Abstract
Time-series are critical to understanding long-term natural variability in the oceans. Bacterial communities in the euphotic zone were investigated for over a decade at the San Pedro Ocean Time-series station (SPOT) off southern California. Community composition was assessed by Automated Ribosomal Intergenic Spacer Analysis (ARISA) and coupled with measurements of oceanographic parameters for the surface ocean (0–5 m) and deep chlorophyll maximum (DCM, average depth ∼30 m). SAR11 and cyanobacterial ecotypes comprised typically more than one-third of the measured community; diversity within both was temporally variable, although a few operational taxonomic units (OTUs) were consistently more abundant. Persistent OTUs, mostly Alphaproteobacteria (SAR11 clade), Actinobacteria and Flavobacteria, tended to be abundant, in contrast to many rarer yet intermittent and ephemeral OTUs. Association networks revealed potential niches for key OTUs from SAR11, cyanobacteria, SAR86 and other common clades on the basis of robust correlations. Resilience was evident by the average communities drifting only slightly as years passed. Average Bray-Curtis similarity between any pair of dates was ∼40%, with a slight decrease over the decade and obvious near-surface seasonality; communities 8–10 years apart were slightly more different than those 1–4 years apart with the highest rate of change at 0–5 m between communities <4 years apart. The surface exhibited more pronounced seasonality than the DCM. Inter-depth Bray-Curtis similarities repeatedly decreased as the water column stratified each summer. Environmental factors were better predictors of shifts in community composition than months or elapsed time alone; yet, the best predictor was community composition at the other depth (that is, 0–5 m versus DCM).
Similar content being viewed by others
Introduction
Investigations into temporal dynamics of marine microbial communities have revealed remarkable similarities and dissimilarities between ocean basins and have provided insight into the complex ecology of microbes (as reviewed in Ducklow et al. (2009); Fuhrman (2009); Giovannoni and Vergin (2012)). A holistic understanding of microbes in the ocean requires knowledge of the following: which microbes are present, when they occur, how much they contribute to the community and what environmental factors facilitate their distribution. Knowledge of each parameter will improve models of the microbial loop and microbial roles in the sea.
Seasonal and monthly patterns of variation have been observed using molecular methods at multiple aquatic time-series sites, which suggest that environmental change elicits a biological response; many, but not all, have also shown recurrence (Acinas et al., 1997; Li, 1998; Morris et al., 2005; Fuhrman et al., 2006; Alonso Sáez et al., 2007; Kan et al., 2007; Treusch et al., 2009; Campbell et al., 2011; Eiler et al., 2011; Gilbert et al., 2012; Robidart et al., 2012). For example, seasonality in Synechococcus ecotypes was observed in the Southern California Bight, primarily for clades I and IV (Tai and Palenik, 2009), and in the Chesapeake Bay (Cai et al., 2010). Distribution of Prochlorococcus ecotypes at the Hawaii Ocean Time-series (HOT) and Bermuda Atlantic Time-series Study (BATS) was similar except during annual deep water column mixing events at BATS (Morris et al., 2005; Treusch et al., 2009; Malmstrom et al., 2010; Eiler et al., 2011). SAR11 ecotypes were also dominant yet variable over time at BATS and HOT (Morris et al., 2002; Carlson et al., 2009; Eiler et al., 2009; Giovannoni and Vergin, 2012). Seasonal patterns were also seen using next-generation sequencing (for example, Andersson et al., 2010; Fortunato et al., 2012; Gilbert et al., 2012).
In this study, we examined bacterial community structure in light of traditional ecological metrics for community composition, membership, phylogeny, persistence and connectivity (recently reviewed in Shade and Handelsman (2012)). Prior research at the San Pedro Ocean Time-series (SPOT) has focused on determining the temporal variability of bacterial, archaeal and protistan taxa and the development of ecological networks that link these communities (Countway and Caron, 2006; Fuhrman et al., 2006; Beman et al., 2010; Countway et al., 2010; Beman et al., 2011; Steele et al., 2011). Here, we assessed inter-annual, seasonal and monthly variability and resilience of bacterial communities in the surface water and deep chlorophyll maximum (DCM), as revealed by Automated Ribosomal Intergenic Spacer Analysis (ARISA), over a full decade. ARISA allowed for repeated detection of the same microbes over time and their relative abundance within the bacterial community; OTUs (operational taxonomic units) were identified by their unique fragment lengths, and each was assigned an identity through the coupling of 16S-ITS sequences from SPOT and elsewhere to an observed (or predicted) ARISA fragment length. We discuss the roles of specific bacterial taxa using correlated partners from network analysis and as members of a newly defined core microbiome at SPOT, as well as the diversity within key bacterial groups, and temporal patterns in relative abundances of taxonomically related OTUs. We also determine which of the measured environmental parameters best explain the observed community structure.
Materials and methods
Sample Collection
Seawater was collected approximately monthly from August 2000 to January 2011 at 0–5 m and the DCM (average 28.2 m, range 7–45 m), as determined from in situ fluorescence, at SPOT (33°33′N, 118°24′W). After losses due to weather conditions or due to equipment failure, 103 months were sampled at 0–5 m and 89 months at DCM over 126 months. DNA was extracted by phenol-chloroform from ∼10 l of seawater serially filtered through a 142 mm Type A/E glass-fiber filter (Pall Life Sciences; Ann Arbor, MI, USA) and a 0.22 μm Durapore GVWP (Millipore, Billerica, MA, USA). Results shown here are from the 0.22 μm filters, representing free-living bacteria and some picoeukaryotes (the largest and attached bacteria and most eukaryotes were removed by the A/E). Bacterial production was measured in triplicate 10 ml seawater samples by [3H]thymidine and [3H]leucine incorporation with a conversion factor of 2 × 1018 cells mol−1 of thymidine and 1.5 × 1017 cells mol−1 of leucine (Fuhrman et al., 2006). Bacterial and viral abundances were enumerated by epifluorescence microscopy from 2 ml formalin-fixed samples (Noble and Fuhrman, 1998; Patel et al., 2007). Environmental parameters were determined using standard oceanographic methods (Brown et al., 2005; Fuhrman et al., 2006; Beman et al., 2010; Steele et al., 2011). Estimates for chlorophyll-a concentrations and primary production were downloaded for the grid area surrounding SPOT from NOAA Coastwatch: (a) SeaWiFS, 0.04167 degrees, West US science quality for chlorophyll-a; and (b) SeaWiFS and Pathfinder, 0.1 degrees, global, experimental data sets for primary productivity (Hooker and McClain, 2000).
Bacterial Community Fingerprinting
ARISA
Amplification for ARISA (Fisher and Triplett, 1999) was modified from that performed in the study by Brown et al (2005) as follows (final volume of 50 μl): 2 ng DNA, 1 × buffer, 2.5 mM MgCl2, 0.2 mM each DNTP, 0.2 mg ml−1 BSA (Sigma-Aldrich, St Louis, MO, USA; A7030), 0.8 μM each of universal 16S primer (1392F: (5′-[C/T]ACACACCGCCCGT-3′)) and bacterial 23S primer (125R: 5′-[TET]GGGTT[C/G/T]CCCCATTC(A/G)G-3′)), and 5 units AmpliTaq Gold (Life Technologies, Grand Island, NY, USA). PCR was initiated with 10 min at 95 °C (hot-start), followed by 30 cycles of 95 °C for 40 s, 56 °C for 40 s and 72 °C for 90 s, with a final 7-min extension at 72 °C. All samples before 2008 were re-amplified for this study to improve fragment resolution from 1 bp to 0.1 bp; samples from 2008–2011 were analyzed as collected. PCR products were verified by agarose gel electrophoresis, concentrated to 10 μl with Zymo Research’s Clean & Concentrator-5 (Irvine, CA, USA) as per the manufacturer’s instructions and quantified by Picogreen (Life Technologies). Fragments, with a 0.1-bp (apparent size) resolution, were detected by slab gel electrophoresis (Chow and Fuhrman 2012; Needham et al., 2013); 10 ng μl−1 was loaded in duplicate on non-adjacent lanes. Fragments, with minimum 0.01% area, were dynamically binned with maximum bin sizes of 1 bp (390–450 bp), 2 bp (450–650 bp), 3 bp (650–900 bp) and 5 bp (900–1200 bp) (Fuhrman et al., 2006; Ruan et al., 2006b; Steele et al., 2011). Bins <0.1 bp were manually merged with the adjacent bin.
Peak Identification
Each ARISA bin was identified where possible using 16S-ITS sequences (Brown et al., 2005; Needham et al., 2013). Our identities were based on 16S-ITS ARISA clones from SPOT, the Atlantic river plume and the Pacific Ocean, which were analyzed similar to environmental samples to determine an observed length; Genbank accession numbers for new 16S-ITS clone sequences included in this study are: KF227401–KF227794. Calculated lengths were determined for additional published sequences from two linear regressions (400–800, 800–1200 bp) of observed lengths versus empirical lengths of base pair counts (Needham et al., 2013). Sequences were searched using BLAST (word size, 7) against Greengenes (McDonald et al., 2011), the Ribosomal Database Project (RDP, (Cole et al., 2009)) and the SILVA108 truncated SSU database (Pruesse et al., 2007) to construct a hybrid taxonomy using the top hit (by e-value with minimum alignment length 200 bp and percentage identity 97%). For each sequence, we determined the following: (1) explicit taxonomic ranks from Greengenes; (2) SILVA identifier from the lowest identified rank, excluding ‘uncultured’ or ‘unidentified’; (3) SAR11 clades from RDP release 10; and (4) cyanobacteria ecotype from ITS sequences, designation of known isolates (Rocap et al., 2002; Brown and Fuhrman 2005) or phylogenetic placement of 16S rDNA sequences.
Identities were assigned by matching environmental lengths to 16S-ITS sequences, with priority to SPOT and/or surface waters, in the following order: (1) observed ARISA length of SPOT clones from 5 m across the seasons (see Brown et al., 2005); (2) published cyanobacterial ITS sequences as noted above; (3) in silico amplification of marine isolate genomes (from megx.net EnvO-Lite annotations (Kottmann et al., 2009), photic zone); (4) observed ARISA lengths of 16S-ITS clones from the (a) central Pacific and (b) tropical Atlantic Oceans; (5) observed ARISA lengths from 16S-ITS clones from surface waters of the Indian Ocean: 0.8–3.0 μm and 3.0–20 μm size fractions (A. Allen, pers. comm); (6) observed ARISA length of SPOT clones from 150 m and 890 m; and (7) in silico amplification of marine isolate genomes originating below the euphotic zone. Dual identifications were retained if an ARISA OTU matched multiple sequences with divergent taxonomy; however, the top hit only was used if one sequence was more numerous than a second at 2:1 or higher. See Supplementary Table S4 for additional taxonomic information on ARISA OTUs presented in this study.
Data Analysis
Statistics
Community composition data were normalized by total peak area per month for each depth to determine relative abundances (also referred to as percentage contributions). The top 100 OTUs, by average abundance, from each depth were assessed for seasonality by fitting a linear model, lm() function in R, with day length and rate of change for day length as independent variables and logit-transformed relative abundance data as dependent variables (adjustment factor of 0.001, car package). Only OTUs with P<0.05 were considered seasonal. The following analyses were completed in PRIMER-E v6 (Clarke 1993; Clarke and Gorley, 2006): (1) Bray-Curtis similarity for bacterial communities and Euclidean distance for environmental measurements; (2) SIMPER: to determine an individual OTU's contribution to the overall (dis)similarity between samples or within groups; (3) RELATE: to compare Bray-Curtis and/or Euclidean (dis)similarity matrices by a comparative nonparametric Mantel-type test (Spearman, 999 permutations); and (4) BIO-ENV: to identify which environmental factors best described variations in bacterial community structure (Spearman, 99 permutations). Environmental data were transformed as follows before analyses in PRIMER-E: log(value+0.01) for NO2, NO3, PO4 and P*, bacterial production by thymidine and leucine incorporation, calculated turnover time, chlorophyll-a (bottle and satellite); square-root for bacterial and viral abundance and the virus:bacteria ratio; no transformation for salinity, temperature, sea surface height differential, primary production (satellite), day length and monthly change in day length. Missing environmental data were filled with the overall mean of the transformed data. Discriminant function analyses (DFA) and time-series plots were calculated in Systat11 using the 50 most important OTUs, as determined in PRIMER-E on the basis of relative abundance and occurrence. Global correlations and linear regressions of average Bray-Curtis similarities per month lag and environmental parameters (untransformed) were determined in Sigmaplot11 (San Jose, CA, USA).
Network Analysis
We used Local Similarity Analysis (eLSA) to identify intervals of correlation between OTUs and environmental parameters: analysis settings included a minimum occurrence of 5 months, normalization of variables by ‘percentileZ’ method, use of pmix (determined theoretical P-value followed by permutation testing (n=2000) for any P <0.1 to decrease computation time while maintaining accuracy) and linear interpolation of missing values (Ruan et al., 2006a; Xia et al., 2011, 2013). eLSA correlations with P <0.05 and q <0.10 were visualized in Cytoscape v2.8.2 (Shannon 2003; Smoot et al., 2011); q-values were calculated to determine a false-discovery rate (Storey 2002). Network statistics and comparisons were determined for undirected networks with Network Analyzer (Assenov et al., 2008) and Advanced Network Merge plugins, respectively. Random undirected networks of equal nodes and edges were constructed by the Erdős–Rényi model using the Random Network plugin (Steele et al., 2011). Modules were identified with the AllegroMCODE plugin using default settings (AllegroViva, Santa Clara, CA, USA).
Results and Discussion
Seasonal environmental variability in the euphotic zone at SPOT
SPOT displayed repeated seasonal stratification (Figure 1) as demonstrated by increased sea surface temperature from 14 °C in winter to 20 °C in late summer and a deepening of the average DCM depth in June (32 m) through September (33.5 m) while remaining at ∼14 °C. The average DCM depth was 28.2 m (range: 7–45 m). Compared with other ocean time-series locations, the mixed layer (and total water column) at SPOT is much shallower than that at HOT and BATS because of regional hydrography and closeness of the location to the shore. During winter mixing from December to March, the apparent mixed layer depth was equal to or slightly below the DCM and did not rise above the DCM until April. Thus, the seasonal position of the mixed layer depth relative to the DCM at SPOT lies between trends observed at HOT and BATS (as reviewed in Giovannoni and Vergin (2012)) – mixing was not as pronounced as at BATS, nor was stratification as pronounced as at HOT.
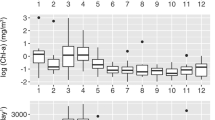
Environmental variability over 10 years in the euphotic zone at SPOT. Monthly averages are shown (X-axis=calendar month), where error bars indicate standard error of the mean, for the following: (a) prokaryotic cellular abundance, (b) viral abundance, (c) bacterial production by leucine (Leu) and thymidine (Tdr) incorporation, (d) chlorophyll-a concentration, (e) DCM sampling depth, (f) seawater temperature, (g) salinity, (h) phosphate, (i) nitrite and (j) nitrate. Filled symbols are 0–5 m; open symbols, DCM.
Regional upwelling near SPOT has been previously linked to local productivity and particle export (Collins et al., 2011). Chlorophyll-a concentrations were the highest in the surface ocean during winter and elevated in April and summer in the DCM (Figure 1d). Bacterial abundance and production peaked in April, whereas nutrient concentrations increased through spring before declining to a minimum by October (Figure 1). April bacterial production rates (between 1.3 × 105 to 5.5 × 105 cells ml−1 per day) were double to 10-fold higher than those during the rest of the year, with the lowest rates during winter. Bacterial abundance ranged from 1.2 × 106 to 3 × 106 cells ml−1, with annually repeated increases in spring. Viral abundance varied within an order of magnitude (2 × 107–4 × 107 particles per ml) with little to no pattern. This observation is in contrast to repeated seasonal increases in viral abundance at BATS (Parsons et al., 2012). In summary, onset of seasonal stratification coincided in late spring with high bacterial productivity and abundance – activity likely due to water column stabilization following increased nutrient availability from winter to spring mixing events.
Individual bacterial OTUs revealed seasonality and persistence
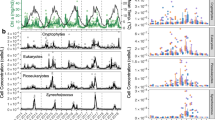
Some OTUs exhibited seasonal patterns (n=22 (0–5 m) and n=20 (DCM), nine common to both depths), whereas others were persistently abundant (Figure 2). Over ten years, 414 OTUs were observed over both depths (0–5 m: 407; DCM: 396). On average, 106±2 (±s.e.m.) OTUs were detected each month (range: 0–5 m, 54–174; DCM, 57–162). Average relative abundance of all OTUs was 0.9%, although the distribution was highly skewed. Seasonal OTUs collectively averaged 23.1%±1.1 (s.e.m.) and 13.8%±0.8 of the community (range: 0–5 m, 2.26–63.2%; DCM, 2.27–37.7%). Some OTUs peaked in fall (August–October, Figures 2a and c) or spring (March–May, Figures 2d and f) and others (for example, SAR11_686.9 and Flavobacteria NSb_726.4) peaked in summer (not shown).
Average monthly contribution of individual ARISA OTUs demonstrates seasonality (a-f) and persistence (g-i). Seasonal OTUs included those that peak in late summer and fall (a-c) and in spring (d-f). Note that one seasonal OTU is a chloroplast from a photosynthetic picoeukaryote (d). The three most abundant OTUs on average are observed consistently year round (g-i). Filled circles, 0–5 m; open circles, DCM. Error bars indicate the standard error of the mean. Y-axis denotes percentage of the total community as measured by ARISA.
The observed contributions of cyanobacteria and SAR11 were also temporally variable, consistent with previous studies (Morris et al., 2005; Kan et al., 2007; Carlson et al., 2009; Tai and Palenik, 2009; Cai et al., 2010; Malmstrom et al., 2010; Gilbert et al., 2012). Cyanobacteria collectively comprised 4.7% (0–5 m) and 2.2% (DCM) on average, and up to 31.8% (5 m) and 18.5% (DCM) (Supplementary Figure S1). In 0–5 m, the relative abundance of cyanobacterial OTUs increased as the DCM depth deepened in late summer to fall. High-light Prochlorococcus OTUs were the largest cyanobacterial contributors – one high-light Prochloroccus OTU from clade I, Pro_HL(I)_828.8, dominated the cyanobacteria at 0–5 m and shared dominance in the DCM with another high-light clade I ecotype, Pro_HL(I)_831.8. A low-light Prochlorococcus OTU (Pro_LL(I)_912.5) was a sporadically high contributor in the DCM, often in the latter half of 2003–2009. Synechococcus OTUs were present year round, but increased in spring following upwelling and times of higher productivity in contrast to reported decreases after upwelling events in Monterey Bay (Paerl et al., 2012). The high proportion of Prochlorococcus OTUs and limited presence of Synechococcus are consistent with other observations that SPOT is oligotrophic.
Cumulative SAR11 OTUs had high relative abundance – 35.7% (0–5 m) and 32.0% (DCM) on average and up to 66.6% (0–5 m) or 63.3% (DCM) in a single month (Supplementary Figure S2), comparable to the mean contribution of 38% in the photic zone at HOT (Eiler et al., 2009). SAR11 Surface Clade 1 OTUs (666.4, 670.5 and 686.9) and SAR11 662 (clade undetermined) were dominant; SAR11 Surface clade 4 OTU (703.7) occurred as a consistently minor contributor at <1% in 86 months and 72 months with a maximum of 7.3% and 3.8% in 0–5 m and DCM, respectively. Peaks in cumulative SAR11 relative abundances occurred in late summer similar to BATS (Carlson et al., 2009) and opposite to winter peaks at HOT and the Western English Channel (Eiler et al., 2009; Gilbert et al., 2012; Giovannoni and Vergin, 2012).
The five most abundant OTUs (Actinobacteria OCS155_435.5 and 4 SAR11 OTUs) and the remaining SAR11 and cyanobacterial OTUs together comprise ∼50% of the community (monthly average: 0–5 m, 50.4%; DCM, 47.4%) and up to 77% and 81% in a single month, respectively (Supplementary Figures S1-S3). Relative contributions of the top five OTUs at each depth were (1) significantly correlated between depths (P<0.05, Supplementary Table S1), (2) totaled ∼33% on average and (3) peaked cumulatively at 63.8% (0–5 m) and 76.7% (DCM) in a single month (Figures 2g and i, Supplementary Figure S3). These abundant OTUs were responsible for over 50% of the observed intra-depth similarity (by SIMPER: 52.1%, 0–5 m; 55.4%, DCM) and as such are key microbes of the euphotic zone.
Defining the microbial community by persistence and rarity
In general, more OTUs were rare and infrequent, yet cumulatively represented only a small fraction of the photic zone community. The bacterial community included persistent (>75% of months), intermittent (25–75%) and ephemeral (<25%) OTUs (Figure 3): 60% (0–5 m) and 58% (DCM) of OTUs were ephemeral, 33.4% (0–5 m) and 35% (DCM) were intermittent and only 6–7% were persistent. Average abundance was <1.2% for ephemeral and intermittent OTUs as compared with >10% for persistent OTUs. Persistent OTUs exhibited the highest average relative abundance, similar to the Western English Channel, Station ALOHA and a freshwater lake (Gilbert et al., 2009; Eiler et al., 2011; Caporaso et al., 2012; Eiler et al., 2012).
Defining the euphotic zone microbiome at SPOT. The number of OTUs (a, Y-axis) and the average contribution per OTU (b, Y-axis) are shown relative to the OTU’s percentage frequency (X-axis) for each depth. Percentage frequency was determined by dividing the number of months an OTU was observed by the total months sampled (n=103 (0–5 m), 89 (DCM)). Filled symbols, 0–5 m; open symbols, DCM. (c) A generalized depiction of the number of OTUs observed by percentage frequency category (Persistent, Intermittent and Ephemeral). Circles are drawn to scale according to their proportion of the community (average of 0–5 m and DCM). (d) Taxonomic summary at Class level of all OTUs by percentage frequency category.
The taxonomic distribution of OTUs as persistent, intermittent or ephemeral was consistent at the Class level between depths (Figure 3d); some Classes were more prone to persistence, whereas others were more fleeting over the 10 years we observed. Persistent OTUs included members of the Alphaproteobacteria, SAR406, Actinobacteria, Flavobacteria, chloroplasts, Deltaproteobacteria, Gammaproteobacteria and Synechococcophycideae (Cyanobacteria). Betaproteobacteria, Oscillatoriophycideae and Sphingobacteria were only observed as intermittent OTUs. Ephemeral OTUs also included Verrucomicrobiae and Chlorobia. All these taxonomic groups are known key factors of oceanic microbial communities over space and time (for example, Morris et al., 2002; Treusch et al., 2009; Zinger et al., 2011; Gilbert et al., 2012; Morris et al., 2012; Yilmaz et al., 2012). A smaller percentage of ephemeral OTUs (40%) were identified compared with persistent OTUs (96.5%), as expected, because clone libraries and sequence databases used for identification are more likely to include common organisms.
Ecological networks illustrate potential niches
Depth-specific association networks, constructed using eLSA from OTU co-occurrence patterns, uncovered complex interactions within bacterial communities. Correlations (mathematical interactions) between bacterial taxa were more numerous compared with environmental parameters, as seen previously at SPOT and other locations (Steele et al., 2011; Eiler et al., 2012; Gilbert et al., 2012). The resulting networks were highly interconnected, more so than by random chance alone (Supplementary Table S2). Clustering coefficients and the clustering coefficient ratio (Cl/Clr) were higher than observed in the previous 4-year DCM network and random networks of equal size; these values align with previously observed ratios from food webs and functional microbial networks (as summarized in Steele et al., 2011). The higher clustering coefficients, as compared with random networks, and minimal distance (the shortest path) between any two OTUs support our previous argument for small-world properties in microbial ecological networks (Watts and Strogatz 1998; Montoya et al., 2006; Steele et al., 2011), such that each OTU is closely linked to all other OTUs in highly clustered cliques.
We characterized potential ecological niches by identifying interconnected clusters (Figure 4) or connections centered on specific taxa (Figure 5). For example, one interconnected cluster from the 0–5 m network consisted of three bacterial clusters (Clusters I-III, Figure 4) linked to biological measurements (Cluster B). Cluster I was connected primarily through negative delayed correlations, Cluster II through positive correlations and Cluster III through positive and positive delayed correlations (see also Supplementary Tables S3 and S4). Cluster I appears to precede Clusters II and III, following the direction of time-delayed correlations (shown by arrows). OTUs in Cluster III were positively correlated with delays, such that this cluster may reflect a succession of OTUs. Roseobacter spp. have commonly been observed in areas or at times of high productivity (Buchan and Gonzalez, 2005; Morris et al., 2012); Roseobacter OTU 987.8 (Cluster II) may represent a similar observation here, as it was positively correlated with bacterial productivity (directly) and abundance (indirectly). Negative and delayed correlations between clusters suggest that each component within the larger network indicates a separate niche, each with its own set of ecological relationships.
Highly interconnected clusters of bacterial OTUs and environmental parameters reveal underlying niches in the surface ocean. Circles, ARISA OTUs; squares, biotic; hexagons, abiotic. Solid lines, positive LS; dashed lines, negative LS; arrow, 1-month delayed LS correlations that point toward the lagging OTU. Additional LSA correlation and node details are provided in Supplementary Tables S3 and S4.
Intersection network of LS correlations that occurred in both 0–5 m and DCM for the top five ARISA OTUs. Primary connections between the top five OTUs and others only are shown, excluding all secondary connections. Squares highlight the five most abundant OTUs; circles represent other ARISA OTUs as labeled. Only direction and delay are shown for LS correlations, as the values themselves differed between depths. Solid lines, positive LS; dashed lines, negative LS; arrow, 1-month delayed LS correlations that point toward the lagging OTU.
Despite the presence of almost all OTUs in each depth, the unique nature of each depth’s association network suggests that individual OTU–OTU relationships differ, on the basis of abundance of co-occurring microbes or environmental constraints. Coherent associations were determined from an intersection network of LS correlations observed in both depths. Both value (positive or negative) and direction (delayed or no delay) were considered when determining whether an interaction was unique or shared between depths. Only 12.4% (0–5 m) and 6.8% (DCM) of LS correlations were shared (Supplementary Table S2). The top five OTUs, however, did have consistent correlations with many other bacteria in both depths (Figure 5). Bacterial OTUs tended to be negatively correlated to SAR11. Actino_OCS155_435.5 was positively correlated to other Actinobacteria, SAR86 and SAR11 OTUs. Most correlated OTUs were uncommon, and some are thought to be relatively copiotrophic. The four most abundant SAR11 OTUs each clustered separately, implying that their preferred conditions may have different potential competitors or partners, consistent with the distribution of SAR11 ecotypes in the ocean (Brown et al., 2012). Future comparisons in phylogeny and relative abundances of OTUs included in this network with other long-term time-series sites may aid the definition of truly global interactions covering time and space.
Ten-year seasonal and annual trends in overall community composition
We previously identified predictable seasonal differences and annual recurrence of communities by discriminant function analysis (DFA) from a 4.5-year survey of the surface ocean (Fuhrman et al., 2006), and here our extended 10-year data set in both the surface and DCM depths displayed annual recurrence in 0–5 m only, despite seasonality in both depths (Supplementary Figure S4). DFA relies upon selecting individual OTUs to optimally distinguish months. Both communities were significantly positively autocorrelated at 1 month and negatively autocorrelated at 4–6 months (‘opposite’ seasons). Positive autocorrelation was statistically significant in 0–5 m at 10 ‘months’ (equivalent to 1 year due to missing data) but not significant in the DCM. Missing data for several months interspersed throughout the time series, and the significant gap in DCM observations (10/2006–2/2008) may limit our interpretation; restricting analysis to 8/2000–9/2006 in the DCM resulted in similar patterns (Supplementary Figure S4 C-F).
Seasonality and annual recurrence in the bacterial community structure were observed by long-term trends in Bray-Curtis similarity over time in 0–5 m, although this was not as apparent in the DCM despite an underlying seasonal trend by DFA (Figure 6). Bray-Curtis similarity is weighted by the relative abundance of each OTU and included all OTUs unlike DFA. Bacterial communities were on average 40.7±0.2% (Bray-Curtis similarity±s.e.m.) and 40.9±0.2% similar in 0–5 m and DCM, respectively. The highest average Bray-Curtis similarity occurred between communities 1 month apart, 50.7% (0–5 m) and 47.2% (DCM), consistent with positive autocorrelation by DFA at 1 month. Annual recurrence in 0–5 m was also demonstrated by identifying local maxima at yearly intervals (12, 24, 48, etc.; average 41.6%) and local minima for opposing seasons (6, 18, 30, etc.; average 38.4%); the difference in these two groups of averages was statistically significant (the Mann–Whitney U-test, P=0.005). Comparison of each depth’s Bray-Curtis similarity matrix with a distance matrix of elapsed time (by absolute number of days) was significantly correlated (RELATE: rho=0.156 (0–5 m), 0.123 (DCM), P=0.01), but to a lesser extent than to the Bray-Curtis similarity matrix of the other depth (rho=0.347, P=0.01). Correlation of the complete Bray-Curtis similarity matrices to a monthly distance model matrix (ignoring years) was low but statistically significant in 0–5 m (rho=0.067, P<0.05). Correlation for the DCM was not significant and may be due to the large fluctuations at >118-month time lags, perhaps stochastic (fewer data points) or due to an El Niño. Winter 2009–2010 was exceptionally warm, whereas winter 2010–2011 was exceptionally cool as compared with winter 2000–2001 (Oceanic Niño Index, NOAA). DFA and Bray-Curtis similarities suggest that adjacent months are more similar to each other compared with the same month from different years, but communities in opposite seasons repeatedly differed the most in the surface. These observations confirmed previously noted patterns on annual recurrence in bacterial communities at SPOT in the surface ocean (Fuhrman et al., 2006) and at BATS, ALOHA and the Western English Channel (Treusch et al., 2009; Eiler et al., 2011; Gilbert et al., 2012), with the last also exhibiting recurrence (Gilbert et al., 2012). The surface ocean thus exhibited more pronounced and predictable seasonal patterns compared with the DCM, perhaps due to the relative isolation of the DCM from direct atmospheric forcing, reduced annual temperature range or potential noise in the data due to the temporally variable depth of the DCM.
Seasonal and inter-annual patterns in Bray-Curtis community similarity. Average pairwise community similarity (Y-axis) was calculated from all OTUs for all months in (a) 0–5 m and (b) DCM. Time lag (X-axis) indicates the number of months between the communities compared. Linear regressions were calculated from average similarities for the following: (1) 0–5 m: from 1–48-month lags (A, black circles and solid line); (2) 0–5 m: 49–125-month lags (A, gray circles and dashed line); (3) 0–5 m: 1–125-month lags (A, all circles and no line shown); and (4) DCM: 1–118-month lags (B, black circles and solid line). Lags of 119–125 months in the DCM (B, white circles) were excluded.
Bray-Curtis comparisons also uncovered long-term trends: average similarity between all pairs of months changed relatively little irrespective of dates (ignoring seasonality in 0–5 m); yet there was a discernible decrease over longer time lags (Figure 6). In both depths, average similarity declined ∼0.5–0.6% per year over 10 years (equations 3 and 4, Figure 6). The decline is about 1% per year for communities <48 months apart in 0–5 m (equation 1), followed by almost no change in communities >48 months apart (equation 2). This slight decline with increasing lags does suggest that community composition overall had measurably changed over the course of this 10-year study.
Coupling Community Structure and Environmental Changes
Coherence of euphotic zone depths, assessed by Bray-Curtis similarities between co-occurring surface and DCM communities, was negatively related to the temperature difference between them (Figure 7). Temperature is an established predictor of variability in community structure on spatial scales (Pommier et al., 2007; Fuhrman et al., 2008; Yilmaz et al., 2012) and may be similarly predictive for spatiotemporal variation. Between-depth Bray-Curtis similarity was not correlated to bacterial production, bacterial or viral abundance, or nutrient concentrations from either depth. Our results thus suggest that the two depths were relatively homogenized during periods of winter mixing; as the euphotic zone warmed and stratified through the summer, bacterial communities diverged under different local conditions, only to be mixed together again the following winter – a pattern also observed at BATS (Carlson et al., 2009; Treusch et al., 2009).
Chlorophyll-a concentrations, bacterial production rates and nutrient concentrations correlated significantly with whole community shifts and for subsets of persistent, intermittent and ephemeral OTUs. Biological community variation was more related to measured environmental parameters (RELATE, 0–5 m: rho=0.221, P=0.01; DCM: rho=0.114, P=0.03) than to distance by time in elapsed days or months (rho<0.16, P<0.05). The environmental measurements that best explained the Bray-Curtis similarity matrices for all OTUs or persistent OTUs were identical at 0–5 m and included chlorophyll-a (satellite), bacterial production rates (leucine and thymidine), nitrate and day length change per month (BEST: all, rho=0.267, P=0.01; persistent, rho=0.257, P=0.01). Intermittent OTUs at 0–5 m correlated with the above parameters, except for nitrate, which was replaced by chlorophyll-a (bottle) or phosphate (rho=0.257, P=0.01); ephemeral OTUs correlated with nitrate, P* and chlorophyll-a (bottle) (rho=0.158, P=0.02). In the DCM, surface chlorophyll-a (satellite), bacterial production rates (thymidine only), calculated turnover time of leucine (bacterial production), phosphate and sea surface height differential by satellite provided the best fit for all OTUs (all: rho=0.268, P=0.01), but no statistically significant combination was observed for the DCM’s persistent OTUs. DCM intermittent OTUs correlated with surface chlorophyll-a (satellite), nitrite, bacterial production rates (thymidine), calculated turnover time (leucine) and day length change per month (rho=0.224, P=0.02); ephemeral OTUs correlated with calculated turnover time (leucine), salinity and sea surface height differential.
Network analysis revealed individual OTUs associated with key predictive environmental parameters – specifically for bacterial production rates and abundance (Figure 4) and for salinity, temperature and nutrient concentrations (Figure 8). ARISA OTUs – including SAR86, SAR11, Synechococcus and Actinobacteria (OCS155) – were positively correlated with few delays to salinity and temperature in the DCM. Most correlations between ARISA OTUs and salinity or temperature at 0–5 m were negative or delayed, suggesting that the OTUs shown were somewhat seasonal. Indeed, seasonality for several OTUs shown (that is, SAR86_402.4both. OCS155_419.5both, Owenwe_594.15 m, SAR324_5195 m, OTU_632.6DCM, SAR11_676.95 m, SAR11_682.45 m, SAR11_S2_716.85 m, SAR11_S2_718.4DCM, OTU_8085 m) was independently determined (Figures 2 and 8). Almost all OTUs included here, with a few SAR11 or Actinobacteria OTUs as exceptions, were intermittent or ephemeral, suggesting that these organisms are limited by nutrients or by other factors essential for growth (such as vitamins or trace metals).
Network analysis uncovered specific bacterial relationships in (a) 0–5 m and (b) DCM to environmental parameters such as salinity, temperature and nutrient concentrations. Only LS correlations >0.35 or <−0.35 for 0–5 m and >0.4 or <−0.4 for DCM are shown, resulting in networks of approximately equal size and complexity. Circles, ARISA OTUs; squares, biotic; hexagons, abiotic. Node size is proportional to an OTU’s percentage frequency in each depth; scale bars are indicated in the lower left (0–5 m) and right (DCM) corners. Solid lines, positive LS; dashed lines, negative LS; arrow, 1-month delayed LS correlations that point toward the lagging OTU. Additional node details are provided in Supplementary Table S4.
Methodological Considerations
We recognize that ARISA, like all PCR-based methods, has potential quantitative biases and other drawbacks and also has relative strengths compared with studies on short 16S rRNA sequences. Rare taxa are not easily identified by ARISA (Figure 3d), as identities are based primarily on clone libraries and genomic sequences. ARISA primers do miss a few known groups like Planctomycetes and SAR202. Occasionally (<15% of identified OTUs), unrelated taxa may be lumped into the same ARISA OTU, which adds noise to interpreting correlations and thus results in conservative conclusions. More importantly for phylogenetic resolution, the ITS region that leads to different observed ARISA lengths has been widely used to differentiate cyanobacterial (for example, Rocap et al., 2002; Brown and Fuhrman, 2005) and SAR11 ecotypes (Brown et al., 2012), which are difficult to resolve by 16S rRNA sequence alone. In silico analysis showed that our original ARISA protocol, since improved, had practical phylogenetic resolution similar to full-length 16S rRNA sequences at 99% similarity level (Brown and Fuhrman 2005). A direct comparison of ARISA clone libraries from SPOT (containing nearly full 16S, ITS and partial 23S rRNA sequences) found that SAR11 sequences from 38 clones resolved to six different SAR11 OTUs at 97% 16S sequence similarity versus 15 ARISA OTUs. Sixteen cyanobacteria clones could be resolved to two OTUs at 97 or 98% similarity versus seven ARISA OTUs. The impact of potential PCR biases (which can occur with both ARISA and tag sequences) was greatly reduced for the large majority of our quantitative analysis by comparing increases and decreases of individual OTUs – for example, as trends over time or as rank correlations between OTUs. ARISA was also previously shown to be an accurate estimator of Prochlorococcus spp. abundance as compared with counts by flow cytometry over time (Brown et al., 2005). Overall, ARISA, as we have applied it (with improved 0.1 bp fragment size, in duplicate, with equal DNA concentration at each step, and backed by >1000 16S-ITS clones from this location and throughout the ocean at multiple time points and depths), remains surprisingly appropriate for detecting fine-resolution microbial patterns especially for the moderately abundant to dominant organisms.
Conclusion
Notwithstanding seasonal variation, a relatively stable core microbial community persisted in both depths such that an average ∼40% pairwise date-to-date similarity was maintained over a decade (declining ∼0.6% per year). Results from both Bray-Curtis similarity and DFA suggest that the DCM community was less seasonal than 0–5 m, but monthly and long-term trends were apparent in both. Our previous observation of recurrence in the surface ocean remains true over a decade of measurements, despite a discernible long-term shift in community structure. Although community membership was consistent between the surface and DCM (as detected by ARISA), the interactions between their constituents differed. This observation may be due to the higher seasonal variation observed in 0–5 m from changing environmental conditions and due to separation from the DCM by seasonal stratification of the water column, such that microbe–microbe interactions are differentially influenced by environmental characteristics or by other biological controls. For example, the growth and activity of cyanobacteria, as well as the activity of phages and heterotrophic bacteria, have been shown to be dependent on light and nutrient conditions (Sher et al., 2011; Weinbauer et al., 2011). Our application of ARISA to this 10-year time series at SPOT illustrates that this method continues to be well suited for investigating complex ecological patterns especially for moderately abundant to dominant organisms and for addressing the effects of diversity within key bacterial groups like SAR11 or cyanobacteria on their ecological role in the ocean. Further exploration of microbe–microbe interactions under varied environmental conditions and co-occurring communities would aid in the interpretation of ecological or association networks developed from fingerprinting and sequencing data.
References
Acinas SG, Rodríguez Valera F, Pedrós-Alió C . (1997). Spatial and temporal variation in marine bacterioplankton diversity as shown by RFLP fingerprinting of PCR amplified 16S rDNA. FEMS Microbiol Ecol 24: 27–40.
Alonso Sáez L, Balagué V, Sà EL, Sánchez O, González JM, Pinhassi J et al (2007). Seasonality in bacterial diversity in north-west Mediterranean coastal waters: assessment through clone libraries, fingerprinting and FISH. FEMS Microbiol Ecol 60: 98–112.
Andersson AF, Riemann L, Bertilsson S . (2010). Pyrosequencing reveals contrasting seasonal dynamics of taxa within Baltic Sea bacterioplankton communities. ISME J 4: 171–181.
Assenov Y, Ramírez F, Schelhorn S, Lengauer T, Albrecht M . (2008). Computing topological parameters of biological networks. Bioinformatics 24: 282–284.
Beman JM, Sachdeva R, Fuhrman JA . (2010). Population ecology of nitrifying Archaea and Bacteria in the Southern California Bight. Environ Microbiol 12: 1282–1292.
Beman JM, Steele JA, Fuhrman JA . (2011). Co-occurrence patterns for abundant marine archaeal and bacterial lineages in the deep chlorophyll maximum of coastal California. ISME J 5: 1077–1085.
Brown MV, Fuhrman JA . (2005). Marine bacterial microdiversity as revealed by internal transcribed spacer analysis. Aquat Microb Ecol 41: 15–23.
Brown MV, Lauro FM, DeMaere MZ, Muir L, Wilkins D, Thomas T et al (2012). Global biogeography of SAR11 marine bacteria. Mol Syst Biol 8: 595.
Brown MV, Schwalbach MS, Hewson I, Fuhrman JA . (2005). Coupling 16S-ITS rDNA clone libraries and automated ribosomal intergenic spacer analysis to show marine microbial diversity: development and application to a time series. Environ Microbiol 7: 1466–1479.
Buchan A, Gonzalez J . (2005). Overview of the Marine Roseobacter Lineage. Appl Environ Microbiol 71: 5665–5677.
Cai H, Wang K, Huang S, Jiao N, Chen F . (2010). Distinct Patterns of Picocyanobacterial Communities in Winter and Summer in the Chesapeake Bay. Appl. Environ. Microbiol 76: 2955–2960.
Campbell BJ, Yu L, Heidelberg JF, Kirchman DL . (2011). Activity of abundant and rare bacteria in a coastal ocean. Proc Natl Acad Sci USA 108: 12776.
Caporaso JG, Paszkiewicz K, Field D, Knight R, Gilbert JA . (2012). The Western English Channel contains a persistent microbial seed bank. ISME J 6: 1089–1093.
Carlson CA, Morris R, Parsons R, Treusch AH, Giovannoni SJ, Vergin KL . (2009). Seasonal dynamics of SAR11 populations in the euphotic and mesopelagic zones of the northwestern Sargasso Sea. ISME J 3: 283–295.
Chow C-ET, Fuhrman JA . (2012). Seasonality and monthly dynamics of marine myovirus communities. Environ Microbiol 14: 2171–2183.
Clarke KR . (1993). Non-parametric multivariate analyses of changes in community structure. Aust J Ecol 18: 117–143.
Clarke KR, Gorley R . (2006) PRIMER v6: User Manual/Tutorial. PRIMER-E: Plymouth: UK.
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ et al (2009). The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37: D141–D145.
Collins LE, Berelson WM, Hammond DE, Knapp A, Schwartz R, Capone DG . (2011). Particle fluxes in San Pedro Basin, California A four-year record of sedimentation and physical forcing. Deep-Sea Res. (1 Oceanogr Res Pap) 58: 898–914.
Countway PD, Caron DA . (2006). Abundance and distribution of Ostreococcus sp. in the San Pedro Channel, California, as revealed by quantitative PCR. Appl Environ Microbiol 72: 2496–2506.
Countway PD, Vigil PD, Schnetzer A, Moorthi SD, Caron DA . (2010). Seasonal analysis of protistan community structure and diversity at the USC Microbial Observatory (San Pedro Channel, North Pacific Ocean). Limnol Oceanogr 55: 2381–2396.
Ducklow HW, Doney SC, Steinberg DK . (2009). Contributions of Long-Term Research and Time-Series Observations to Marine Ecology and Biogeochemistry. Annu Rev Marine Sci 1: 279–302.
Eiler A, Hayakawa DH, Church MJ, Karl DM, Rappé MS . (2009). Dynamics of the SAR11 bacterioplankton lineage in relation to environmental conditions in the oligotrophic North Pacific subtropical gyre. Environ Microbiol 11: 2291–2300.
Eiler A, Hayakawa DH, Rappé MS . (2011). Non-random assembly of bacterioplankton communities in the subtropical north pacific ocean. Front Microbio 2: 1–10.
Eiler A, Heinrich F, Bertilsson S . (2012). Coherent dynamics and association networks among lake bacterioplankton taxa. ISME J 6: 330–342.
Fisher MM, Triplett EW . (1999). Automated approach for ribosomal intergenic spacer analysis of microbial diversity and its application to freshwater bacterial communities. Appl Environ Microbiol 65: 4630–4636.
Fortunato CS, Herfort L, Zuber P, Baptista AM, Crump BC . (2012). Spatial variability overwhelms seasonal patterns in bacterioplankton communities across a river to ocean gradient. ISME J 6: 554–563.
Fuhrman JA . (2009). Microbial community structure and its functional implications. Nature 459: 193–199.
Fuhrman JA, Hewson I, Schwalbach MS, Steele JA, Brown MV, Naeem S . (2006). Annually reoccurring bacterial communities are predictable from ocean conditions. Proc Natl Acad Sci USA 103: 13104–13109.
Fuhrman JA, Steele JA, Hewson I, Schwalbach MS, Brown MV, Green JL et al (2008). A latitudinal diversity gradient in planktonic marine bacteria. Proc Natl Acad Sci USA 105: 7774–7778.
Gilbert JA, Field D, Swift P, Newbold L, Oliver A, Smyth T et al (2009). The seasonal structure of microbial communities in the Western English Channel. Environ Microbiol 11: 3132–3139.
Gilbert JA, Steele JA, Caporaso JG, Steinbrück L, Reeder J, Temperton B et al (2012). Defining seasonal marine microbial community dynamics. ISME J 6: 298–308.
Giovannoni SJ, Vergin KL . (2012). Seasonality in Ocean Microbial Communities. Science 335: 671–676.
Hooker SB, McClain CR . (2000). The calibration and validation of SeaWiFS data. Prog Oceanogr 45: 427–465.
Kan J, Suzuki MT, Wang K, Evans SE, Chen F . (2007). High Temporal but Low Spatial Heterogeneity of Bacterioplankton in the Chesapeake Bay. Appl Environ Microbiol 73: 6776–6789.
Kottmann R, Kostadinov I, Duhaime MB, Buttigieg PL, Yilmaz P, Hankeln W et al (2009). Megx.net: integrated database resource for marine ecological genomics. Nucleic Acids Res 38: D391–D395.
Li W . (1998). Annual average abundance of heterotrophic bacteria and Synechococcus in surface ocean waters. Limnol Oceanogr 43: 1746–1753.
Malmstrom RR, Coe A, Kettler GC, Martiny AC, Frias-Lopez J, Zinser ER et al (2010). Temporal dynamics of Prochlorococcus ecotypes in the Atlantic and Pacific oceans. ISME J 4: 1252–1264.
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A et al (2011). An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J 6: 610–618.
Montoya JM, Pimm SL, Solé RV . (2006). Ecological networks and their fragility. Nature 442: 259–264.
Morris RM, Frazar CD, Carlson CA . (2012). Basin-scale patterns in the abundance of SAR11 subclades, marine Actinobacteria (OM1), members of the Roseobacter clade and OCS116 in the South Atlantic. Environ Microbiol 14: 1133–1144.
Morris RM, Rappé MS, Connon SA, Vergin KL, Siebold WA, Carlson CA et al (2002). SAR 11 clade dominates ocean surface bacterioplankton communities. Nature 420: 806–810.
Morris RM, Vergin KL, Cho JC, Rappé MS, Carlson CA, Giovannoni SJ . (2005). Temporal and spatial response of bacterioplankton lineages to annual convective overturn at the Bermuda Atlantic Time-series Study site. Limnol. Oceanogr 50: 1687–1696.
Needham DM, Chow C-ET, Cram JA, Sachdeva R, Parada A, Fuhrman JA . (2013). Short-term observations of marine bacterial and viral communities: patterns, connections and resilience. ISME J 7: 1274–1285.
Noble RT, Fuhrman JA . (1998). Use of SYBR Green I for rapid epifluorescence counts of marine viruses and bacteria. Aquat. Microb. Ecol 14: 113–118.
Paerl R, Turk K, Beinart R, Chavez FP, Zehr J . (2012). Seasonal change in the abundance of Synechococcus and multiple distinct phylotypes in Monterey Bay determined by rbcL and narB quantitative PCR. Environ Microbiol 14: 580–593.
Parsons RJ, Breitbart M, Lomas MW, Carlson CA . (2012). Ocean time-series reveals recurring seasonal patterns of virioplankton dynamics in the northwestern Sargasso Sea. ISME J 6: 273–284.
Patel A, Noble RT, Steele JA, Schwalbach MS, Hewson I, Fuhrman JA . (2007). Virus and prokaryote enumeration from planktonic aquatic environments by epifluorescence microscopy with SYBR Green I. Nat Protoc 2: 269–276.
Pommier T, Canbäck B, Riemann L, Boström KH, Simu K, Lundberg P et al (2007). Global patterns of diversity and community structure in marine bacterioplankton. Mol Ecol 16: 867–880.
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J et al (2007). SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35: 7188–7196.
Robidart JC, Preston CM, Paerl RW, Turk KA, Mosier AC, Francis CA et al (2012). Seasonal Synechococcus and Thaumarchaeal population dynamics examined with high resolution with remote in situ instrumentation. ISME J 6: 513–523.
Rocap G, Distel DL, Waterbury JB, Chisholm SW . (2002). Resolution of Prochlorococcus and Synechococcus ecotypes by using 16S-23S ribosomal DNA internal transcribed spacer sequences. Appl Environ Microbiol 68: 1180–1191.
Ruan Q, Dutta D, Schwalbach MS, Steele JA, Fuhrman JA, Sun F . (2006a). Local similarity analysis reveals unique associations among marine bacterioplankton species and environmental factors. Bioinformatics 22: 2532–2538.
Ruan Q, Steele JA, Schwalbach MS, Fuhrman JA, Sun F . (2006b). A dynamic programming algorithm for binning microbial community profiles. Bioinformatics 22: 1508–1514.
Shade A, Handelsman J . (2012). Beyond the Venn diagram: the hunt for a core microbiome. Environ Microbiol 14: 4–12.
Shannon P . (2003). Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Research 13: 2498–2504.
Sher D, Thompson JW, Kashtan N, Croal L, Chisholm SW . (2011). Response of Prochlorococcus ecotypes to co-culture with diverse marine bacteria. ISME J 5: 1125–1132.
Smoot ME, Ono K, Ruscheinski J, Wang P-L, Ideker T . (2011). Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27: 431–432.
Steele JA, Countway PD, Xia L, Vigil PD, Beman JM, Kim DY et al (2011). Marine bacterial, archaeal and protistan association networks reveal ecological linkages. ISME J 5: 1414–1425.
Storey JD . (2002). A direct approach to false discovery rates. J Roy Stat Soc B Met 64: 479–498.
Tai V, Palenik B . (2009). Temporal variation of Synechococcus clades at a coastal Pacific Ocean monitoring site. ISME J 3: 903–915.
Treusch AH, Vergin KL, Finlay LA, Donatz MG, Burton RM, Carlson CA et al (2009). Seasonality and vertical structure of microbial communities in an ocean gyre. ISME J 3: 1148–1163.
Watts DJ, Strogatz SH . (1998). Collective dynamics of ‘small-world’ networks. Nature 393: 440–442.
Weinbauer MG, Bonilla-Findji O, Chan AM, Dolan JR, Short SM, Simek K et al (2011). Synechococcus growth in the ocean may depend on the lysis of heterotrophic bacteria. J Plankton Res 33: 1465–1476.
Xia LC, Ai D, Cram JA, Fuhrman JA, Sun F . (2013). Efficient Statistical Significance Approximation for Local Association Analysis of High-Throughput Time Series Data. Bioinformatics 29: 230–237.
Xia LC, Steele JA, Cram JA, Cardon ZG, Simmons SL, Vallino JJ et al (2011). Extended local similarity analysis (eLSA) of microbial community and other time series data with replicates. BMC Syst. Biol 5: S15.
Yilmaz P, Iversen MH, Hankeln W, Kottmann R, Quast C, Glöckner FO . (2012). Ecological structuring of bacterial and archaeal taxa in surface ocean waters. FEMS Microbiol Ecol 81: 373–385.
Zinger L, Amaral-Zettler LA, Fuhrman JA, Horner-Devine MC, Huse SM, Welch DM et al (2011). In: Global Patterns of Bacterial Beta-Diversity in Seafloor and Seawater Ecosystems. Gilbert. JA (ed.) PLoS One 6: e24570.
Acknowledgements
The authors acknowledge the efforts of the entire USC Microbial Observatory team, especially Troy Gunderson, Mark Brown, Ian Hewson, Mike Schwalbach, Mahira Kakajiwala, Sheila O’Brien, Tu My To, Henry Ho, Diane Kim, Adriane Jones, Pete Countway, Li Xia, Fengzhu Sun, David Caron and the captain and crews of R/V Seawatch and R/V Yellowfin. We thank Andrew Allen for generously providing Indian Ocean clones and 16S-ITS sequences. This work was supported by the NSF Graduate Research Fellowship Program (awarded to CET Chow, DM Needham, and AE Parada), and Microbial Observatory and Dimensions in Biodiversity Programs, grants 0703159 and 1136818.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Chow, CE., Sachdeva, R., Cram, J. et al. Temporal variability and coherence of euphotic zone bacterial communities over a decade in the Southern California Bight. ISME J 7, 2259–2273 (2013). https://doi.org/10.1038/ismej.2013.122
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.122
Keywords
This article is cited by
-
Seasonal bacterial niche structures and chemolithoautotrophic ecotypes in a North Atlantic fjord
Scientific Reports (2022)
-
Structure and membership of gut microbial communities in multiple fish cryptic species under potential migratory effects
Scientific Reports (2020)
-
Explore mediated co-varying dynamics in microbial community using integrated local similarity and liquid association analysis
BMC Genomics (2019)
-
Differential co-occurrence relationships shaping ecotype diversification within Thaumarchaeota populations in the coastal ocean water column
The ISME Journal (2019)
-
Temporal and spatial dynamics of Bacteria, Archaea and protists in equatorial coastal waters
Scientific Reports (2019)