Abstract
The gastrointestinal tract of mammals contains a complex microbial community that influences numerous aspects of health and development. It is postulated that establishment of this community during early life has long-term consequences on the health status of adults. Potential influences on colonization are expected to include environmental microbes, diet and the developmental changes of the host. Denaturing gradient gel electrophoresis was used to follow the individual community dynamics of 24 piglets over the period of 3–36 days after birth. The community of piglets older than 31 days was inferred to show high stability relative to the first 28 days post birth. The stable day 36 community showed significant correlation between cohabiting piglets, but not between siblings. This cohabitation effect was not observable in 1- or 2-week-old piglets but was strongest at either 3 or 4 weeks post birth. The onset of this change after 2 weeks is predicted to be after the development of key induction elements of the immune system and before significant levels of piglet sIgA were observable (4 weeks). The outcome is altered community dynamics that result in significant similarity between the stable communities that develop in cohabiting pigs. We conclude that for a finite period in their development, the outcome of gut colonization in piglets is greatly influenced by the immediate environment. The implication is that mammals have a developmental window, in which the developing host–gut microbiota interaction will be simultaneously more amenable to engineering and more susceptible to disturbance.
Similar content being viewed by others
Introduction
Animals naturally harbour enormous numbers of microorganisms within their lower gastrointestinal tract (collectively referred here as the intestinal microbiota). The presence and activity of these microbes impacts the host animal in a variety of ways. For example, microbial colonization is necessary for several aspects of postnatal development (Alam et al., 1994; Stappenbeck et al., 2002; Mazmanian et al., 2005) and the activity of intestinal microbes contributes to host nutrition and metabolism through vitamin synthesis (Gustafsson, 1959), increased degradation of polysaccharides (Salyers et al., 1977) and catabolism of toxins (Swanson et al., 1987). However, the role of variation in the composition of the intestinal microbiota in determining the outcomes for the host and how to manage this is not well understood.
Early culture-based studies concluded that the broad process and outcomes of intestinal colonization were similar in diverse animal species (Smith, 1965). The first colonizers are acquired during birth and over the first few days, populations of anaerobes increase to become dominant (Stark and Lee, 1982). This neonatal microbiota is relatively dynamic and compositionally distinct to that of postweaned (maternally independent) young. For example, in breast-fed humans, the neonatal community can be dominated by bifidobacteria, which are minor or absent in adults (Yoshioka et al., 1983). Intensive culture efforts suggested the intestinal microbiota of humans could include up to 400 species (Moore and Holdeman, 1974).
The advent of culture-independent approaches to assessing community structure has given more detail to this picture. Large-scale, culture-independent studies of community structure, based on ribosomal RNA sequence analysis, have been conducted for intestinal microbiota in humans, mice and pigs (Leser et al., 2002; Eckburg et al., 2005; Ley et al., 2005). Available data show the microbial communities of all three have similar composition at coarse phylogenetic resolution (taxa defined at >85% rRNA identity) and the majority of taxa belong to just two bacterial divisions, the Bacteroidetes and the Firmicutes. Considerable diversity is seen at finer scales of observation. More than half the taxa reported in each of the human, pig and mouse studies were unique to that study, if taxa were defined at relatedness of >95% rRNA identity (approximately genus) and uniqueness to individuals was seen at finer scales. As a consequence, investigations of the significance of community composition to an individual's health must focus on fine-scale diversity at the species and strain level (in rRNA terms, this approximates 97% and 99% identity, respectively). At this scale of resolution, there is evidence that community structure is a source of variation in host outcomes (Backhed et al., 2004; Robosky et al., 2005). This has given rise to the concept that diseases of relevance to humans, or growth performance issues relevant to animal production, can be linked to the quality of an individual's intestinal microbiota.
Our interest is in how individual microbiotas become established since the processes that lead to the development of a stable gut microbiota are anticipated to have life-long consequences for the individual (Tannock, 2005). The potential to inadvertently trigger undesirable outcomes of the colonization process, such as a microbiota predisposing to disease (for example, obesity or immunological disorders), or to engineer the establishment of a desirable microbiota, is largely unknown (Ley et al., 2006). It is, therefore, important that colonization processes are better understood to determine which factors have long-term consequences for gut community structure. Some studies have looked at the composition of microbiota at distinct ages (Konstantinov et al., 2006) but very few studies have looked in any detail at community dynamics during the establishment of an individual's intestinal microbiota.
The establishment of the gut community begins during birth and although the infant is exposed to many different microbes from the environment, only a subset of these will become established in that individual. Potentially important factors include the environmental microbial exposure, diet and host physiology. The relative importance of these is likely to vary with the age of the individual. The environmental microbial exposure during the immediate postnatal period will be dominated by the mother's microbiota but its relative importance will decline with the increasing independence of the infant. In mammals, diet (intake of breast milk followed by weaning) and its impact on the microbial community structure is known to change with time. For example, large differences in the microbiota of breastfed human infants compared to formula-fed infants have been reported (Harmsen et al., 2000) and differences in the porcine gut community associated with weaning (Konstantinov et al., 2006). However, later in life, changes to the diet produce only temporary or no apparent change to community structure in adults (Mai et al., 2004; Tannock et al., 2004). Finally, the immune system undergoes considerable postnatal development and this may be expected to result in a decrease in the potential for environmental microbes to colonize as the infant develops. Understanding the dynamics of microbial colonization requires a degree of control of these factors and the repeated sampling of individuals over the period from birth to complete immune development and/or independence from the mother. There are few studies that have used sampling intensities of weekly or better over this period (Favier et al., 2002; Inoue et al., 2005b) and only one that examined >10 individuals (Palmer et al., 2007). The general dynamics of microbial colonization remain very poorly known for any animal model.
We have used pigs as our model system and followed community dynamics of individuals through denaturing gradient gel electrophoresis (DGGE) fingerprinting of fecal samples collected over their first 36 days of life. Levels of fecal IgA were monitored in one cohort of pigs as an indicator of immunological development. The microbial assemblage present in the feces will not be identical to any specific community within the intestinal tract but stool samples are the only practical way to obtain longitudinal samples reflective of an individual's intestinal microbiota. The precocial nature of piglets allows their separation from the mother shortly after birth, thus allowing a degree of control over diet and environmental microbial exposure. The rapid development of pigs means that community dynamics of individual's microbiota can be tracked from near-birth until development of an adult-like immune capacity with relatively few samples. Electrophoretic fingerprinting techniques, such as DGGE, are capable of discriminating between species and sometimes strains of the same species. They differ from sequencing approaches in that classification into operational taxonomic units is on the basis of DNA mobility rather than sequence relatedness. As a consequence, they do not directly yield insight into the composition of a microbial community at any point, but they are a powerful means of detecting patterns of change in composition through space and time. Previous studies have shown that within-gel comparisons of electrophoretic fingerprints are highly reproducible and capable of resolving stable inter-individual differences in intestinal microbiota over time (Simpson et al., 1999, 2002; Favier et al., 2002; Inoue et al., 2005b) Here, we have shown that the intestinal microbiota of piglets >30 days old is stable relative to that of younger piglets. A significant correlation was observed between housing environment and community structure. Surprisingly, this correlation had a very sudden onset, occurring after 14 days and before 21 days of age. This finding suggests that environmental microbial exposure within a specific developmental window is a major determinant of adult microbial community structure.
Materials and methods
Animals and sample collection
Data were collected from a total of 35 piglets over three animal trials. The study protocol was approved by the University of Sydney Animal Ethics Committee. The first two trials, each with 12 piglets, were in 2004. Piglets were obtained from a commercial piggery: Hillcrest piggery, Menangle, NSW, Australia for the first trial (pigs 1–12) and Boen Boe piggery, Mittagong, NSW, Australia for the second trial (pigs 13–24). Male domestic piglets (Sus scrofa, Landrace/Large White cross) were obtained at 3 days of age from a total of five different litters and the sow but not the sire was identified for each piglet (refer to Supplementary Table S1). All piglets were housed in pairs and kept in the same temperature-controlled room on a 12 h light and dark cycle. Unrelated piglets of similar weight were cohoused wherever possible. Each group was fed a standard diet of soy/whey/casein (55:9:6) sow milk replacer (Wombaroo Food Products, Glen Osmond, South Australia, Australia) until 36 days of age. Milk intake for all piglets was 285 ml kg−1 day−1 during the first 2 weeks and 230 ml kg−1 day−1 for the remaining weeks. Fecal samples (n=102) were obtained by directly monitoring all piglets. Each sample was collected immediately after defecation and could, therefore, be assigned to an individual piglet. Gut contents from piglets were also obtained directly from the distal colon following euthanization at 36 days of age. All samples were immediately frozen at −20 °C. Prior to DNA extraction, 1 g of the fecal sample was homogenized in 5 ml of 10 mM TE buffer (1 mM EDTA, pH 7.5). This was then used for DNA extraction and DGGE analysis.
A further 72 fecal samples were collected from 11 individual male piglets during a third trial in 2006. The piglets were obtained from Boen Boe piggery, Mittagong, NSW, Australia and originated from three litters (refer to Supplementary Table S2). The piglets were housed and fed as described above. The fecal samples were used to measure fecal IgA concentration. Samples were collected as above but were not homogenized prior to fecal IgA extraction. Samples from one piglet (pig no. 25) were used for DNA extraction and DGGE analysis to examine daily variation and to correlate IgA with gut community structure.
For analysis, samples were grouped into series by time of collection as described in Supplementary Tables S1 and S2. For the sample set used for DGGE analysis, samples collected at 7–9 days were classed as week 1, 14–18 days as week 2, 22–24 days as week 3, 28–32 days as week 4 and samples obtained at euthanisation at 36 days as week 5. For IgA analysis, samples were collected at 3–7 days, 10–13 days, 20–21 days, 28 days and 36 days.
DNA extraction
Extraction of DNA from the homogenised fecal samples was carried out, using the FastPrep system (Bio101, La Jolla, CA, USA) with modifications as described previously (Yeates et al., 1998). Briefly, 0.5 ml of homogenized fecal sample was placed in a screw-cap tube with a single 5 mm glass bead and 0.3 g each of 150–212 and 472–600 μm glass beads (Sigma-Aldrich, St Louis, MO, USA). Cells were lysed in a Bio101 FP-120 Fastprep machine (Bio101) on a speed setting of 5.5 for 30 s and the DNA was isolated as per the protocol for the FastDNA Spin Kit for soil (Bio101). The DNA was suspended in TE buffer and was stored at −20 °C. Two independent DNA extractions were made per fecal homogenate and reproducibility was confirmed by DGGE analysis.
PCR amplification for DGGE
PCR primers F-968-GC and R-1401 (Nubel et al., 1996) were used to amplify the V6-V8 segment of the 16S rDNA. Each 25 μl reaction volume contained 1 × thermopol buffer (New England BioLabs, Beverley, MA, USA), 5 mM deoxynucleoside triphosphates (New England BioLabs), 20 pmoles F-968-GC, 10 pmoles R-1401, 1U Taq polymerase DNA (New England BioLabs) and 1 μl of fecal DNA. The program used was as follows: 1 min of initial denaturation at 94 °C, followed by 30 cycles of denaturation (94 °C for 30 s), annealing (56 °C for 30 s) and extension (72 °C for 1 min) with a final extension for 7 min at 72 °C). Reproducibility was assessed by independent amplification of the same DNA sample.
DGGE analysis
DGGE analysis was performed using the DCode system (Bio-Rad Laboratories, Hercules, CA, USA). Electrophoresis was done using a 16 × 16 cm 1-mm-thick gel that contained 8% polyacrylamide (ratio of acrylamide to bisacrylamide was 37.5:1) in 1 × TAE buffer (40 mM Tris–acetate, 1 mM EDTA; pH 7.4). A gradient of 40%–70% denaturant was used to separate PCR fragments where 100% denaturant was defined as 7 M urea and 40% (v/v) formamide. The gels were run at 80 V for 16 h at 60 °C. After electrophoresis, the gels were silver stained as described (Sambrook, 2001). Gels were scanned using a GS-800 calibrated densitometer (Bio-Rad).
Analysis of DGGE profiles
The digitized gel images were analysed using Quantity One (version 4. 6.1; Bio-Rad). The software was used to detect bands in each lane, using a match tolerance of 2%. A similarity matrix was constructed using Dice's similarity coefficient. This is defined as [2j/(a + b)] × 100, where j is the number of bands in common between two lanes, and (a+b) is the total band number of both lanes. Dendrograms were constructed by the unweighted pair group method, using arithmetic averages (UPGMA). The significance of comparisons between groups of littermates and cohabitating piglets was assessed using a two-tailed unpaired Student's t-test. All analyses were conducted within the same gel (that is, lanes from different gels were not directly compared).
Fecal immunoglobulin extraction
Fecal samples collected in 2006 were dried by vacuum centrifuge. The dry weight was measured and a subsample of feces was homogenized in 5% non-fat milk with phosphate-buffered saline (Sigma-Aldrich). The fecal suspension was centrifuged at 14 680 r.p.m. The supernatant was retained and a protease inhibitor cocktail (Sigma-Aldrich) was added prior to storage at −20 °C.
A sandwich IgA enzyme-linked immunosorbent assay (Bethyl Laboratories Inc., Montgomery, TX, USA) was used to quantify the IgA concentration of piglet feces collected in 2006. Briefly, an ELISA plate was coated with goat anti-pig IgA antibody and incubated at room temperature. After washing, standards and samples were added to designated wells in duplicate. Following further washings, goat anti-Pig IgA HRP-conjugated antibody was added to each well as a detection antibody. Following a further incubation period, tetramethyl-benzidine enzyme substrate (3,3′,5,5′-tetramethyl-benzidine) was added to each well. After 5 min, the reaction was stopped with 2 M H2SO4 and individual well absorbencies were determined at 450 nm.
Pig reference serum (Bethyl Laboratories Inc.) was used to construct a standard curve from which the IgA concentration of the fecal extracts could be determined. The optimal dilution for each fecal extract was determined and using this dilution, the samples were independently diluted and assayed twice. The results were expressed as microgram of IgA per gram of (dry weight) feces and standard error of the mean (s.e.m.) was calculated. Reproducibility was assessed by extracting a single fecal sample three times and assaying each extract in duplicate.
Results
Sample collection and reproducibility
A total of 174 fecal samples were collected from 35 piglets over 3 trials (see Supplementary Data, Supplementary Tables S1 and S2). Collection of samples on a specific day was difficult, particularly so for younger piglets where defecation was infrequent. For most piglets in this study, between three and six samples were collected at approximately weekly intervals. To assess temporal changes at a finer scale, one piglet from trial 3 was intensively monitored with a total of 25 samples collected at intervals ranging from 4 h to 3 days.
The immune system of piglets undergoes significant postnatal development and has been suggested as a factor influencing gut community structure (Bailey et al., 2001). To confirm that the sampling period was sufficient to observe effects of immunological development, we assayed for IgA content in fecal samples collected in the third trial in 2006 (storage of the 2004 samples affected assay reproducibility). Among the 11 sampled piglets, fecal IgA levels showed a characteristic profile, whereby moderate levels were present initially (days 3–7), low to very low levels from days 8 to 21, and moderate-to-high levels from 22 to 36 days (refer to Supplementary Figure S1). This is consistent with the gradual clearance of maternal IgA ingested during the first 3 days, followed by development of the capacity to secrete IgA into the intestinal lumen after about 21 days of age. We conclude that the duration of the sample period (36 days) is sufficient to observe the development and consequences of immunological cross talk between the host and the microbiota.
To determine that differences in banding profiles represented changes to the gut microbial community and not technical artifacts, the reproducibility of DNA extraction and DGGE analysis of fecal samples was assessed. Following sample homogenization in TE buffer, subsamples of fecal homogenate were independently extracted and then these DNA templates independently amplified. DGGE analysis of PCR products derived from the same original fecal sample gave banding patterns that were indistinguishable when loaded on the same gel and analysed using Quantity One software (data not shown). We conclude that the DNA extraction and PCR amplification gives a reproducible sample of the fecal assemblage. On any one gel up to 22 samples could be reliably compared.
Piglet gut microbiota shows temporal variation and individuality
It is important to note that observable changes to the DGGE band profiles primarily reflect changes in which populations have highest relative abundance within the sample, rather than changes in absolute presence/absence from the sample (Muyzer and Smalla, 1998; Zoetendal et al., 1998). Temporal variation in an individual was assessed by pairwise comparisons of the DGGE fingerprint of each fecal sample to its adjacent samples in the temporal sequence from the same pig. For the majority of pigs in the study, samples were only available at weekly intervals and very little evidence of increased stability over time was seen. For example, in a set of five piglets whose communities were sampled at weekly intervals, the average similarity between successive community samples from the same piglet over 5 weeks (61.3%±6%) was not significantly different from the average similarity between piglets at 5 weeks of age (56%±6%). This suggests that either pigs do not show an increase in relative community stability over time or that the increase does not occur until after 4 weeks. To explore this further, one pig (no. 25) was intensively sampled in the third trial. Analysis of this sample set showed that prior to 28 days of age, the average pairwise similarity between two adjacent time points (66.7%±14.8%) was comparable to the temporal variation seen with weekly samples in other pigs. In stark contrast, the pairwise comparisons between fingerprints of adjacent samples collected between day 31 and day 36 showed average similarity of 94.5% with a range of 85.7–100. This period coincided with the consistent presence of high levels of IgA in the fecal samples (Figure 1). We conclude that within an individual, the intestinal microbiota shows temporal variation up to 28 days, a significant increase in stability beyond 30 days and that weekly samples are sufficient to resolve major community dynamics.
Fecal IgA concentrations and bacterial community structure in an individual piglet over days 4–36. (a) Temporal change in IgA concentration (diamonds) and community structure (squares) over time. IgA levels were determined by enzyme-linked ImmunoSorbent assay; IgA detected before 10 days is likely to be mostly derived from maternal fluids ingested during the first 3 days and IgA after 22 days to be produced by the piglet. Bacterial community turnover at each day is the pairwise similarity between the community fingerprint of that day's sample and the preceding sample. (b) Image of the DGGE fingerprints of the bacterial community in each sample.
At the end of the sampling period (36 days), all individuals had a unique community fingerprint. The average community fingerprint similarity between cohabiting piglets was 62%, compared to 51% for piglets housed in separate pens. Comparisons between siblings or non-siblings, irrespective of housing, also showed average similarities of approximately 50% (Table 1). This difference between cohabiting piglets and all other cohorts was significant in unpaired two-tailed Student's t-test (P<0.05). The piglets included in this study were separated from their mothers at the age of 3 days and housed in a controlled environment for the duration of the sampling period. This observation implies that the shared environmental exposure of cohabitants during the sampling period had a greater impact on the outcomes of colonization than the shared maternal environment of siblings during the first 3 days. We explored this phenomenon further by examination of average pairwise similarity between cohorts of cohabiting/noncohabiting, or sibling/non-sibling, with samples collected at different temporal sampling points.
Interindividual differences are present at 1 week of age independent of environmental or maternal relationship
We looked for evidence of maternal correlation during the early stages of community establishment, as piglets from the same litter are predicted to have received a similar maternally derived inoculation at birth. In the samples collected at 1 week, there was no significant difference between piglets grouped by housing arrangement or maternal relationship (Table 1). The most notable observation was that the average similarity between individuals at this time point (28.5%) was significantly lower than that for any other set of samples in the study. If samples were grouped by temporal relationship (that is, comparison of all samples collected at the same time point) then average interindividual community fingerprint similarity ranged from 38.3% to 57.0% over 2–5 weeks (Table 1). Therefore, at 1 week of age, an individual's gut microbiota is apparently more distinctive than at any other point in its life. When taken together with the lack of correlation between maternally related or housing-related communities, this suggests that at 1 week of age, the intestinal microbiota of all individuals in our study has developed essentially and independently.
Community structure developed significant environmental dependence within a defined temporal window of piglet development
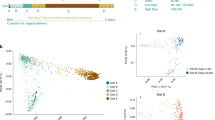
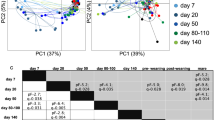
Comparative analyses over time of piglet fecal samples grouped by environmental relationship (cohabiting/non-cohabiting piglets) or maternal relationship (sibling/non-sibling) showed a remarkable pattern, whereby the pairwise similarity of cohabiting piglets was sharply increased in 3 or 4 weeks relative to observations in weeks 1 and 2, and remained high through to the end of the study at week 5 (Figure 2). When cluster analyses of all community fingerprints at each time point were performed, the impact of this ‘cohabitation effect’ was most obvious in the week 3 and 4 sample sets, where the majority of cohoused pairs comprised a distinct clade in dendrograms (Figure 2). To test the significance of this, we tabulated all available pairwise fingerprint comparisons. In unpaired two-tailed student's t-test, the average similarity of fingerprints from cohabiting piglets was significantly greater than those from non-cohabiting piglets or siblings at weeks 3 (P<0.0001), 4 (P<0.0001) and 5 (P<0.05) (Table 1). This implies that the community of each pig follows a distinct trajectory that reflects a bottleneck after week 2 and before week 4 (see Supplementary Figure S3 for a timeline). The outcome of this community development is strongly dependent on the immediate environment experienced after separation from the mother. In the intensively sampled pig, an unusually large and sudden shift in community composition was observed to occur between days 17 and 20. Increasing participation of the host in regulation of microbial community structure by either selective tolerance or intolerance of populations (immunological development), or selective encouragement of microbial populations by expression of growth or adhesion factors (gut tissue development) are plausible candidates for this type of interaction (Bry et al., 1996).
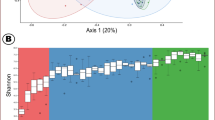
(a) DGGE profiles of bacterial communities in fecal samples obtained from 3-week-old piglets. Above each lane, similarly patterned dots indicate pigs derived from the same litter and brackets indicate cohabiting piglets. A schematic representation of all recorded bands and their assigned mobilities is shown in Supplementary Figure S2. (b) Dendrogram of community DGGE fingerprint similarities between 3-week-old piglets. Dendrogram was constructed by the unweighted pair group method using arithmetic averages (UPGMA). Cohabiting piglets are indicated by brackets. The maximal observed pairwise similarity for each piglet and sample point, where this was seen, are indicated. Pairs of piglets where the maximum was >85% for at least one sample point are indicated by shading.
Discussion
It is useful to conceptually separate the development of the gut community into two stages with a transitional phase in between. In this model, the first stage is essentially a microbial ecosystem in which the role of the piglet host is predominantly in setting physicochemical environmental parameters. The second stage is conceptually a different type of ecosystem where the gut microbiota are most usefully considered as part of a ‘super-organism’ and the role of the piglet has expanded from host to participant. It is this second stage that most directly leads to the stable gut microbiome of an adult pig.
The gut community of young piglets has attributes of a predominant microbial ecosystem: it is dynamic and independently acquired in each piglet
Colonization of the mammalian gut commences at birth. Immediately following birth, the gastrointestinal tract and oral cavities of infants contains bacteria, mostly derived from the mother (Bettelheim et al., 1974; Brook et al., 1979). There are presumably few or no barriers to microbes from other environments that are rapidly colonizing the infant at this stage. It is assumed that an individual neonatal gut community will be a subset of a very much larger meta-community comprised of all those bacterial groups, capable of colonizing the infant gut. It has been predicted that communities in physically similar environments, such as the intestines of the same host species, will have different compositions if they are formed at random from a larger seeding community (Curtis and Sloan, 2004). Our data are consistent with this model showing a high level of individuality in 1- and 2-week-old piglets (Table 1). This suggests that there is considerable randomness to the process of acquiring microbes. The resultant individuality of gut communities has the potential to influence the health and well being of the host. In adult rats or mice, differences between the microbiota of individuals have been predicted or identified as a contributing factor in metabolic and physiological variations between individuals, such as metabolite profiles (Robosky et al., 2005), obesity (Ley et al., 2005; Turnbaugh et al., 2006) and drug response (Clayton et al., 2006). However, in these examples, the phenotypes are inferred to reflect comparatively stable interindividual differences of mature gut microbiotas. In the very young piglets, the gut community was also highly dynamic. The important issue here is how do the individual differences become established? It is in this context that our observation of the ‘cohabitation effect’ is highly relevant.
The cohabitation effect reflects the onset of transition from a microbial ecosystem to a super-organism ecosystem
In essence, the cohabitation effect reflects that cohoused piglets suddenly develop very similar communities. One possible explanation is behavioural. Coprophagy would result in sudden exchange of microbiotas by penmates and if extensive, the ingested microbiota could conceivably result in ‘swamping’ of the community fingerprints. However, this would require the behavioural trait suddenly emerging in 3-week-olds simultaneously, and in hundreds of hours of the piglets’ observation, no instances of coprophagy were observed. Our favoured explanation is that a host response triggers the change in microbial community structure by engineering an environmental change that selectively advantages a subset of the community. This fits our conceptual model of a transitional period in the development of the host–intestinal microbiota association. At the age at which the cohabitation effect is observable (approximately 21 days), the community is still dynamic and a stable host–microbiota association has not formed. However, the community turnover in piglets at this age is nonrandom. The selective nature of the community change implies that the host response must somehow be informed by interaction with the microbiota. The persistence of a significant relationship between cohabitation and community structure in older piglets (>30 days) with a relatively stable community structure implies that the changes that occur in this transitional period will have long-term (possibly life-long) consequences.
Host factors influencing community turnover at transition will include the developing immune system
In all pigs examined, a community transition occurred after week 2 (sample points on days 14–18) and before week 4 (samples on days 28–30). Our observations, and available data on the maturation of the pig immune system, are consistent with this, reflecting a window in immune development. In piglets the key elements of the induction system in the mucosa develop over the first 2 weeks after birth and an almost adult organization of Peyer's patches is typically present by 12 days (Bailey et al., 2001). The cohabitation effect is always observed after this. The final stage of pig immune system maturation is thought to start around 28 days of age when the CD8+ T cells infiltrate the intestinal tissue and the mucosal immune system has a largely adult architecture by 6 weeks of age (Bailey et al., 2001). At this age, IgA-secreting plasma cells are also present in the intestine (Bianchi et al., 1999). In our study, IgA synthesis in piglets was established by 28 days (moderate to high levels in all nine sampled pigs) and not evident in the 21-day samples (Supplementary Figure S1). Thus, the onset of the cohabitation effect is predicted to occur after the development of the induction components of the mucosal immune system and evidently before the maturation of IgA secretion capacity. This suggests that the full response capacity of the immune system is not necessary to affect the community changes underlying the cohabitation effect.
Piglets are born with little or no passive immunity, as there is no transfer of antibodies in utero, but they do acquire maternal antibody from colostrum and milk (Westrom et al., 1984). It has been suggested that in mice, the lag time between the decline in IgA levels at weaning and the maturation of mucosal immune system allows an increase in bacterial colonization (Inoue et al., 2005a). We observed detectable levels of fecal IgA in the earliest samples (3–7 day-olds). This is almost certainly maternal IgA ingested by our piglets during the first 3 days. In samples collected from 8- to 21-day-old piglets, IgA was either not detectable or very low (Supplementary Figure S1). This probably reflects a combination of clearance of ingested maternal IgA combined with little or no synthesis of IgA. Both the week 2 and 3, the sample sets fall within the middle of this low-IgA period. The absence of the cohabitation effect from the week 2 samples suggests that low levels of maternal IgA are not sufficient by themselves to give rise to the community change. The later appearance of the cohabitation effect, after the induction system, is likely to be developed, suggesting the capacity to process antigens is also an important requirement. The most difficult aspect of the phenomenon to explain is that different organisms dominated the community, in independently housed pairs of piglets. We consider it highly unlikely that the same organisms newly acquired from the immediate external environment could simultaneously become dominant members of the microbiota in different pigs. We postulate that the community change that underlies the cohabitation effect reflects the first ‘decisions’ by the host as to tolerance/nontolerance of microbial antigens and the primary microbial beneficiaries (in terms of population increase) are organisms that have somehow been ‘seen’ (for example, low-abundance members of both communities) by both pigs prior to this point.
Environmental impacts on early microbial colonization of the gastrointestinal tract potentially have long-lasting impacts on the host
The stability of the gut community in adults provides the host with a life-long phenotype and this phenotype may affect an individual's response to some medical treatments and propensity towards disease (Backhed et al., 2004; Clayton et al., 2006; Ley et al., 2006). It has been proposed that the establishment of this gut community represents an opportunity to engineer specific outcomes for the adult host phenotype, as well as a period of risk when disturbance could lead to undesirable outcomes (Ley et al., 2006). Further support for the practical value of understanding this process comes from observations that outcomes of inoculation with probiotic strains are age dependent (Inoue et al., 2007). Our study illustrates that processes that occur during a relatively short developmental window in pigs have a significant effect on the nature of the stable community that appears after 5 weeks. The implication is that at the ages of 2–3 weeks in pigs (and corresponding developmental periods in other animals), the gut community is likely to be relatively more amenable to engineering and perhaps also more susceptible to disturbance than at other ages.
References
Alam M, Midtvedt T, Uribe A . (1994). Differential cell kinetics in the ileum and colon of germfree rats. Scand J Gastroenterol 29: 445–451.
Backhed F, Ding H, Wang T, Hooper LV, Koh GY, Nagy A et al. (2004). The gut microbiota as an environmental factor that regulates fat storage. Proc Natl Acad Sci USA 101: 15718–15723.
Bailey M, Plunkett FJ, Rothkotter HJ, Vega-Lopez MA, Haverson K, Stokes CR . (2001). Regulation of mucosal immune responses in effector sites. Proc Nutr Soc 60: 427–435.
Bettelheim KA, Breadon A, Faiers MC, Ofarrell SM, Shooter RA . (1974). Origin of O serotypes of Escherichia coli in babies after normal delivery. J Hyg 72: 67–70.
Bianchi AT, Scholten JW, Moonen Leusen BH, Boersma WJ . (1999). Development of the natural response of immunoglobulin secreting cells in the pig as a function of organ, age and housing. Dev Comp Immunol 23: 511–520.
Brook I, Barrett CT, Brinkman CR, Martin WJ, Finegold SM . (1979). Aerobic and anaerobic bacterial flora of the maternal cervix and newborn gastric fluid and conjunctiva: a prospective study. Pediatrics 63: 451–455.
Bry L, Falk PG, Midtvedt T, Gordon JI . (1996). A model of host–microbial interactions in an open mammalian ecosystem. Science 273: 1380–1383.
Clayton TA, Lindon JC, Cloarec O, Antti H, Charuel C, Hanton G et al. (2006). Pharmaco-metabonomic phenotyping and personalized drug treatment. Nature 440: 1073–1077.
Curtis TP, Sloan WT . (2004). Prokaryotic diversity and its limits: microbial community structure in nature and implications for microbial ecology. Curr Opin Microbiol 7: 221–226.
Eckburg PB, Bik EM, Bernstein CN, Purdom E, Dethlefsen L, Sargent M et al. (2005). Diversity of the human intestinal microbial flora. Science 308: 1635–1638.
Favier CF, Vaughan EE, De Vos WM, Akkermans ADL . (2002). Molecular monitoring of succession of bacterial communities in human neonates. Appl Environ Microbiol 68: 219–226.
Gustafsson BE . (1959). Vitamin K deficiency in germfree rats. Ann NY Acad Sci 78: 166–174.
Harmsen HJM, Wildeboer-Veloo ACM, Raangs GC, Wagendorp AA, Klijn N, Bindels JG et al. (2000). Analysis of intestinal flora development in breast-fed and formula-fed infants by using molecular identification and detection methods. J Pediatr Gastroenterol Nutr 30: 61–67.
Inoue R, Nishio A, Fukushima Y, Ushida K . (2007). Oral treatment with probiotic Lactobacillus johnsonii NCC533 (La1) for a specific part of the weaning period prevents the development of atopic dermatitis induced after maturation in model mice, NC/Nga. Br J Dermatol 156: 499–509.
Inoue R, Otsuka M, Ushida K . (2005a). Development of intestinal microbiota in mice and its possible interaction with the evolution of luminal IgA in the intestine. Exp Anim 54: 437–445.
Inoue R, Tsukahara T, Nakanishi N, Ushida K . (2005b). Development of the intestinal microbiota in the piglet. J Gen Appl Microbiol 51: 257–265.
Konstantinov SR, Awati AA, Williams BA, Miller BG, Jones P, Stokes CR et al. (2006). Post-natal development of the porcine microbiota composition and activities. Environ Microbiol 8: 1191–1199.
Leser TD, Amenuvor JZ, Jensen TK, Lindecrona RH, Boye M, Moller K . (2002). Culture-independent analysis of gut bacteria: the pig gastrointestinal tract microbiota revisited. Appl Environ Microbiol 68: 673–690.
Ley RE, Backhed F, Turnbaugh P, Lozupone CA, Knight RD, Gordon JI . (2005). Obesity alters gut microbial ecology. Proc Natl Acad Sci USA 102: 11070–11075.
Ley RE, Peterson DA, Gordon JI . (2006). Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 124: 837–848.
Mai V, Katki HA, Harmsen H, Gallaher D, Schatzkin A, Baer DJ et al. (2004). Effects of a controlled diet and black tea drinking on the fecal microflora composition and the fecal bile acid profile of human volunteers in a double-blinded randomized feeding study. J Nutr 134: 473–478.
Mazmanian SK, Liu CH, Tzianabos AO, Kasper DL . (2005). An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 122: 107–118.
Moore WE, Holdeman LV . (1974). Human fecal flora: the normal flora of 20 Japanese-Hawaiians. Appl Microbiol 27: 961–979.
Muyzer G, Smalla K . (1998). Application of denaturing gradient gel electrophoresis (DGGE) and temperature gradient gel electrophoresis (TGGE) in microbial ecology. Antonie Van Leeuwenhoek 73: 127–141.
Nubel U, Engelen B, Felske A, Snaidr J, Wieshuber A, Amann RI et al. (1996). Sequence heterogeneities of genes encoding 16S rRNAs in Paenibacillus polymyxa detected by temperature gradient gel electrophoresis. J Bacteriol 178: 5636–5643.
Palmer C, Bik EM, DiGiulio DB, Relman DA, Brown PO . (2007). Development of the human infant intestinal microbiota. Plos Biology 5: 1556–1573.
Robosky LC, Wells DF, Egnash LA, Manning ML, Reily MD, Robertson DG . (2005). Metabonomic identification of two distinct phenotypes in Sprague–Dawley (Crl:CD(SD)) rats. Toxicol Sci 87: 277–284.
Salyers AA, West SE, Vercellotti JR, Wilkins TD . (1977). Fermentation of mucins and plant polysaccharides by anaerobic bacteria from the human colon. Appl Environ Microbiol 34: 529–533.
Sambrook JR, Russell DW . (2001). Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Press: New York, pp A9.5–A9.7.
Simpson JM, Kocherginskaya SA, Aminov RI, Skerlos LT, Bradley TM, Mackie RI et al. (2002). Comparative microbial diversity in the gastrointestinal tracts of food animal species. Integr Comp Biol 42: 327–331.
Simpson JM, McCracken VJ, White BA, Gaskins HR, Mackie RI . (1999). Application of denaturant gradient gel electrophoresis for the analysis of the porcine gastrointestinal microbiota. Journal of Microbiological Methods 36: 167–179.
Smith HW . (1965). The development of the flora of the alimentary tract in young animals. J Pathol Bacteriol 90: 495–513.
Stappenbeck TS, Hooper LV, Gordon JI . (2002). Developmental regulation of intestinal angiogenesis by indigenous microbes via Paneth cells. Proc Natl Acad Sci USA 99: 15451–15455.
Stark PL, Lee A . (1982). The microbial ecology of the large bowel of breast-fed and formula-fed infants during the first year of life. J Med Microbiol 15: 189–203.
Swanson SP, Nicoletti J, Rood Jr HD, Buck WB, Cote LM, Yoshizawa T . (1987). Metabolism of three trichothecene mycotoxins, T-2 toxin, diacetoxyscirpenol and deoxynivalenol, by bovine rumen microorganisms. J Chromatogr 414: 335–342.
Tannock GW . (2005). Commentary: remembrance of microbes past. Int J Epidemiol 34: 13–15.
Tannock GW, Munro K, Bibiloni R, Simon MA, Hargreaves P, Gopal P et al. (2004). Impact of consumption of oligosaccharide-containing biscuits on the fecal microbiota of humans. Appl Environ Microbiol 70: 2129–2136.
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI . (2006). An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444: 1027–1031.
Westrom BR, Svendsen J, Ohlsson BG, Tagesson C, Karlsson BW . (1984). Intestinal transmission of macromolecules (Bsa and Fitc-labeled dextrans) in the neonatal Pig—influence of age of piglet and molecular-weight of markers. Biol Neonate 46: 20–26.
Yeates C, Gillings MR, Davison AD, Altavilla N, Veal DA . (1998). Methods for microbial DNA extraction from soil for PCR amplification. Biol Proced Online 1: 40–47.
Yoshioka H, Iseki K, Fujita K . (1983). Development and differences of intestinal flora in the neonatal period in breast-fed and bottle-fed infants. Pediatrics 72: 317–321.
Zoetendal EG, Akkermans ADL, De Vos WM . (1998). Temperature gradient gel electrophoresis analysis of 16S rRNA from human fecal samples reveals stable and host-specific communities of active bacteria. Appl Environ Microbiol 64: 3854–3859.
Acknowledgements
We acknowledge support from the NHMRC and AJH from the Selby Research Trust. The assistance of Muhsin Karim in the care of the piglets and collection of samples is gratefully acknowledged. CT is the recipient of an Australian Postgraduate Award (APA).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Rights and permissions
About this article
Cite this article
Thompson, C., Wang, B. & Holmes, A. The immediate environment during postnatal development has long-term impact on gut community structure in pigs. ISME J 2, 739–748 (2008). https://doi.org/10.1038/ismej.2008.29
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2008.29
Keywords
This article is cited by
-
Parity changed fecal microbiota of sows and its correlation with milk long-chain fatty acid profiles
Applied Microbiology and Biotechnology (2024)
-
Cefquinome shows a higher impact on the pig gut microbiome and resistome compared to ceftiofur
Veterinary Research (2023)
-
Age influences the temporal dynamics of microbiome and antimicrobial resistance genes among fecal bacteria in a cohort of production pigs
Animal Microbiome (2023)
-
An insight into the commercial piglet’s microbial gut colonization: from birth towards weaning
Animal Microbiome (2022)
-
Immunohistochemical visualisation of the enteric nervous system architecture in the germ-free piglets
Journal of Molecular Histology (2022)