrano, Departamento de Química Orgánica, Universidad de Sevilla (Spain) for the analysis of sugar composition of B1 and B2 macroscopic growths. We also thank Professor Peter Golyshin and Dr Michail Yakimov for their excellent support in relation to the critical revision of the manuscript and the analysis of metabolic capacities, respectively.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Méndez-García, C., Mesa, V., Sprenger, R. et al. Microbial stratification in low pH oxic and suboxic macroscopic growths along an acid mine drainage. ISME J 8, 1259–1274 (2014). https://doi.org/10.1038/ismej.2013.242
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.242
Keywords
This article is cited by
-
Temporal patterns of bacterial communities in the Billings Reservoir system
Scientific Reports (2024)
-
Evolutionary patterns of archaea predominant in acidic environment
Environmental Microbiome (2023)
-
Genome-resolved metagenomics inferred novel insights into the microbial community, metabolic pathways, and biomining potential of Malanjkhand acidic copper mine tailings
Environmental Science and Pollution Research (2023)
-
Enhanced terrestrial Fe(II) mobilization identified through a novel mechanism of microbially driven cave formation in Fe(III)-rich rocks
Scientific Reports (2022)
-
Microbial diversity in extreme environments
Nature Reviews Microbiology (2022)