Abstract
Microbial mats in sulfidic cave streams offer unique opportunities to study redox-based biogeochemical nutrient cycles. Previous work from Lower Kane Cave, Wyoming, USA, focused on the aerobic portion of microbial mats, dominated by putative chemolithoautotrophic, sulfur-oxidizing groups within the Epsilonproteobacteria and Gammaproteobacteria. To evaluate nutrient cycling and turnover within the whole mat system, a multidisciplinary strategy was used to characterize the anaerobic portion of the mats, including application of the full-cycle rRNA approach, the most probable number method, and geochemical and isotopic analyses. Seventeen major taxonomic bacterial groups and one archaeal group were retrieved from the anaerobic portions of the mats, dominated by Deltaproteobacteria and uncultured members of the Chloroflexi phylum. A nutrient spiraling model was applied to evaluate upstream to downstream changes in microbial diversity based on carbon and sulfur nutrient concentrations. Variability in dissolved sulfide concentrations was attributed to changes in the abundance of sulfide-oxidizing microbial groups and shifts in the occurrence and abundance of sulfate-reducing microbes. Gradients in carbon and sulfur isotopic composition indicated that released and recycled byproduct compounds from upstream microbial activities were incorporated by downstream communities. On the basis of the type of available chemical energy, the variability of nutrient species in a spiraling model may explain observed differences in microbial taxonomic affiliations and metabolic functions, thereby spatially linking microbial diversity to nutrient spiraling in the cave stream ecosystem.
Similar content being viewed by others
Introduction
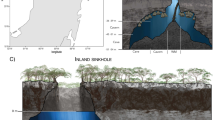
Karst terrains extend over ∼20% of the earth's surface (Ford and Williams, 2007), and caves are common features of karst terrains that allow study of subsurface habitats and ecosystems. Caves systems with surface entrances, but relative hydrological isolation from the surface, share basic physicochemical conditions, including complete darkness, year-round thermal stability, and relatively constant humidity. Caves are also generally isolated from the influence of solar UV radiation and physical weathering due to wind, rain, or freezing. Compared to our understanding of biogeochemical cycling in photosynthetic microbial mats (Ferris et al., 1997; Ward et al., 1998), our knowledge of nutrient cycling in aphotic systems is limited, especially in the context of how nutrient cycling and microbial diversity might be linked. But, the redox-stratified microbial mats from a hydrogen sulfide (H2S)-bearing, anoxic to dysoxic cave stream habitat in Lower Kane Cave, Wyoming, USA (Figure 1), provide an opportunity to evaluate the relationships among habitat geochemistry, nutrient cycling, ecosystem function, and microbial diversity.
(a) General cave location in Wyoming and plan-view cave map from Engel et al. (2004a). (b and c) Representative photographs from the Upper Spring transect, from the spring orifice (b) through the microbial mats downstream (c). Dashed arrows indicate flow direction. Numbers in circles represent sample locations, in meters.
The microbial mats in Lower Kane Cave consist of filamentous and web-like morphotypes in contact with moving stream water. The filamentous surface covers a dense, gel-like mat interior. Dissolved oxygen profiles reveal that the first 3–5 mm of the mats are oxygenated, but that the mat interior is anoxic (PO2<10 Pa) (Engel et al., 2004a). Similar microbial mat and biofilm stratification has been reported recently from the Frasassi Caves in Italy (Macalady et al., 2008). On the basis of the previous full-cycle rRNA investigations from Lower Kane Cave, the aerobic portions of the mats are dominated by uncultured representatives from the Epsilonproteobacteria class and culturable representatives from Thiothrix spp. from the class Gammaproteobacteria (Engel et al., 2003, 2004a). These groups form the bulk of the mats and provide chemolithoautotrophically fixed carbon and other nutrients to the ecosystem, but the diversity and dynamics of anaerobic microbial guilds, and their physical and functional relationships to aerobic groups, remain unknown.
In this study, we focused on the diversity, spatial arrangement, and abundance of anaerobic guilds to better evaluate ecosystem function and nutrient cycling in the microbial mats. We combined the full-cycle 16S rRNA approach, including clone library construction and fluorescence in situ hybridization (FISH), with the most probable number (MPN) method to correlate to guild function. Stream water advection provided an opportunity to evaluate the oxidation state transformations between mobile (for example dissolved in moving stream water) and immobile (for example incorporated as relatively stationary biologic forms) dissolved solutes associated with abiotic and biotic processes. With the expanded view of microbial diversity, the nutrient spiraling concept (Webster and Patten, 1979) was evaluated using carbon and sulfur geochemical and stable isotope data. The distance of solute transport before biogeochemical transformation has been evaluated earlier in stream reaches through nutrient enrichment (Payn et al., 2005) or transport-based analyses (Runkel, 2007), and has been used to describe nitrogen and phosphorus cycling and transport in surface stream ecosystems (Newbold et al., 1981, 1982; Ensign and Doyle, 2006) and algal biofilms (Paul et al., 1991), but to our knowledge this is the first time that nutrient spiraling has been compared with the spatial patterns of aerobic and anaerobic microbial groups within aphotic microbial mats.
Materials and methods
16S rRNA gene diversity and analyses
The microbial diversity of the anaerobic portion of the mats was evaluated using earlier established methods to describe the aerobic portions of the mats (Engel et al., 2003, 2004a; Meisinger et al., 2007). Specific details regarding these methods, including sampling site information, geochemical analyses, DNA extraction, amplification, and sequencing, as well as FISH preparation and sample examination, and MPN estimations, are provided in the Supplementary material (abbreviated ‘SM’). Nearly full-length (>1100 bp) 16S rRNA gene sequences from this study were proof-read and evaluated by the ARB software (Ludwig et al., 2004), which consists of ∼200 000 16S rRNA gene sequences that represent relevant relatives and outgroups to the retrieved clones according to BLAST searches in GenBank (http://www.ncbi.nlm.nih.gov/; Altschul et al., 1990), the Ribosomal Database Project (RDP) (http://rdp.cme.msu.edu/; Cole et al., 2007), and the SILVA databases (http://www.arb-silva.de/; Pruesse et al., 2007). Chimeric clone sequences were identified by the ARB software using a partial treeing approach with two different filter sets that split the full-length sequence into two parts (for example Escherichia coli positions 1–750 and 751–1500) and performed separate treeing calculations on each section. If the two parts of the clone sequences were affiliated to different groups, then the sequences were identified as chimeric; ∼10% of the sequences were identified as chimeric and were not included in further analyses. Nonchimeric sequences were submitted to GenBank under the accession numbers AM490641–AM490771.
Operational taxonomic units (OTUs) at the 98% similarity level were determined using DOTUR (Schloss and Handelsman, 2005). Coverage of the clone library was determined by rarefaction curves generated by aRarefactWin (Analytic Rarefaction ver. 1.3, http://www.uga.edu/strata/software/software.html). Sequences from the cave, close relatives, and outgroups were aligned by comparing select regions of the target gene sequences with alignment positions from multiple data sets (Ludwig et al., 2004) and the resulting overall alignment was visually refined. A general phylogeny was reconstructed using the neighbour-joining distance method (Figure 2). For more refined analyses, sequences for major microbial groups were separated to reconstruct phylogenies using distance, maximum parsimony, and randomized a(x)ccelerated maximum likelihood methods (Stamatakis et al., 2005). For each of these groups, a consensus tree was constructed in ARB from all treeing procedures (Supplementary Figures SM1–SM10).
Neighbor-joining phylogeny of 16S rRNA gene sequences (⩾1100 base pairs) from each of the representative bacterial taxonomic diversity from Lower Kane Cave and closest relatives. Major groups are indicated, and the tree was rooted using four archaeal sequences (AY822003, M60880, L07510, and AF255604).
Distribution patterns from FISH and MPN counts
Twenty-nine rRNA-targeted fluorescently labeled oligonucleotide probes were used for FISH to screen for different microbial groups present in 30 different mat samples distributed along the cave (Supplementary Table SM1). Although detection was limited to groups with at least 105 cells ml–1 in the mats and a relatively high ribosome content (Schleifer, 2004), this approach allowed for screening major taxonomic groups from many different aerobic and anaerobic mat samples without generating clone libraries. Cultivable microbes capable of fermentation, sulfate reduction, sulfur reduction, iron reduction, nitrate-reducing sulfide oxidation, and methanogenesis were enumerated using the MPN method (Hurely and Roscoe, 1983). Details for each enrichment medium are included in the Supplementary material.
Assessment of nutrient cycling and spiraling
Applicability of a nutrient spiraling model was evaluated using previous analyses of aqueous geochemistry and stable isotopes from cave stream waters and microbial mats (Engel et al., 2004a, 2004b). We focused on the Upper Spring transect, in which cave spring water had pH values of 7.1±0.1 and a nearly constant temperature of 21.5 °C. Stream velocity was previously determined from salt-dilution traces, which were approved in 2002 by the Bureau of Land Management as conservative and nondestructive to the microbial mats and cave ecosystem (which includes a federally endangered species); these restrictions currently limit use of radioactive or isotopic tracers. Engel et al. (2004b) measured dissolved and atmospheric gases to determine the theoretical CTS= concentrations in the cave stream if sulfide loss was due to (1) volatilization of gas-phase H2S from the cave stream and not considering turbulence, (2) autoxidation of CTS= along the stream reach (Engel et al., 2004b), or (3) a net first-order loss of CTS= along the stream estimated from CTS= at the upstream and downstream ends of the reach:

where C is CTS= (shortened for clarity), C0 is CTS= at the upstream end of the reach, t is the transport time along the reach, and k is the first-order rate of CTS= transformation. Observed longitudinal distributions of CTS= were compared with the three simple models of sulfide loss listed above (volatilization, autoxidation, and net first-order loss). These comparisons were used to evaluate spatial variability in sulfide production and consumption along the stream, and to compare these patterns to the microbial community structure and function.
Results
Microbial phylogenetic diversity
At least 17 major taxonomic bacterial groups and one archaeal group could be identified based on RFLP screening and partial (<600 bp) 16S rRNA screening of all retrieved (346) clones from the 203 g mat sample (Figure 2; detailed results in Supplementary Table SM2). Rarefaction estimates indicate that coverage barely approached saturation (Figure 3) and that the anaerobic mat community described by this work was more diverse than the aerobic mat samples reported earlier (Engel et al., 2004a). Of the OTUs identified, only ∼7% were ≥98% similar to previously identified groups in GenBank, corresponding to species-level relationships, and 122 novel OTUs were delineated based on 98% sequence identity. More than 50% of the clones were affiliated with three taxonomic groups: the Deltaproteobacteria (δ-Proteobacteria), Chloroflexi, and Gammaproteobacteria (γ-Proteobacteria). Nearly full-length sequences (>1100 bp) were retrieved for 135 representative clones for more detailed phylogenetic evaluation (Figure 2).
Rarefaction curves based on operational taxonomic unit designation showing the diversity of the 203 g 16S rRNA gene sequence bacterial clone library from this study and of the clones from 10 libraries from the aerobic mat samples described in Engel et al. (2004a).
Distribution and ecophysiology within the microbial mats
As clone libraries do not provide distribution information (Juretschko et al., 2002), FISH offered a more realistic view of the distribution and abundance of the dominant and tentatively active (that is with relatively high ribosome contents) groups identified by the clone library approach. Different levels of prokaryotic diversity were targeted with 29 oligonucleotide probes (Supplementary Table SM1), and all but 7 of the 29 probes yielded positive probe signals in at least one of the samples. No signals were observed using the probes targeting Archaea (Arch344), Chloroflexi (CFX1223 and GNSB941), nitrifying bacteria (Ntspa662 and Ntspa712), and Verrucomicrobia (EUBIII) (Table 1). Depending on the sample type, and of the 30 samples examined, Bacteria were detectable by FISH using the general EUB338mix probe for only 18 mat samples. Many of the samples were not suitable for FISH due to prevalent suspended or entrained inorganic material. Higher percentages of detectable cells could be identified by FISH in aerobic samples versus the anaerobic samples (Supplementary Figure SM11). Most of the aerobic and anaerobic mat samples analyzed had high abundances of six probe-targeted groups: the γ-Proteobacteria, Epsilonproteobacteria (ɛ-Proteobacteria), Acidobacteria, α-Proteobacteria, δ-Proteobacteria, and the Firmicutes (Table 1; Supplementary Figure SM11). Filaments were the most targeted cell type, whereby positive signals were observed with probes for ɛ-Proteobacteria, δ-Proteobacteria, γ-Proteobacteria, α-Proteobacteria, and Bacteroidetes. The remaining probe-targeted groups were found in <30% of the investigated mat samples. As functional gene primers are not established for many possible metabolic lifestyles, construction of functional gene clone libraries would have been too time-consuming for the purposes of this study. Therefore, as a way to understand guild distribution in the microbial mats, MPN was used to screen and evaluate potentially lower abundance groups representing different metabolic life styles. Compared to the overall cell estimates of ∼1010 cells ml–1 from the aerobic portions of the mats based on cell carbon content (Engel et al., 2004a), the MPN results enumerated five to six orders of magnitude less cells for each of the various anaerobic guilds (Figure 4).
Most probable number (MPN) estimates for anaerobic microbial guilds from Lower Kane Cave water, sediment, and microbial mat samples compared with dissolved oxygen and sulfide concentrations for each of those samples. Samples are along a transect of the Upper Spring (from 189 to 215 m). Number at left represents site location, in meters, from the back of the cave forward (the spring orifice is the lowest number). White mats are generally considered to be aerobic, while gray mats are anaerobic. SRP, sulfate-reducing prokaryotes (groups I and II, and autotrophic); S0RP, sulfur-reducing prokaryotes; DIRB, dissimilatory iron-reducing bacteria. Error bars are 95% confidence intervals calculated by the MPN program (Hurely and Roscoe, 1983). For nitrate-reducing sulfur-oxidizing bacteria (NRSOB) and methanogens, biomass estimates were not made and therefore only the presence of these groups is indicated; see text for methodological details.
Screening of the mat samples by FISH indicated that at least members affiliated to the δ-Proteobacteria and Firmicutes were present (Table 1; Supplementary Figure SM11), and although the MPN results did not provide information regarding which sulfate-reducing microbial groups (abbreviated as SRP) were enriched, cell estimates for formatotrophic and acetotrophic SRP were equally high (up to 106 cells ml–1) from almost all samples (Figure 4). Autotrophic SRP were not detected in all samples and generally had up to four orders of magnitude fewer cells. SRP cell abundance increased from the Upper Spring orifice downstream through the microbial mats. SRP were considerably less abundant in the cave stream (Figure 4). The aerobic and anaerobic mat samples had nearly equivalent SRP MPN estimates. Sulfate reduction is a polyphyletic metabolic process (Loy et al., 2002), and although several lineages within the δ-Proteobacteria class, the phyla Thermodesulfobacteria, Nitrospirae (for example Thermodesulfovibrio), Firmicutes (for example Desulfotomaculum), and some Archaea (for example Archaeoglobus) gain energy from sulfate reduction, FISH results indicated positive hybridization signals with the δ-proteobacterial and Firmicutes probes in 70% of the mat samples (Table 1; Supplementary Figure SM11). Firmicutes represented ∼7% of the 16S rRNA gene clones, and none were related to known sulfate-reducers (for example, Desulfotomaculum) (Supplementary Table SM2). The majority of clones in the library was affiliated with the δ-Proteobacteria related to environmental sequences retrieved from soil, sediments, the Frasassi Cave sulfidic stream microbial mats, and to cultured SRP like the filamentous Desulfonema spp., Desulfomonile spp., and Chondromyces spp. (Figure 2). S0RP taxonomic diversity is poorly known (Schauder and Kroger, 1993; Kletzin et al., 2004; Kodama et al., 2007). Several species are capable of disproportionating sulfur to sulfide and sulfate, including Desulfocapsa thiozymogenes (Finster et al., 1998), which was retrieved in the clone library (Figure 2). Up to 102 S0RP cells ml–1 were enumerated from MPN (Figure 4).
MPN revealed that fermenting microbes were present in 80% of the mat samples, at abundances up to 106 cells ml–1 in the downstream samples (Figure 4). As fermentation is not limited to select taxonomic groups, different mixed-acid fermentation reactions were assessed (see Supplementary material). Of the 28 mixed-acid fermentation cultures, eight distinct types of heterotrophic growth were identified, predominately from downstream gray filament samples in which ∼70% of the mixed-acid fermentation enrichments produced acidity and CO2 gas. Twenty-eight percent of the mixed-acid fermentation enrichments generated H2S. Ten percent of the enrichments grew on the TSI agar, but did not decrease medium pH or produce gas, suggesting one of four possibilities: (1) no fermentation; (2) fermentation but no decrease in pH or CO2 production because of 2,3-butanediol or ethanol production; (3) only lactose fermentation; or (4) peptone catabolism. Hence, in addition to identifying different organic acid utilization pathways, we could also distinguish between butanediol and ethanol production that is common among Gram-positive bacteria like the Firmicutes (Talarico et al., 2005), which represented 7% of the clone library. This group has not been found in high abundances from sulfidic cave systems previously, but it has been identified from cave-wall microbial communities and cave sediments (Laiz et al., 2003; Schabereiter-Gurtner et al., 2003; Cacchio et al., 2004; Chelius and Moore, 2004; Ikner et al., 2007).
Fourteen percent of the 16S rRNA gene sequence clones were related to uncultured Chloroflexi species (Figure 2; Supplementary Table SM2). Despite Chloroflexi being retrieved from the Frasassi Caves (Engel et al., 2007) and the H2S-bearing Zodletone Spring, Oklahoma (Elshahed et al., 2003), the cave clones were more related to clones retrieved from soil, river sediments, or sludge (90–98% sequence similarity), and distantly related to cultured, anaerobic, filamentous strains (88–90% sequence similarity). FISH revealed that none of the microbial mats showed unambiguous positive hybridization signals to standard Chloroflexi-targeting fluorescently labeled probes (Table 1). However, preliminary studies using horse-radish peroxidase labeled Chloroflexi targeting probes (CFX 1223) and subsequent catalyzed reporter deposition-FISH (Pernthaler et al., 2002) have recently revealed that a significant number of filamentous cells in the mats hybridized with this probe (Meisinger et al., unpublished data). Because of their distant relationship to a large number of uncultured specimens found in anaerobic environments and to culturable nonobligate anaerobic species, like the novel candidate genera Leptolinea and Caldilinea in subclass I and VI of Chloroflexi (Kohno et al., 2002; Yamada et al., 2005) (Supplementary Table SM2), it is plausible that the novel Chloroflexi groups have an anaerobic lifestyle.
The importance of sulfur-oxidizing bacterial groups to sulfuric acid speleogenesis has been made apparent (Engel et al., 2004b; Macalady et al., 2006). The γ-Proteobacteria are predominately affiliated with the filamentous and sulfur-oxidizing genera Thiothrix spp. and Beggiatoa spp., and these groups were retrieved from the clone libraries (Figure 2), and FISH revealed that 70% of all mats had positive hybridization signals with the γ-proteobacterial probes, mostly to filaments, as well as to short and long rods and cocci (Table 1; Supplementary Figure SM11). The anaerobic metabolism of nitrate reduction linked to sulfide oxidation was evaluated by MPN. Half of the nitrate-reducing sulfide-oxidizing bacterial (NRSOB) enrichments had positive growth, predominately from the gray mat samples (Figure 4). As there are currently no FISH probes for this guild because the metabolism is polyphyletic, it was not possible to use FISH to identify NRSOB distribution patterns. But, 60% of the screened mat samples had positive signals for long filaments associated with the ɛ-Proteobacteria, which could indicate the presence of NRSOB (Macalady et al., 2006).
The remaining 16S rRNA clones represented a range of taxonomic and metabolic groups (Figure 2). None of the clones were closely related (>99% sequence identity) to known cultured organisms (Supplementary Table SM2). Nearly 10% of the clones were affiliated with Bacteroidetes/Chlorobi members that were related (89–98% similarity) to clones retrieved from lake, sludge, and tar-oil contaminated aquifers (Supplementary Table SM2). The Bacteriodetes/Chlorobi were only detected in downstream samples by FISH (Supplementary Figure SM11). Despite members of this large group being common in caves, including nonsulfidic examples, their functional role is generally poorly understood and attributed to fermentation or metal cycling (Angert et al., 1998; Schabereiter-Gurtner et al., 2003; Chelius and Moore, 2004; Macalady et al., 2006; Ikner et al., 2007). Up to 103 cells ml–1 of dissimilatory iron-reducing bacteria were identified from anaerobic sediment in the Upper Spring from MPN estimates, but were also detected from the aerobic mat samples downstream (Figure 4). None were detected from spring or stream water. Although Meisinger et al. (2007) hypothesized that some Acidobacteria may be capable of iron reduction, such as those distantly related to Geothrix fermentans, very few clones were retrieved that were closely related to known dissimilatory iron-reducing bacteria (Supplementary Table SM2). Cells hybridized with Acidobacteria-specific probes were detected by FISH in 60% of the samples, and in some instances, this was the most abundant group identified by FISH (Supplementary Figure SM11). Several of the remaining OTUs could not be assigned to a particular division, and given the low sequence identities (86–95%) it is possible that novel groups, even at the familial level, may exist due to affiliations with basal or unlabeled groups (Figure 2).
Diverse methanogens have been identified from ɛ-Proteobacteria- and Thiothrix-dominated microbial mats from photic sulfidic springs (Moissl et al., 2002; Elshahed et al., 2004). However, there are strong indicators that methanogenic guilds have low population sizes in Lower Kane Cave, including trace detectable CH4 in the stream water (Engel et al., 2004a), low cell yields from MPN enrichments (Figure 4), low numbers of archaeal clones using three different primer pairs, and dubious fluorescent signals when using Archaea-specific FISH probes. All of the retrieved clones belonged to the Euryarchaeota, and were most closely related to a methanogenic freshwater sediment clone (Supplementary Table SM2). Interestingly, the rest of the clones amplified by PCR for Archaea were closely related to Thiothrix spp. FISH results were inconclusive for the Archaea because cell numbers may be too low to be detected by FISH, cell inactivity, or low ribosomal content, or because the Archaea probes, ARC344 and in particular Arch915, had unspecific probe signals to certain filaments. When Arch915 was combined with the γ-Proteobacteria probe Gam42a and Thiothrix spp.-specific probes, the filaments had overlapping signals. These results support several previous studies that indicate the present probe (Arch 915) and primer sets (A112f-A934b) used to target Archaea may yield erroneous (nonarchaeal) results (Amann and Fuchs, 2008).
Sulfide dynamics and nutrient spiraling
Spiraling metrics are typically based on travel distance along the stream (as defined by Newbold et al., 1981), assuming an average influence of flow velocity on solute concentrations along a stream reach. Stream velocity was relatively constant from ∼190 to 204 m at 0.5 m s–1, but decreased to 0.2 m s–1 downstream of the microbial mat terminus (from ∼204 to 219 m). We considered changes in CTS= relative to transport time to account for the distinct change in velocity, yielding a more traditional time-based kinetic model. We returned the time-based kinetic analyses to a variably scaled spatial spiraling context (Figure 5) to directly compare locations of CTS= processing and specific microbial guilds along the stream (Doyle, 2005). Although the concentrations of dissolved bicarbonate and sulfate did not vary appreciably along the cave stream, dissolved oxygen increased while CTS= values decreased (Figures 5b and c). The mats abruptly terminate between 203 and 204 m where dissolved oxygen exceeded 45 μmol l−1and CTS= was half the initial concentration (∼28.7 s). From Equation (1), k was estimated as 0.02 s−1 from the difference between upstream and downstream CTS=. From 0 to 5 s, and beyond 30 s travel time, the observed CTS= values generally follow a first-order CTS= loss curve, although the observed values are slightly less than the expected values from 0 to 5 s (Figure 5c, black curve). Between 5 and 30 s, however, observed CTS= differed from the first-order loss curve. At any given dissolved oxygen concentration, the observed CTS= values were lower than would be expected for CTS= if loss was due to either autoxidation or volatilization processes (Figure 5c, gray curves) (Engel et al., 2004b).
(a) Dissolved bicarbonate and sulfate concentrations. (b) Dissolved oxygen and total dissolve sulfide (CTS=) concentrations for the Upper Spring transect. (c) Observed CTS= versus modeled CTS=/CTS=0 based on a first-order loss from t1/2 of 28.7 s (black line), and on autoxidation and volatilization loss (gray lines) according to Engel et al. (2004b). The broken x axis indicates where there is a change in velocity. The shaded box in all panels denotes the stream reach with microbial mats (from 0 to ∼30 s, or 190–204 m).
Discussion
In the Lower Kane Cave stream system, the microbial mats are likely to alter the hydrodynamics of a system, increase the contact time of water and solutes with both aerobic and anaerobic portions of the mat, and thereby compress nutrient consumption and production spirals along the cave stream (Battin et al., 2003). So, the susceptibility of a molecule to metabolic transformation, such as from a mobile to immobile phase, increases as solute transport into mats from the water column becomes limiting. Gas-phase nutrients, such as H2S, O2, and CO2, have the greatest potential to be limiting, and we hypothesized that the resulting variations in the uptake (consumption) and release (production) rates of these nutrients downstream should influence the longitudinal distribution of specific metabolic groups. We combined the full-cycle 16S rRNA approach with the MPN method to realize the physical and functional distribution of anaerobic microbial guilds and communities in the microbial mats. Our phylogenetic investigations uncovered the potential for undescribed biodiversity, and some groups have not been identified previously in subsurface systems. These results made evaluating the mechanisms that control changes in diversity difficult (Doebeli and Dieckmann, 2003; Mizera and Meszena, 2003; Hansen et al., 2007). But, the spatial distribution of microbial diversity was placed into a nutrient spiraling framework by interpreting hydraulic, geochemical, and biological processes that balance nutrient and energy flow along the cave stream (Webster and Patten, 1979; Newbold et al., 1981, 1982; Runkel, 2007).
Consideration of stable carbon and sulfur isotope ratio analyses from the microbial mats and cave waters illustrate that nutrient spiraling influences changes in microbial diversity, and that a single net fractionation cannot be applied to the system due to the diversity of biological processes that offset the isotopic composition along stream (Figure 6). For Lower Kane Cave, chemolithoautotrophically fixed carbon serves as the trophic basis of this microbial ecosystem because photosynthetically produced carbon is unlikely to provide any significant energy to the system due to the cave's isolation from the surface (Engel et al., 2004a; Porter et al., 2009). The δ13C values for microbial biomass are typical of autotroph-dominated biomass because they reflect the large, irreversible enzymatic discrimination against 13C from a dissolved inorganic carbon reservoir (Figure 6a). The dissolved inorganic carbon δ13C values decrease by ∼2‰ from the spring orifice (190 m) to the end of the mats at ∼204 m (Engel et al., 2004a). Porter et al. (2009) indicate that at least 30% of the autotrophically fixed carbon in Lower Kane Cave microbial mats is cycled through a microbial detrital loop. From 0 to 10 s (∼190–195 m), where measured CTS= values are only slightly lower than expected (given experimental error) from the first-order loss curve (Figure 5c), the δ13C values for upstream samples, which consist predominately of ɛ-Proteobacteria and few other groups (Supplementary Figure SM11), differ by <5‰ (Figure 6a). By ∼16–17 s (∼197 m), however, the differences between δ13C values from the white and gray mat samples increases to >7‰. As excretion, respiration, and heterotrophic cycling of carbon should have negligible effects on carbon isotope values, the differences in δ13C values could be linked to changes in the abundances of different autotrophic ɛ- and γ-proteobacterial groups and variations in the rates and expression of 13C discrimination during autotrophy (Engel et al., 2004a). From 20 to 30 s (∼200–205 m), the white and gray δ13C biomass compositions are less variable, suggesting that the proximity of heterotrophs to autotrophically produced organic carbon may cause immediate sequestration of carbon rather than its downstream transport (Battin et al., 2003). In other words, the biomass δ13C decreases with each carbon spiral along the stream because of a feedback effect between autotrophic fractionation (negative for 13C) and the continual respiration of autotrophically fixed organic carbon from upstream. Carbon spiraling should homogenize the δ13C in the organic and inorganic carbon pools with increasing distance downstream, as well as cause the ensuing decrease in the δ13C dissolved inorganic carbon downstream, because CO2 respired upstream is used by downstream autotrophs.
(a) Carbon isotope compositions summarized from Engel et al. (2004a) for white (aerobic) and gray (anaerobic) filamentous microbial mat samples. (b) Sulfur isotope compositions for microbial biomass and the dissolved sulfide from the cave stream (Engel et al., 2007). The broken x axis indicates a change in velocity.
The δ34S values for CTS= in the cave stream decrease by ∼1.6‰ along the cave stream, and white and gray biomass δ34S values are variable (Figure 6b). The gray mat samples from the middle of the stream have the highest abundance of sulfate-reducing and heterotrophic groups (Figure 4), and these samples also correspond to a reach where measured CTS= exceeds the first-order CTS= decay loss (Figure 5c). The increase in CTS= could not be due to abiotic processes, so the large and diverse community of SRP and fermenters are responsible for producing autochthonous sulfide. As autochthonous H2S would have very low δ34S values compared with allochthonous dissolved sulfide in the cave stream, consumption of this autochthonous sulfide by sulfur-oxidizing community would shift the average δ34S values for microbial biomass downstream by more than ±1‰ (Fry et al., 1988; Mandernack et al., 2003). Differences in autochthonous H2S uptake rates downstream, as well as differences in retention of sulfur-containing compounds within the mats downstream, may result in compositional variability. The mean sulfur content of white filaments is ∼30% w/w, whereas gray filaments only have ∼2% sulfur (Engel et al., 2004a). In contrast, most cells have values approximately equal to 1% in the absence of stored sulfur (Fagerbakke et al., 1996). Engel et al. (2007) confirmed that sulfur in white filaments is cyclo-octasulfur (S8), or elemental sulfur, which accumulates as a byproduct of sulfur oxidation, but that sulfur in the gray filaments is mostly as monosulfane from the accumulation of amino acids.
These correlations among microbial abundance and diversity, CTS= dynamics, and the isotopic composition of water and mats provide a basic understanding that nutrient spiral dimensions lengthen or shorten (that is loosen or tighten; Webster and Patten, 1979) in different segments of the stream, even if we do not yet fully understand uptake and turnover dynamics between the aerobic and anaerobic portions of the mats. For instance, it is possible that some groups may facultatively switch from autotrophy to mixotrophy as autochthonous organic carbon availability increases. Therefore, given the complexity of this natural system, future work will require a more detailed characterization of the hydrology (Doyle, 2005; Runkel, 2007), nutrient addition experiments, and the introduction of conservative hydrologic tracers to quantify how spiraling dimensions change with shifts in community composition and metabolism, or how biological or hydraulic retention influence spiral lengths due to reactive transport and nutrient retention and storage within the microbial mats. More detailed modeling of the system will also require characterizing gas (H2S) transfer across the stream–air interface, which is not implemented in the more commonly used active solute transport models (for example OTIS; Runkel, 1998). Lastly, microbial functional processes should also be explored and quantified, such as from functional genomics and expression studies.
In conclusion, this study expands the known diversity of some microbial groups to caves and the subsurface, even though the microbial diversity of various sulfidic caves and aquifers has been investigated for more than a decade (Sarbu et al., 1996; Vlasceanu et al., 1997, 2000; Hose et al., 2000; Barton and Luiszer, 2005; Macalady et al., 2006). In Lower Kane Cave, anaerobic mat communities were more taxonomically and metabolically diverse relative to the aerobic mat communities. Although groups like SRP and fermenters present a fraction of the whole cell abundance of the system, their metabolic impact is reflected and recorded in the geochemistry (that is isotopically) of the system. Changes in the forms and abundances of nutrients influence the overall functional diversity with distance downstream, and this may result in the emergence of potentially specialized groups of differing genetic and functional compositions downstream (Battin et al., 2003; Mizera and Meszena, 2003; Parnell et al., 2009). Evaluation of carbon and sulfur nutrient spiraling in Lower Kane Cave improves our understanding of ecosystem function in aphotic habitats and will help us to understand microbial processes occurring in other modern, redox-stratified microbial mat systems.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ . (1990). Basic local alignment search tool. J Mol Biol 215: 403–410.
Amann R, Fuchs BM . (2008). Single-cell identification in microbial communities by improved fluorescence in situ hybridization techniques. Nature Rev Microbiol 6: 339–348.
Angert ER, Northup DE, Reysenbach A-L, Peek AS, Goebel BM, Pace NR . (1998). Molecular phylogenetic analysis of a bacterial community in Sulphur River, Parker Cave, Kentucky. Am Mineral 83: 1583–1592.
Barton HA, Luiszer F . (2005). Microbial metabolic structure in a sulfidic cave hot spring: potential mechanisms of biospeleogenesis. J Cave Karst Stud 67: 28–38.
Battin TJ, Kaplan LA, Newbold JD, Hansen CME . (2003). Contributions of microbial biofilms to ecosystem processes in stream mesocosms. Nature 426: 439–442.
Cacchio P, Contento R, Ercole C, Cappuccio G, Preite Martinez M, Lepidi A . (2004). Involvement of microorganisms in the formation of carbonate speleothems in the Cervo Cave (L'Aquila-Italy). Geomicrobiol J 21: 497–509.
Chelius MK, Moore JC . (2004). Molecular phylogenetic analysis of Archaea and bacteria in Wind Cave, South Dakota. Geomicrobiol J 21: 123–134.
Cole JR, Chai B, Farris RJ, Wang Q, Kulam-Syed-Mohideen AS, McGarrell DM et al. (2007). The ribosomal database project (RDP-II): introducing myRDP space and quality controlled public data. Nucleic Acids Res 35: D169–D172.
Doebeli M, Dieckmann U . (2003). Speciation along environmental gradients. Nature 421: 259–264.
Doyle MW . (2005). Incorporating hydrologic variability into nutrient spiraling. J Geophy Res 110: G01003, doi:01010.01029/02005JG000015.
Elshahed MS, Najar FZ, Roe BA, Oren A, Dewers TA, Krumholz LR . (2004). Survey of archaeal diversity reveals an abundance of halophilic Archaea in a low-salt, sulfide- and sulfur-rich spring. Appl Environ Microbiol 70: 2230–2239.
Elshahed MS, Senko JM, Najar FZ, Kenton SM, Roe BA, Dewers TA et al. (2003). Bacterial diversity and sulfur cycling in a mesophilic sulfide-rich spring. Appl Environ Microbiol 69: 5609–5621.
Engel AS, Lee N, Porter ML, Stern LA, Bennett PC, Wagner M . (2003). Filamentous ‘Epsilonproteobacteria’ dominate microbial mats from sulfidic cave springs. Appl Environ Microbiol 69: 5503–5511.
Engel AS, Lichtenberg H, Prange A, Hormes J . (2007). Speciation of sulfur from filamentous microbial mats from sulfidic cave springs using X-ray absorption near-edge spectroscopy. FEMS Microbiol Lett 269: 54–62.
Engel AS, Porter ML, Stern LA, Quinlan S, Bennett PC . (2004a). Bacterial diversity and ecosystem function of filamentous microbial mats from aphotic (cave) sulfidic springs dominated by chemolithoautotrophic ‘Epsilonproteobacteria’. FEMS Microbiol Ecol 51: 31–53.
Engel AS, Stern LA, Bennett PC . (2004b). Microbial contributions to cave formation: new insight into sulfuric acid speleogenesis. Geology 32: 369–372.
Ensign SH, Doyle MW . (2006). Nutrient spiraling in streams and river networks. J Geophys Res 111: G04009; doi:10.1029/2005JG000114.
Fagerbakke KM, Heldal M, Norland S . (1996). Content of carbon, nitrogen, oxygen, sulfur, and phosphorous in native aquatic and cultured bacteria. Aq Microbial Ecol 10: 15–27.
Ferris MJ, Nold SC, Revsbech NP, Ward DM . (1997). Population structure and physiological changes within a hot spring microbial mat community following disturbance. Appl Environ Microbiol 63: 1367–1374.
Finster K, Liesack W, Thamdrup B . (1998). Elemental sulfur and thiosulfate disproportionation by Desulfocapsa sulfoexigens sp. nov., a new anaerobic bacterium isolated from marine surface sediment. Appl Environ Microbiol 64: 119–125.
Ford DC, Williams P . (2007). Karst Geomorphology and Hydrology. J Wiley & Sons, Inc.: Sussex, pp 576.
Fry B, Ruf W, Gest H, Hayes JM . (1988). Sulfur isotope effects associated with oxidation of sulfide by O2 in aqueous solution. Chem Geol 73: 205–210.
Hansen SK, Rainey PB, Haagensen JAJ, Molin S . (2007). Evolution of species interactions in a biofilm community. Nature 445: 533–536.
Hose LD, Palmer AN, Palmer MV, Northup DE, Boston PJ, DuChene HR . (2000). Microbiology and geochemistry in a hydrogen-sulphide rich karst environment. Chem Geol 169: 399–423.
Hurely MA, Roscoe ME . (1983). Automated statistical analysis of microbial enumeration by dilution series. J Appl Bacteriol 55: 157–164.
Ikner LA, Toomey RS, Nolan G, Neilson JW, Pryor BM, Maier RM . (2007). Culturable microbial diversity and the impact of tourism in Kartchner Caverns, Arizona. Microbial Ecol 53: 30–42.
Juretschko S, Loy A, Lehner A, Wagner M . (2002). The microbial community composition of a nitrifying-denitrifying activated sludge from an industrial sewage treatment plant analyzed by the full-cycle rRNA approach. System Appl Microbiol 25: 84–99.
Kletzin A, Urich T, Müller F, Bandeiras TM, Gomes CM . (2004). Dissimilatory oxidation and reduction of elemental sulfur in thermophilic archaea. J Bioenerg Biomembr 36: 77–91.
Kodama Y, Ha LT, Watanabe K . (2007). Sulfurospirillum cavolei sp. nov., a facultatively anaerobic sulfur-reducing bacterium isolated from an underground crude oil storage cavity. Int J Syst Evol Microbiol 57: 827–831.
Kohno T, Sei K, Mori K . (2002). Characterization of Type 1851 organism isolated from activated sludge samples. Water Sci Technol 46: 111–114.
Laiz L, Gonzalez-Delvalle M, Hermosin B, Ortiz-Martinez A, Saiz-Jimenez C . (2003). Isolation of cave bacteria and substrate utilization at different temperatures. Geomicrobiol J 20: 479–489.
Loy A, Lehner A, Lee N, Adamczyk J, Meier H, Ernst J et al. (2002). Oligonucleotide microarray for 16S rRNA gene-based detection of all recognized lineages of sulfate-reducing prokaryotes in the environment. Appl Environ Microbiol 68: 5064–5081.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al. (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Macalady JL, Dattagupta S, Schaperdoth I, Jones DS, Druschel GK, Eastman D . (2008). Niche differentiation among sulfur-oxidizing bacterial populations in cave waters. ISME J 2: 590–601.
Macalady JL, Lyon EH, Koffman B, Albertson LK, Meyer K, Galdenzi S et al. (2006). Dominant microbial populations in limestone-corroding stream biofilms, Frasassi Cave system, Italy. Appl Environ Microbiol 72: 5596–5609.
Mandernack KW, Krouse HR, Skei JM . (2003). A stable sulfur and oxygen isotopic investigation of sulfur cycling in an anoxic marine basin, Framvaren Fjord, Norway. Chem Geol 195: 181–200.
Meisinger DB, Zimmermann J, Ludwig W, Schleifer K-H, Wanner G, Schmid M et al. (2007). In situ detection of novel Acidobacteria in microbial mats from a chemolithoautotrophically based cave ecosystem (Lower Kane Cave, WY, USA). Environ Microbiol 9: 1523–1534.
Mizera F, Meszena G . (2003). Spatial niche packing, character displacement and adaptive speciation along an environmental gradient. Evol Ecol Res 5: 363–382.
Moissl C, Rudolph C, Huber R . (2002). Natural communities of novel Archaea and bacteria with a string-of-pearls-like morphology: molecular analysis of the bacterial partners. Appl Environ Microbiol 68: 933–937.
Newbold JD, Elwood JW, O'Neill RV, Van Winkle W . (1981). Measuring nutrient spiralling in streams. Can J Fish Aquat Sci 38: 860–863.
Newbold JD, Mulholland PJ, Elwood JW, O'Neill RV . (1982). Organic carbon spiralling in stream ecosystems. Oikos 38: 266–272.
Parnell JJ, Crowl TA, Weimer BC, Pfrender ME . (2009). Biodiversity in microbial communities: system scale patterns and mechanisms. Mol Eco 18: 1455–1462; doi: 10.1111/j.1365-294X.2009.04128.x.
Paul BJ, Duthie HC, Taylor WD . (1991). Nutrient cycling by biofilms in running waters of differing nutrient status. J North Am Benthol Soc 10: 31–41.
Payn RA, Webster JR, Mulholland PJ, Valett HM, Dodds WK . (2005). Estimation of stream nutrient uptake from nutrient addition experiments. Limnol Oceanogr Meth 3: 174–182.
Pernthaler A, Pernthaler J, Amann R . (2002). Fluorescence in situ hybridization and catalyzed reporter deposition for the identification of marine bacteria. Appl Environ Microbiol 68: 3094–3101.
Porter ML, Engel AS, Kane TC, Kinkle BK . (2009). Productivity-diversity relationships from chemolithoautotrophically based sulfidic karst systems. Int J Speleol 3: 27–40.
Pruesse E, Quast C, Knittel K, Fuchs B, Ludwig W, Peplies J et al. (2007). SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acid Res 35: 7188–7196.
Runkel RL . (1998). One dimensional transport with inflow and storage (OTIS): a solute transport model for streams and rivers. U.S. Geological Survey Water-Resources Investigations Report 98-4018. US Geological Survey, Denver, Colorado (http://co.water.usgs.gov/otis).
Runkel RL . (2007). Toward a transport-based analysis of nutrient spiraling and uptake in streams. Limnol Oceanogr Meth 5: 50–62.
Sarbu SM, Kane TC, Kinkle BK . (1996). A chemoautotrophically based cave ecosystem. Science 272: 1953–1955.
Schabereiter-Gurtner C, Saiz-Jimenez C, Piñar G, Lubitz W, Rölleke S . (2003). Phylogenetic diversity of bacteria associated with Paleolithic paintings and surrounding rock walls in two Spanish caves (Llonín and La Garma). FEMS Microbiol Ecol 1606: 1–13.
Schauder R, Kroger A . (1993). Bacterial sulfur respiration. Arch Microbiol 195: 491–497.
Schleifer K-H . (2004). Microbial diversity: facts, problems and prospects. System Appl Microbiol 27: 3–9.
Schloss PD, Handelsman J . (2005). Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microbiol 71: 1501–1506.
Stamatakis AP, Ludwig T, Meier H . (2005). RAxML-II: a program for sequential, parallel & distributed inference of large phylogenetic trees. Concurrency Comp Practice Exp 17: 1705–1723.
Talarico LA, Gil MA, Ingram LO, Maupin-Furlow JA . (2005). Construction and expression of an ethanol production operon in gram-positive bacteria. Microbiol 151: 4023–4031.
Vlasceanu L, Popa R, Kinkle B . (1997). Characterization of Thiobacillus thioparus LV43 and its distribution in a chemoautotrophically based groundwater ecosystem. Appl Environ Microbiol 63: 3123–3127.
Vlasceanu L, Sarbu SM, Engel AS, Kinkle BK . (2000). Acidic cave-wall biofilms located in the Frasassi Gorge, Italy. Geomicrobiol J 17: 125–139.
Ward DM, Ferris MJ, Nold SC, Bateson MM . (1998). A natural view of microbial biodiversity within hot spring cyanobacterial mat communities. Microbiol Mol Biol Rev 62: 1353–1370.
Webster JR, Patten BC . (1979). Effects of watershed perturbation on stream potassium and calcium dynamics. Ecol Mono 19: 51–72.
Yamada T, Sekiguchi Y, Imachi H, Kamagata Y, Ohashi A, Harada H . (2005). Diversity, localization, and physiological properties of filamentous microbes belonging to Chloroflexi subphylum I in mesophilic and thermophilic methanogenic sludge granules. Appl Environ Microbiol 71: 7493–7503.
Acknowledgements
The Bureau of Land Management, Cody Office, granted permission for the work. Field assistance was given by PC Bennett, SA Engel, T Dogwiler, K Mabin, M Edwards, J Deans, and HH Hobbs, III. Partial funding was provided from the Department Geological Sciences of The University of Texas at Austin (to ASE), the Life in Extreme Environments (LExEn) program of the NSF (EAR-0085576 to PC Bennett and LAS at The University of Texas at Austin), and from the Technische Universität München and the Helmholtz Foundation for the ‘virtual institute for isotope biogeochemistry–biologically mediated processes at geochemical gradients and interfaces in soil—water systems’ (to DBM and NML). Continued funding to ASE was provided by the College of Basic Sciences, Louisiana State University. Insightful contributions for the nutrient spiraling concept were provided by K Simon and H Bao, and valuable discussions were also provided by PC Bennett, M Wagner (University of Vienna), W Liebl and W Ludwig (both at TUM, Technische Universität München).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Rights and permissions
About this article
Cite this article
Engel, A., Meisinger, D., Porter, M. et al. Linking phylogenetic and functional diversity to nutrient spiraling in microbial mats from Lower Kane Cave (USA). ISME J 4, 98–110 (2010). https://doi.org/10.1038/ismej.2009.91
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2009.91
Keywords
This article is cited by
-
Topsoil and subsoil bacterial community assemblies across different drainage conditions in a mountain environment
Biological Research (2023)
-
Competition-cooperation in the chemoautotrophic ecosystem of Movile Cave: first metagenomic approach on sediments
Environmental Microbiome (2022)
-
Studies on Geomicrobiology in some Indian Caves
Journal of the Geological Society of India (2022)
-
Composition and functional profiles of microbial communities in two geochemically and mineralogically different caves
Applied Microbiology and Biotechnology (2021)
-
Co-occurrence pattern and function prediction of bacterial community in Karst cave
BMC Microbiology (2020)