Abstract
Although invasive plants can drive ecosystem change, little is known about the directional nature of belowground interactions between invasive plants, native roots, bacteria, archaea and fungi. We used detailed bioinformatics and a recently developed root assay on soils collected in fescue grassland along a gradient of smooth brome (Bromus inermis Leyss) invasion to examine the links between smooth brome shoot litter and root, archaea, bacteria and fungal communities. We examined (1) aboveground versus belowground influences of smooth brome on soil microbial communities, (2) the importance of direct versus microbe-mediated impacts of plants on soil fungal communities, and (3) the web of roots, shoots, archaea, bacteria and fungi interactions across the A and B soil horizons in invaded and non-invaded sites. Archaea and bacteria influenced fungal composition, but not vice versa, as indicated by redundancy analyses. Co-inertia analyses suggested that bacterial–fungal variance was driven primarily by 12 bacterial operational taxonomic units (OTUs). Brome increased bacterial diversity via smooth brome litter in the A horizon and roots in the B horizon, which then reduced fungal diversity. Archaea increased abundance of several bacterial OTUs, and the key bacterial OTUs mediated changes in the fungi’s response to invasion. Overall, native root diversity loss and bacterial mediation were more important drivers of fungal composition than were the direct effects of increases in smooth brome. Critically, native plant species displacement and root loss appeared to be the most important driver of fungal composition during invasion. This causal web likely gives rise to the plant–fungi feedbacks, which are an essential factor determining plant diversity in invaded grassland ecosystems.
Similar content being viewed by others
Introduction
Invasive plant species are important drivers of ecosystem change through their influence on belowground microbial communities (van der Putten et al., 2001). Soil microbial interactions are critical determinants of aboveground diversity (Klironomos, 2002), yet little is known about how invasion alters the directional nature of interactions among microbial groups (although see Schröter et al., 2004; Scherber et al., 2010). For example, the switch from negative plant–soil microbial feedbacks in native habitats to positive feedbacks in invaded habitats may be crucial for invasive plant success (Callaway et al., 2004; Diez et al., 2010). How this switch occurs is still open to debate; however, it is likely linked to aboveground carbon supply (Bradley et al., 2006) or changes in the composition and structure of soil microbial communities and altered nutrient cycling (Elgersma et al., 2011; Shannon-Firestone et al., 2015).
Several recent studies suggest that plant invasion success is linked to suppression and/or promotion of microbial species (Bever et al., 2012; van der Putten et al., 2013). For example, plant invasion may selectively suppress dominant bacterial species, allowing for rare and/or ammonia-oxidizing bacteria (AOB) to increase in relative abundance, which in turn has an impact on nitrogen cycling (Ehrenfeld, 2010; Piper et al., 2015a). However, plant effects are not constant among plant species. In particular, some invasive species may decrease fungal abundance (Vogelsang and Bever, 2009), thus increasing the bacteria to fungi ratio in the soil; other species may increase particular fungal groups (Kourtev et al., 2002). Temporal lags between invasion and accumulation of species-specific pathogens in the soil may also allow invasive plants to flourish at the expense of native species (Klironomos, 2002), although in some instances pathogenic fungi may prevent invasion (Reinhart et al., 2005). Despite the clear importance of plant–microbial interactions during invasion, the causal chain linking changes in aboveground to belowground diversity and the evolution of plant–microbial feedbacks is not clear. Some authors suggest that plant effects are dampened as they percolate through soil trophic levels (Scherber et al., 2010), whereas others suggest that plant effects are limited to specialist microbial groups (van der Putten et al., 2007).
Fungal communities are critical in any consideration of plant–soil feedbacks, as fungi appear to determine vegetation composition through structuring plant–plant interactions (Bennett and Cahill, 2016) and plant abundance (Reinhart et al., 2005). However it is not clear how directional relationships among fungi, bacteria and archaea evolve during invasion. Fungi may metabolize root exudates, and those metabolites then drive bacterial community composition (Millard and Singh, 2009) or, alternatively, bacteria can suppress fungal pathogens (Kobayashi and Crouch, 2009). Relative to bacteria, much less is known about the relationship of archaea with other microbes. We know archaea are common, although not major, inhabitants of roots (Buée et al., 2009, although see Simon et al., 2000 for an exception), and in some instances fungi may reduce ammonia-oxidizing archaea and AOB abundance via plant root pathways (Chen et al., 2013), although often there is no relationship between archaeal and fungal diversity at broad spatial scales (Prober et al., 2015). This is likely because archaea, unlike fungi, typically respond more to soil and environmental properties than to plant cover (Nielsen et al., 2010). Thus, from a plant–microbe perspective we may conceptualize microbial community responses to plants at a field scale as a gradient, where fungi respond mainly to vegetation and only slightly to soil, bacteria respond strongly to both vegetation and soil, and archaea respond mainly to soil and not to vegetation. However, there is still considerable uncertainty in how the structure of intermicrobe interactions changes in communities undergoing plant invasion.
Smooth brome (Bromus inermis Leyss) invasion lowers plant species diversity in both mixed-grass prairie (Fink and Wilson, 2011) and fescue grasslands (Wilson and Pinno, 2013). This invasion alters soil conditions (that is, increased nitrogen availability; Piper et al., 2015b), thereby initiating a cascade of soil community changes, including the suppression of dominant members of the bacterial community (Piper et al., 2015a). However, no one has conclusively identified an above- or belowground mechanism linking brome invasion to bacterial diversity. Rather, structural equation models (SEM) have indicated that an unknown direct pathway driven by smooth brome abundance is responsible for the changes in bacterial diversity (Piper et al., 2015a). One hypothesis is that the unknown mechanism may result from the interactions between bacterial and fungal communities (Frey-Klett et al., 2011). These bacterial–fungal interactions could be specific pathogen interactions (for example, as outlined in van der Putten et al., 2007); however, there is some evidence that more general links between the fungal and bacterial communities may have a major role in structuring microbial communities (for example, Siciliano et al., 2014).
Here we present a model to study interkingdom interactions and responses to plant invasion in a smooth brome-invaded fescue grassland. Using samples from the smooth brome invasion gradient, we assessed how the structure of the aboveground plant and its root communities interact with belowground archaea, bacteria and fungi to influence soil ecosystem composition. We hypothesized that smooth brome-structured fungi via roots (cf. Johnson et al., 2004) and that, in turn, the fungi affected other microbes. To address this hypothesis, we asked a series of linked questions: (1) does the influence of invasive smooth brome on soil microbial communities differ between aboveground litter and belowground root effects? (2) What is the relative importance of direct versus microbe-mediated impacts of invasive smooth brome on soil fungal communities? (3) Can we determine the directional nature of the web of roots, litter, archaea, bacteria and fungi interactions across A and B horizons of soils in invaded and non-invaded sites?
Materials and methods
Study area
The field site is a 14.6 ha native fescue prairie undergoing invasion by smooth brome, ~120 km south of Saskatoon, Saskatchewan, Canada (51°12′N, 107°17′W). The site is near the border of the moist mixed and mixed prairie ecoregions and within the Orthic Dark Brown Chernozemic soil order (Soil Classification Working Group, 1998). The grasses Festuca hallii and several species of Hesperostipa, Elymus and Pascopyrum dominate the native prairie at this site. Smooth brome is invading the site from disturbed edges (primarily roads), and many invaded patches can be found in the interior of the site (Piper et al., 2015a, 2015b). At the time of sampling there were no cattle grazing on the site or other management practices applied. The mean annual temperature is 3.5 °C, and 376.9 mm of precipitation falls each year (Rock Point weather station, ~7 km from the field site, Environment Canada, 2016).
Sample collection
We collected plant and soil samples from a broad range of smooth brome cover classes (Piper et al., 2015a, b). A total of sixty 50 × 50 cm plots were established in a stratified random manner. To ensure even sampling across the full range of the brome invasion (0, >0–50, 51–85 and >85%), each plot had 14–16 samples in each of four classes of smooth brome shoot abundance. Sampling locations were determined using the random point generator in ArcMap (ESRI, Redlands, CA, USA). Within each plot, we assessed plant species cover and gathered grass, forb, shrub and litter biomass. Smooth brome biomass was collected separately from other grass species.
Biomass samples were dried for 2 days at 60 °C and weighed. We measured the depth of the A horizon (defined as mineral soil with less than 17% organic carbon) using changes in soil color and texture (Soil Classification Working Group, 1998). Using an AMS soil corer (AMS, Inc., American Falls, ID), we extracted two 5-cm-diameter soil cores from the top 5 cm of each of the A and B horizons at each plot. To minimize cross contamination, A and B horizon samples were collected and analyzed separately because (1) soil horizons are created by fundamentally different soil-forming processes, and (2) ammonia-oxidizing microbes respond differently to smooth brome abundance in each horizon (Soil Classification Working Group, 1998; Piper et al., 2015b). The two soil cores from each horizon were then combined, resulting in one composite soil core from each horizon at each plot. These samples were frozen at –20 °C until they could be analyzed.
To characterize roots, we used a recently developed assay capable of identifying and quantifying plant roots from DNA extracts of mixed-species root samples (Hiiesalu et al., 2012; Lamb et al., 2016; see Appendix S3 in Supplementary Information). Briefly, we modified the trnL primers used previously by other researchers (Taberlet et al., 1991; 2007) and used an improved bioinformatics pipeline that referenced site-specific taxon lists to identify plants to a species level (for detailed information on the approach, see Lamb et al., 2016). Bacterial, archaeal and fungal community compositions were assessed by high-throughput amplicon library sequencing, as described in Piper et al. (2015a) and Siciliano et al. (2014). Briefly, DNA was extracted from 0.5 g of 2-mm sieved soil using the Ultraclean Soil DNA Extraction Kit (MoBio, Carlsbad, CA, USA). DNA concentration was determined using a ultraviolet–visible spectrophotometer (Nanodrop 2000, ThermoScientific, Wilmington, DE, USA). For bacteria, we used the universal 16 S rRNA primer set 515 F/806 R, which amplifies a 291-bp fragment near the bacterial v4 region (Caporaso et al., 2011). Primers contained an Ion Torrent adapter and a unique barcode sequence for sample pooling. Samples were amplified in triplicate using a 25-μl reaction mix containing 18 μl Platinum Blue Supermix (Invitrogen), and sample replicates were pooled and purified using a QiaQuick gel extraction kit (Qiagen Inc., Toronto, ON, Canada). Following pooling, the samples were sequenced according to the Ion PGM 200 Sequencing Kit v2 (Life Technologies). Similarly, fungal and archaeal diversity were assessed via triplicate reactions using ITS1F and ITS4 primers (Gardes and Bruns, 1993; Manter and Vivanco, 2007) for fungi (acceptable for differentiating fungi at the genera level and coarser taxonomic resolutions, cf. Krüger et al., 2009; Stockinger et al., 2009) and A344F and A589R primers for archaea (Teske and Sørensen, 2007). These amplicon libraries were sequenced from the forward primer on the 454 FLX titanium platform (Roche, Branford, CT, USA) at Genome Quebec. We note here, however, that the ITS1F and ITS4 primers may not effectively amplify arbuscular mycorrhizal fungi (Stockinger et al., 2010). Thus, we suggest our results be interpreted in terms of the overall microbial community, and not specifically the arbuscular mycorrhizal fungi community—which is important but beyond the scope of this study.
Bioinformatics processing
The 16 S gene is often used as a species diagnostic for bacteria and archaea (Stackebrandt and Goebel, 1994; Baker et al., 2003), although it has fewer hypervariable domains in fungi. Among the regions of the ribosomal cistron, the internal transcribed spacer (ITS) region has the highest probability of successful identification for the widest range of fungi (Schoch et al., 2012). Therefore, we used the 16 S and ITS sequences to identify and quantify bacterial and archaeal composition and fungal composition, respectively.
Bacterial and archaeal 16S
Data from the Ion torrent and 454 Platforms were processed for bacterial and archaeal 16 S rRNA gene analysis, respectively (see Appendix S2 in Supplementary Information). Seed sequences for each bacterial and archaeal operational taxonomic unit (OTU) were taxonomically identified using the GreenGenes database (August 2013 release) using a naive Bayesian classifier algorithm implemented in mothur (Wang et al., 2007; Schloss et al., 2009). We calculated two measures of alpha diversity: community evenness using Evar (Smith and Wilson, 1996) and observed richness as the total number of OTUs in each sample normalized to a specific number of reads per sample. In addition to these measures of alpha diversity, we calculated phylogenetic diversity for archaea and bacteria by considering the sequence data and the evolutionary relationships in the communities by implementing the UniFrac algorithm (Lozupone and Knight, 2005; Lozupone et al., 2011). The UniFrac approach incorporates evolutionary distance at the sequence level and provides a more robust estimate of community composition than OTU-based approaches. For fungal community composition, we were unable to use the UniFrac approach because the differing read lengths of the intragenic spacer region would bias UniFrac’s sequence-based analysis. Thus, we used principal component analysis to characterize the fungal OTU matrix and used the scores for principal component one as a metric of phylogenetic diversity.
Fungal ITS
ITS sequences were dereplicated and denoised, and chimeric sequences were removed. Remaining sequences (>200 bp) were further dereplicated, such that reads identical with longer reads were added to the longer read count (cluster fragments function in mothur). Next, we used the OTU picking function in QIIME (pick_otus.py with usearch61 and a 0.90 sequence similarity threshold) to generate a list of OTUs and pick_rep_set.py to establish a file of seed sequences for the selected OTUs (Caporaso et al., 2010). These OTUs were classified against the UNITE database (mothur format version –v6_sh_99) to provide a list of taxonomic identifications (Kõljalg et al., 2005). OTUs with identical classification were aggregated into single OTUs and their abundance summed using Primer6 (Clarke, 1993).
Statistical analyses
Overview
We used redundancy analysis (RDA) to identify significant covariates among microbial communities, followed by co-inertia analysis (CoIA) to identify keystone OTUs (addressing questions 1 and 2 in the introduction), and then confirmed these causal assumptions using SEM (question 3 and see SEM of plant, root and soil microbial relationships below).
Rare species
Species that occur in low quantities across a study site (few occurrences and low abundance) generate a large number of zeros in data matrices. Because zeros have high leverage, they contribute heavily to the total inertia of the data matrix (Legendre and Legendre, 2012). From one point of view, researchers may consider rare species as potential indicators of special environmental conditions. However, the primary purpose of ordination is to display the main axes of variation of the data, not to deal with exceptions. On the other hand, researchers may consider the occurrence of rare species as a chance event that should not be heavily favored in the analysis. To mitigate the bias of zero-inflated data, we used a stepwise elimination procedure to remove these species, which progressively downweighted and removed rare species (Figures 1a and b; see Supplementary Appendix S3 in Supplementary Information; Legendre and Legendre, 2012).
Sequence of statistical analyses used to assess functional relationships among smooth brome and soil microbial communities. (a) Distinct data matrices were used for archaea, bacteria, fungi, and explanatory covariates (bottom left) along a brome invasion gradient. (b) OTUs with small occurrence values only appear in a fraction of the sampling plots and generate a lot of zeros in the data matrices. Because zeros have high leverage and contribute heavily to the total inertia of the ordinations, we chose to eliminate rare species using a stepwise elimination procedure (Legendre and Legendre, 2012; see Materials and methods). (c) RDA was used to summarize significant linear relationships among microbial communities, and between communities and above- and belowground explanatory covariates. (d) Principal components analysis (PCA) was run on each microbial data matrix to find the maximal inertia projected on two axes. The reduction in dimensions of each data set resulted in independent structures for each community. CoIA tests for agreement between two structures (microbial and/or root communities) by calculating axes that maximize the covariance between the factorial scores of samples. (e) SEM uses two or more structural equations to model multivariate relationships. Here we use the results of the SEM analyses, based on prior experience and substantive knowledge gained from the previous statistical analyses, to reflect causal dependencies between smooth brome and soil microbial communities in a natural grassland under invasion by smooth brome.
We acknowledge that the topic of rare species removal is a contentious and unresolved issue (for example, Marchant, 1999; Cao et al., 2001) and recognize both the potential ecological significance of some rare species and statistical quagmire of rare species present due to chance (Lynch and Neufeld, 2015). Numerous methods exist for downweighting rare species (for example, Poos and Jackson, 2012), although many lack strong ecological foundations (see a discussion on this topic in Lynch and Neufeld, 2015). Therefore, we ran our RDA analyses on both the full and downweighted data matrices to evaluate the effect of rare species removal on variation in community composition. We found statistical relationships among microbial composition and environmental covariates were stronger (as indicated by higher F-statistics) after rare species removal (Figure 2, Supplementary Figure S4; compare F-values in Tables 1 and 2 of the downweighted data set to the corresponding Supplementary Tables S1 and S2, which are of the full data set). We interpreted the stronger pattern in the downweighted data as an increase in our signal-to-noise ratio, and a winnowing of the full microbial data matrices to only the most influential abundant species. Therefore, we focused all subsequent analyses on the downweighted data sets.
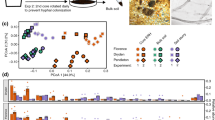
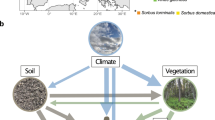
Conceptual model of aboveground (green arrows), belowground (brown and gray arrows), and interkingdom influences (large light blue arrows) on soil microbial communities along cover classes of smooth brome. Conceptual arrows representing aboveground, belowground, and interkingdom influences are based on significant explanatory variables as determined through each RDA. Biplots of RDAs for each kingdom ((a) bacteria, (b) fungi, and (c) archaea) are presented with significant explanatory variables (bold blue arrows within biplots; P⩽0.05). Box color in each RDA represents smooth brome cover classes and point shape represents A and B horizons (see legend at top of figure).
As another check of rare species influence, we assessed whether individual bacterial OTU abundance was associated with smooth brome cover, and whether these OTUs were associated with environmental covariates. To do this, we first assessed the relationship between OTU abundance and smooth brome cover via zero-inflated Poisson regression models (Zuur, 2009) followed by Bonferroni correction (Shaffer, 1995), and then ran an RDA using these correlated OTUs as covariates on the environmental data matrix. We found that 88 OTUs were strongly correlated with smooth brome cover (n=88) using a P<0.001 threshold. Of these 88 OTUs, only eight were correlated with the environmental matrix (P<0.05). These OTUs were from the phyla Acidobacteria, Actinobacteria, and Planctomycetes and represented a mean relative abundance of 0.29±0.03%, compared with 1.27±0.04% for the downweighted data OTUs. We included these eight OTUs as endogenous variables in our final SEM models with direct links between brome roots and shoots, and fungal evenness (see below). We found only two significant relationships: (1) smooth brome roots increased the abundance of a bacterial OTU of class Actinobacteria and (2) pH reduced the abundance of a bacterial OTU of class Kueneniae, which itself reduced fungal evenness. Thus, we present our downweighting–RDA–CoIA approach as a means to isolate larger-abundance influential consortia, and present this Poisson–RDA method as an alternative approach to identify influential conditionally rare taxa such as pathogens (see Appendix S4 in Supplementary Information and SEM results).
Soil microbial composition and keystone species
We identified significant covariates among belowground, aboveground, and interkingdom influences on microbial composition using RDA on Hellinger-transformed reduced data (Figure 1c). RDA is a method of summarizing linear relationships in a set of dependent variables influenced by a set of independent variables, using a blend of multiple linear regression and principal components analysis (Legendre and Legendre, 2012). RDA is a constrained ordination method that computes axes that are linear combinations of the explanatory variables (Borcard et al., 2011). Thus, the axes produced using RDA model the variation of the dependent variable, allowing one to test a hypothesis (H0) of absence of a linear relationship between microbial composition and explanatory independent variables (Figure 1c).
The following four groups of explanatory variables were used to model microbial composition (bacteria, archaea, and fungi): (1) soil influences (soil pH, soil organic carbon (SOC), and soil total nitrogen), (2) aboveground plant influences (litter nitrogen, smooth brome shoot abundance), (3) belowground plant influences (root biomass, root evenness, and root abundance of smooth brome), and (4) microbial community influences (ammonia-oxidizing archaea and AOB abundance; bacterial, fungal, and archaeal evenness, richness, and diversity [H′]; Piper et al., 2015a, b). Explanatory variables selected for retention were based on permutation tests (Borcard et al., 2011) and we used a permutational multivariate analysis of variance (PERMANOVA; Anderson, 2001) to test for significant differences among brome cover categories and soil horizon. We assumed a direction of influence based on the nature of explanatory variables. For example, we found bacterial richness was a significant predictor of fungal composition, but fungal diversity was not a predictor of bacterial composition. We interpreted this as a primarily unidirectional influence of bacteria on fungi. Though there may be exceptions to this, our statistical framework supported this directional nature and thus we maintained this direction of influence through subsequent analyses.
We implemented CoIA (Dolédec and Chessel, 1994) using the package ‘ade4’ version 1.7-2 in R (Dray and Dufour, 2007) to examine important correlations between kingdoms as revealed by the RDAs (Figure 1d). In CoIA, multivariate analyses like principal components analysis are processed on each community matrix separately to find the maximal inertia (that is, variance) projected onto two axes. The reduction in dimensions of each data set results in independent structures for each community. CoIA tests for agreement between these independent structures (microbial and/or root communities) by calculating axes that maximize the covariance between the factorial scores of samples. Here we first reduced the dimensionality of the archaeal, bacterial, and fungal matrices using Principal components analysis, and then selected the dominant components (axes). The CoIA then generated new axes by rotation in multidimensional space so as to maximize the covariance between the axes in the two data sets (Dolédec and Chessel, 1994). Two eigenvectors were selected from each kingdom matrix for inclusion into the CoIA. RV coefficients (a multivariate generalization of the Pearson correlation coefficient) were calculated for each CoIA to measure global correlations between matrices. The significance of the associations between kingdom matrices was tested with a Monte Carlo permutation test using 9999 permutations.
Preliminary analyses revealed significant correlations between site distances from the origin of the co-inertia plane and smooth brome cover for the soil A horizon and not the B horizon; therefore, we focused our analyses on the A horizon. Within the A horizon, we used t-tests followed by Bonferroni correction (Shaffer, 1995) to determine which plots contributed most to the common variance (total co-inertia) within the two matrices (that is, sites furthest from the plot origin), as well as the most important species driving the co-inertia (Figure 1d). We then visualized the configuration of the matrix pairs with joint site plots, in which arrows represent the distances between matrices in n-dimensional space and points represent the arrow midpoint (Legendre and Legendre, 2012).
SEM of plant, root, and soil microbial relationships
We used SEM (Grace, 2006; Lamb et al., 2011) to investigate a range of potential mechanisms underlying smooth brome–microbial community relationships (Figure 1e). Whereas many multivariate methods are largely descriptive and more appropriate for exploratory analyses, SEM is capable of testing a network of causal hypotheses and is recommended for the evaluation of multivariate hypotheses (Grace, 2006; Grace et al., 2012). Specifically, we used SEM because it allows the evaluation of simultaneous influences (for example, brome may influence fungal composition both directly and through altering bacterial diversity) rather than individual causes (for example, brome influences fungal composition only directly). The method is thus appropriate for establishing probable causality at the system (for example, brome–archaea–bacteria–fungi) rather than the individual level (for example, brome–fungi).
SEM rely on specification by researchers of a network of a priori causal assumptions based on a scientific body of evidence (for example, that pH is an important driver of bacterial richness and not that bacterial richness determines soil pH), and then testing whether that causal network is consistent with empirical data (Grace, 2006; Pearl, 2012; Shipley, 2016). The postulated causal network gives rise to a series of linear equations, which in turn give rise to an expected covariance matrix. The expected covariance matrix is then compared with the observed covariance matrix arising from the data. A statistically acceptable congruence between the expected (causal model implied) and observed covariance matrices is thus an empirical validation of the causal assumptions used. In other words, an SEM is a ecological theory describing a particular system, and if congruent with the data, the theory is supported. SEM is a well-established tool widely used in the natural and social sciences for testing causal inferences with correlative data sets; it is however critically important that the causal assumptions made by the researcher be well grounded in prior studies, scientific knowledge, logical arguments, and other evidence. An SEM model fitting the data does not prove the causal assumptions used, but replication of a given model across many systems represents a very strong test of the underlying theory (Grace et al., 2012).
The initial (theoretical) SEM used smooth brome shoot biomass and root abundance as measures of the above- and belowground severity of smooth brome invasion; it also used bacterial richness, evenness, and phylogenetic diversity as measures of bacterial diversity. Using RDA, we identified a directional link between archaeal diversity and both bacterial and fungal diversity (for example, Figure 2); thus, we included a directional pathway from archaea to bacteria and fungi in our SEMs. As pH is known to influence microbial community composition (Fierer and Jackson, 2006), we included direct relationships from pH to microbial diversity. Both competitive and mutualistic interactions exist between bacteria and fungi (de Boer et al., 2005). However, our RDA analyses suggested the influence of bacteria on fungi is greater than the reverse in this system, so we included a directional pathway from bacteria to fungi in our model.
We used multigroup SEMs with observed variables, as we collected two subsets of data (A and B horizon) from the same sample points (Grace, 2006; Piper et al., 2015a). In a multigroup SEM, models are initially constrained to ensure path coefficients are equal between groups (Grace, 2006; Lamb et al., 2011). Constraints are then progressively released to improve model fit. Different path coefficients between horizons suggest significant differences in the biological processes represented by that path between horizons. Models were fit using the ‘lavaan’ package v. 0.5–20 in R (Rosseel, 2012). In order to determine differences in biological processes between groups in the multigroup SEM, we initially constrained the models so that path coefficients were equal between horizons. Constraints were progressively released to improve model fit (cf. Piper et al., 2015a). Path coefficient differences between horizons indicated a significant distinction in the biological processes represented by that path between horizons. Before fitting the SEM, we used general linear models with a quadratic term to check for nonlinear relationships among exogenous and endogenous variables; no nonlinearities were found. To equalize variances we standardized variables by dividing raw values by their group maximum.
We constructed our SEM model stepwise. Our initial multigroup model assumed that the effects of smooth brome shoots and roots differed between horizons. We separated roots and shoots as they likely influence nutrient availability differently between soil horizons (Soil Classification Working Group, 1998). We then tested for horizon-specific effects of (1) pH on richness, evenness and phylogenetic diversity of archaea, bacteria, and fungi, (2) archaea on bacteria and fungi, and (3) bacteria on fungi. Improved model fit was assessed based on reduced χ2 and Akaike Information Criterion. There were no horizon-specific effects for pH on bacterial or fungal composition (Supplementary Tables S6–S8). Finally, we fit alternative models for each bacterial OTU that exerted a significant influence on fungal diversity with relaxed parameter constraints identical to those mentioned above.
In our first group of SEMs, we had three metrics of fungal diversity (richness, evenness, and phylogenetic diversity), and evaluated the following four separate causal assumptions: (1) no direct links between smooth brome roots or litter to fungal diversity, (2) direct links between brome litter and fungal diversity, (3) direct links between smooth brome roots and fungal diversity, and (4) direct links between both smooth brome litter, and roots and fungal diversity. Thus, we ran 12 separate models to test each of these potential causal assumptions (that is, 3 × 4; Supplementary Tables S6–S8). Once we found the best model configuration (as determined by Comparative Fit Index, Root Mean Square Error of Approximation, and Standardized Root Mean Square Residuals), we evaluated this model structure for each bacterial OTU of interest identified through the CoIA (n=12), as well as bacterial richness, evenness, and phylogenetic diversity (n=3) for each fungal diversity metric (n=3) for a total of 108 models (12 × 3 × 3). As an additional check on the direction of causality of the diversity measures, we inverted the causal link between fungi and bacteria, that is, that fungal richness, evenness, and phylogenetic diversity drove bacteria. For all three metrics, evenness, richness, and phylogenetic diversity, the χ2 was greater when bacteria were dependent on fungi. For example, for evenness, the χ2 increased from 5.4 to 8.9, for richness the χ2 increased from 14.7 to 21.3 and for phylogenetic diversity the χ2 increased from 6.0 to 6.1. The SEM analyses suggested three classes of relationships among smooth brome and microbial diversity based on frequency of microbial interactions. Specifically, if we found significant relationships between endogenous variables in more than one SEM, we designated this as a ‘class’. For brevity, we present a summary of the three model classes here and an example of each class in the supplementary material (Supplementary Tables S9–S17).
Results
Aboveground, belowground, and interkingdom predictors of microbial diversity
Removing rare OTUs from each microbial matrix resulted in a total of 82 archaeal (from 1339), 29 bacterial (from 6747), and 119 fungal (from 4014) OTUs used in subsequent analyses (Supplementary Figures S1–S3). The range of relative abundances following downweighting changed as follows: archaea: 0.001–1.170% to 0.418–2.666%, bacteria: 0.001–5.151% to 1.438–11.326%, and fungi: <0.001–2.607% to 0.067–3.424%.
Using RDA, we found compositional differences within fungi and bacteria between the A and B horizons; this variance appeared driven by root biomass and SOC (Figures 2a and b, Supplementary Table S2). Archaea composition did not vary between soil horizons (Figure 2c). We identified significant (P<0.05) belowground, aboveground, and interkingdom correlations on bacterial composition (Figure 2a, Table 1), representing 20% of the total variance in the bacterial matrix. These influences included litter nitrogen, root biomass, SOC, and ammonia-oxidizing archaea abundance. Bacteria and archaea both influenced fungal composition (although not vice versa as inferred from the RDAs), along with several belowground and aboveground influences (archaeal and bacterial richness, AOB abundance, root biomass, SOC, and litter nitrogen); the constrained variance represented 11% of the total variance in fungal composition (Figure 2b). Of the potential explanatory variables, litter nitrogen was the only significant predictor of archaeal composition (Figure 2c; Table 1), although it represented only 2% of the variance. Microbial community compositions of archaea, bacteria, and fungi were similar within microbial groups within a given brome cover class (PERMANOVA, brome cover class explained variance: 2% for each congener, P<0.023; Table 2). Bacterial and fungal communities were similar within horizons (explained variance: 10% for bacteria and 6% for fungi, P<0.001).
Interkingdom soil microbial relationships
The association between bacteria and fungi was significant as indicated by the CoIA (RV=0.365, P<0.001); the first axis represented 47% of the variance. Bacterial–fungal variance in the co-inertia plane was driven mainly by 12 bacterial OTUs. Six of these OTUs in the phyla Acidobacteria, Actinobacteria, and Verrucomicrobia were particularly influential as indicated by significant distances (P<0.05) from the biplot origin (Figure 3a; Supplementary Table S3), although this relationship appeared independent of smooth brome cover (4% explained variance, P=0.509). The correlation between archaea and bacteria community composition was significant (RV=0.357, P=0.036) for the first and second axes of the co-inertia plane (Figure 3b). The first axis represented 30% of the variance and co-varied with 19 archaeal OTUs. Nine of these OTUs were significantly distant (P<0.05) from the biplot origin (Supplementary Table S4) and all from the Candidatus phylum. There was not a clear separation between invaded and uninvaded plots in terms of smooth brome (2% explained variance, P=0.855). The association between archaea and fungi was significant (RV=0.610, P=0.030): the first axis represented 12% of the variance, although sites were differentiated along the second axis based on smooth brome cover (22% explained variance, P=0.042) and 29 archaeal OTUs (nine significantly distant from the biplot origin (P<0.05)), and represented 8% of the variance (Figure 3c; Supplementary Table S4).
Co-inertia analyses of A horizon interkingdom interactions between (a) bacteria and fungi, (b) archaea and bacteria, and (c) archaea and fungi along a smooth brome invasion gradient. The large boxes contain joint site plots colored by smooth brome cover, where arrows link kingdom diversity. Top left, clockwise: arrow tails represent bacteria and the heads represent fungi; tails represent archaea, heads represent bacteria; tails represent archaea, heads represent fungi. Arrow length indicates the strength of the association between matrices: short arrows indicate strong concordance; long arrows, weak concordance. Small boxes: projection of the kingdom of interest onto the co-inertia plane. Bacterial and archaeal OTUs with significant loadings (P⩽0.05) on the smooth brome gradient are indicated by bold red arrows (Supplementary Tables S3 and S4).
Root influence on soil microbial composition
Roots were significantly linked to both bacteria and fungi via biomass and SOC (Figures 2a and b). In the root–bacteria co-inertia analyses, the first and second axes together represented 47% of the variance, with clear separation between smooth brome-invaded and non-invaded sites (20% explained variance, P=0.022; Figure 4a); however, the concordance was insignificant (RV=0.273, P=0.105). Plots were differentiated into invaded and non-invaded by roots from 13 plant species (Figure 4a). There was significant concordance (RV=0.472, P=0.049) between roots and fungi; the first axis represented 15% of the variance and 10 root species differentiated sites (Figure 4b; Supplementary Table S5). Smooth brome clearly separated invaded and non-invaded sites (29% explained variance, P=0.010). Smooth brome reduced the abundance of several native species (Bouteloua gracilis, Carex duriuscula, C. filifolia, Elymus sp., and Galium boreale), which appeared to be linked to bacterial and fungal community composition (Figure 4b; Supplementary Table S5).
Co-inertia analyses of A horizon root-kingdom interactions along a smooth brome invasion gradient. The large panels are joint site plots colored by smooth brome cover, where arrows link root-kingdom diversity. (a) Arrow tails represent roots, and the heads represent bacteria. (b) Arrow tails represent roots, and heads represent fungi. Arrow length indicates the strength of the association between matrices: short arrows indicate strong concordance; long arrows, weak concordance. Small boxes: projection of roots, fungi, or bacteria of interest onto the co-inertia plane. Bold red arrows indicate root species with considerable (P⩽0.05) loadings on the smooth brome gradient (Supplementary Table S5).
Effects of smooth brome abundance on soil microbial structure
Although including direct pathways between smooth brome and fungi diversity resulted in better fit for the SEMs, as indicated by reduced χ2-values (Supplementary Tables S6–S8), the direct pathways were not significant (Figure 5). There were 12 bacterial OTUs important for structuring fungal communities (Supplementary Table S3), with unique relationships between endogenous variables (Supplementary Figure S5) and common pathways among several other OTUs. The SEM analyses suggested three classes of relationships among smooth brome and microbial diversity based on frequency of microbial interactions. In the first class, archaeal diversity increased abundance of four bacterial OTUs (that is, we found significant path coefficients between archaeal diversity and abundance of four bacterial OTUs), with no other significant linkages identified (Figure 5a). In the second class of relationships, increased smooth brome litter was associated with a decrease in abundance of two bacterial OTUs (Figure 5b). Finally, in the third class, increased smooth brome litter was associated with an increase in bacterial diversity in the A horizon, and increased smooth brome root abundance was associated with an increase in bacterial diversity in the B horizon; this increased bacterial diversity then decreased fungal diversity in the A horizon (Figure 5c). Thus, the smooth brome–bacteria and bacteria–fungi pathways were horizon-specific, and bacteria–fungi relationships were more pronounced in the A horizon—with the exception of two rare bacterial OTUs that were horizon-unspecific (Figures 5d and e). The resulting best models (as determined by Comparative Fit Index, Root Mean Square Error of Approximation and Standardized Root Mean Square Residuals) used fungal evenness as the diversity metric for classes 1 and 2 (Supplementary Tables S10 and S13) and fungal richness for class 3 (Supplementary Table S15).
Multigroup SEMs of smooth brome and pH influences on bacterial and fungal diversity (richness, evenness, and phylogenetic diversity) and interkingdom interactions. (a–c) Influential abundant taxa identified through downweighting, RDA, and CoIA. (d, e) Influential rare taxa identified through Poisson regression and RDA. (a) Increased archaeal diversity increases abundance of four bacterial OTUs. (b) Smooth brome shoots reduced abundance of two bacterial OTUs. (c) Smooth brome shoots and roots increase bacterial diversity resulting in decreased fungal diversity. (d) Smooth brome roots increased abundance of OTU 5113. (e) pH had an inverse effect on OTU 2472 (that is, alkaline pH, reduced abundance; acidic pH, increased abundance) and that OTU reduced fungal diversity. Solid arrows represent significant relationships (P<0.05, dashed lines are nonsignificant), the thickness of the arrow indicates the strength of the relationship and the color indicates the direction of the relationship (green=positive, red=negative). Standardized path coefficients are shown next to pathways. Path coefficients can be interpreted as this: if, in a for example, brome shoot abundance increases by one s.d. from the mean, then fungal diversity would increase by 0.027 s.d.'s from its own mean. Thus, the greater the path coefficient, the larger the effect. Where parameters differ between horizons, two path coefficients are shown: the first is the coefficient for the A horizon and the second for the B horizon. All models achieved adequate fit (Supplementary Table S20). Significant path coefficients between smooth brome roots and shoots, bacterial parameters on fungal diversity and archaeal diversity on bacterial parameters are illustrated in the inset bar graph (Supplementary Tables S9–S17). Bars with a height of 0.025 indicate nonsignificant paths. For details on each model class, see Supplementary Information (fungal evenness: Supplementary Table S10, fungal richness: Supplementary Table S9, fungal phylogenetic diversity: Supplementary Table S11, bacterial OTU 5113: Supplementary Table S18, bacterial OTU 2472: Supplementary Table S19).
Discussion
Biological invasions are thought to be among the most serious threats to biodiversity and ecosystem functioning in terrestrial environments (Mooney and Hobbs, 2000). Invasive species often have an impact on soil microbial communities (van der Putten et al., 2007; Rodrigues et al., 2015), and feedbacks from these microbial communities can be essential for invasion success (van der Putten et al., 2001; Smith et al., 2014). Yet, to our knowledge, no study has examined an ecosystem undergoing invasion to include environmental, plant, and multiple kingdom microbial components along the invasion gradient. We expected a trophic cascade in which smooth brome-structured fungi via roots (cf. Johnson et al., 2004) and fungi in turn influenced other microbes. Instead, we found that bacteria mediated fungal responses to brome invasion, whereas archaea influenced select bacteria. Bacteria were the only significant pathway linking aboveground brome effects to fungal changes associated with invasion.
Horizon- and species-specific effects of invasion
Horizon-specific smooth brome effects suggest a decoupling of litter and root effects of invasive species (Elgersma et al., 2011; Ma et al., 2016), although potential litter leachate effects on roots should not be discounted (Scherer-Lorenzen et al., 2003). Our SEM results suggest smooth brome shoot abundance caused changes in the A horizon bacterial community, whereas roots drove changes in the B horizon (Figure 5c). Effects in the A horizon were driven mainly by plant litter-derived nitrogen and its influence on bacteria (Table 1). Similarly, smooth brome litter drove increased abundances of AOB, archaea, and soil nitrogen at this site (Piper et al., 2015a, b).
Given the importance of the plant root community to vesicular–arbuscular mycorrhizal fungi and fungi’s role as the initial decomposers of plant litter (Johnson et al., 2004), we expected direct plant–fungal links. Rather, we found that aboveground biomass effects cascaded into the fungal community via bacteria, with no direct links to the plant root community. Similarly, in the B horizon, roots were linked to bacteria but not to fungi. Thus, a model in which plants and bacteria mediate effects on fungi is congruent with our field observations.
Several possible mechanisms can account for the mediation of plant root effects on fungal communities. First, bacteria could suppress fungi via antibiotic production (Forsberg et al., 2014) and mediation of root exudation (Rudrappa et al., 2008). Second, plant species lost during invasion may also explain why bacteria were key in mediating root effects on the belowground ecosystem. Specifically, we found the loss of roots of native graminoids, including Bouteloua gracilis, Carex spp., and Elymus spp. (Figure 4), was more important to the structuring of fungal and bacterial communities than was the increase in smooth brome roots following invasion. Displacement of slower growing, resource-conservative native species by invasive species changes the community-weighted mean values of root traits such as root nitrogen allocation, specific root length, and root turnover rates (Smith et al., 2014). The shift in soil nitrogen cycling (Craine et al., 2002) driven by fast-growing (N-exploitative) plants like smooth brome may encourage bacterial biomass growth, compared with slow-growing (N-conservative) plants that promote fungal abundance (de Vries et al., 2012).
Another possibility is that plant species-specific mediation of fungi may also be occurring. For example, the native Bouteloua gracilis is a C4 grass that displays frequent mutualisms with fungi in semi-arid grasslands (Herrera et al., 2010), likely due to regulation of heat stress by endophytic root-associated fungi (McLellan et al., 2007). B. gracilis promotes microbial grazer abundance (Ingham et al., 1985) and then benefits from the excreted ammonia. Smooth brome may also alter fungal communities through depleted phosphorus concentrations in the rhizosphere (Wang et al., 2004) or increased root exudates (Dean Stacy et al., 2005). Further, native Carex species structure fungal diversity in grasslands (Johnson et al., 2004) and across environmental gradients (Pellissier et al., 2014) and promote beneficial dark septate fungi (Johnson et al., 2004). Carex reduction of fungal pathogens of bacteria may increase bacterial richness (Johnson et al., 2003). Finally, Elymus species may form symbiotic relationships with both endophytic fungi (White and Morgan-Jones, 1987) and bacteria (Dalton et al., 2004). Our results suggest that the displacement of native species, along with the associated changes in the trait structure of the plant root community, may be a more important (albeit indirect) driver of fungal communities in invaded areas than increases in smooth brome abundance.
Keystone bacterial and archaeal OTUs
In general, some bacteria influenced fungi, archaea influenced bacteria and fungi did not influence the other two kingdoms. Few species emerged as critical for structuring the between-kingdom interactions. Only 10 plant species roots drove the plant–microbial concordance, 12 bacterial OTUs drove the bacterial–fungal concordance, and 18 archaea OTUs drove archaeal–bacterial concordance. Our statistical approach does not explicitly build networks; therefore, these bacterial and archaeal indicators may be the strongest representatives of more diffuse networks that influence other kingdoms.
Archaea and fungal communities were concordant (RV=0.610, P=0.030), and together they differentiated invaded versus native grassland sites. Ammonia-oxidizing archaea are common at this site and increase during invasion, with subsequent effects on nitrification (Piper et al., 2015b). Despite numerous studies of archaea, bacteria, and fungi (van der Heijden, 2008), to our knowledge, none have investigated interactions among the three kingdoms. Here we show that there were many (n=26) links between archaea and fungi, although very little is known about archaeal–fungal interactions (Prosser and Nicol, 2008). Archaea are common, but not overly abundant in roots (Buée et al., 2009, although see Simon et al., 2000); our work suggests that research on the functional roles of archaea in structuring root-associated fungal communities is warranted.
Conclusion
Invasive plant species can have a major impact on the structure and function of soil microbial communities, which is important as soil microbes have a critical role in the functioning of natural ecosystems (Bissett et al., 2013; van der Putten et al., 2013). The effects of plant invasion differ between soil horizons, with shoot litter having a dominant effect in the A horizon and roots in the B horizon. Furthermore, fungal changes in response to plant invasion were mediated via select bacterial species. Critically, native plant species displacement and root loss appeared to be the most important driver of fungal composition during invasion. Thus, we suggest microbial response to plant invasion may be driven not by the plants that are present, but rather by the plants that are now absent. Future work should include investigation of networks among roots and microbial populations (sensu Banerjee et al., 2016; Weiss et al., 2016) to better understand the roles of keystone taxa in changing ecosystem functioning during invasion.
References
Anderson MJ . (2001). A new method for non-parametric multivariate analysis of variance. Austr Ecol 26: 32–46.
Baker GC, Smith JJ, Cowan DA . (2003). Review and re-analysis of domain-specific 16 S primers. J Microbiol Methods 55: 541–555.
Banerjee S, Kirkby CA, Schmutter D, Bissett A, Kirkegaard JA, Richardson AE . (2016). Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol Biochem 97: 188–198.
Bennett JA, Cahill JF . (2016). Fungal effects on plant-plant interactions contribute to grassland plant abundances: evidence from the field. J Ecol 104: 755–764.
Bever JD, Platt TG, Morton ER . (2012). Microbial population and community dynamics on plant roots and their feedbacks on plant communities. Annu Rev Microbiol 66: 265–283.
Bissett A, Brown MV, Siciliano SD, Thrall PH . (2013). Microbial community responses to anthropogenically induced environmental change: towards a systems approach. Ecol Lett 16: 128–139.
Borcard D, Gillet F, Legendre P . (2011) Numerical Ecology with R. Springer: New York.
Bradley BA, Houghton RA, Mustard JF, Hamburg SP . (2006). Invasive grass reduces aboveground carbon stocks in shrublands of the Western US. Global Change Biol 12: 1815–1822.
Buée M, De Boer W, Martin F, van Overbeek L, Jurkevitch E . (2009). The rhizosphere zoo: an overview of plant-associated communities of microorganisms, including phages, bacteria, archaea, and fungi, and of some of their structuring factors. Plant Soil 321: 189–212.
Callaway RM, Thelen GC, Rodriguez A, Holben WE . (2004). Soil biota and exotic plant invasion. Nature 427: 731–733.
Cao Y, Larsen DP, Thorne RS . (2001). Rare species in multivariate analysis for bioassessment: some consideration. J N Am Benthol Soc 20: 144–153.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7: 335–336.
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ et al. (2011). Global patterns of 16 S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA 108: 4516–4522.
Chen Y-L, Chen B-D, Hu Y-J, Li T, Zhang X, Hao Z-P et al. (2013). Direct and indirect influence of arbuscular mycorrhizal fungi on abundance and community structure of ammonia oxidizing bacteria and archaea in soil microcosms. Pedobiologia 56: 205–212.
Clarke KR . (1993). Non-parametric multivariate analyses of changes in community structure. Austr J Ecol 18: 117–143.
Craine JM, Tilman D, Wedin D, Reich P, Tjoelker M, Knops J . (2002). Functional traits, productivity and effects on nitrogen cycling of 33 grassland species. Funct Ecol 16: 563–574.
Dalton DA, Kramer S, Azios N, Fusaro S, Cahill E, Kennedy C . (2004). Endophytic nitrogen fixation in dune grasses (Ammophila arenaria and Elymus mollis from Oregon. FEMS Microbiol Ecol 49: 469–479.
de Boer W, Folman LB, Summerbell RC, Boddy L . (2005). Living in a fungal world: impact of fungi on soil bacterial niche development. FEMS Microbiol Rev 29: 795–811.
de Vries FT, Manning P, Tallowin JRB, Mortimer SR, Pilgrim ES, Harrison KA et al. (2012). Abiotic drivers and plant traits explain landscape-scale patterns in soil microbial communities. Ecol Lett 15: 1230–1239.
Dean Stacy M, Perryman BL, Stahl PD, Smith MA . (2005). Brome control and microbial inoculation effects in reclaimed cool-season grasslands. Rangeland Ecol Manag 58: 161–166.
Diez JM, Dickie I, Edwards G, Hulme PE, Sullivan JJ, Duncan RP . (2010). Negative soil feedbacks accumulate over time for non-native plant species. Ecol Lett 13: 803–809.
Dolédec S, Chessel D . (1994). Co-inertia analysis: an alternative method for studying species–environment relationships. Freshwater Biol 31: 277–294.
Dray S, Dufour AB . (2007). The ade4 package: implementing the duality diagram for ecologists. J Stat Softw 22: 1–20.
Ehrenfeld JG . (2010). Ecosystem consequences of biological invasions. Annu Rev Ecol Evol System 41: 59–80.
Elgersma KJ, Ehrenfeld JG, Yu S, Vor T . (2011). Legacy effects overwhelm the short-term effects of exotic plant invasion and restoration on soil microbial community structure, enzyme activities, and nitrogen cycling. Oecologia 167: 733–745.
Environment Canada. (2016). National Climate Data and Information Archive. Meteorological Service of Canada. http://climate.weather.gc.ca/.
Fierer N, Jackson RB . (2006). The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103: 626–631.
Fink KA, Wilson SD . (2011). Bromus inermis invasion of a native grassland: diversity and resource reduction. Botany 89: 157–164.
Forsberg KJ, Patel S, Gibson MK, Lauber CL, Knight R, Fierer N et al. (2014). Bacterial phylogeny structures soil resistomes across habitats. Nature 509: 612–616.
Frey-Klett P, Burlinson P, Deveau A, Barret M, Tarkka M, Sarniguet A . (2011). Bacterial-fungal interactions: hyphens between agricultural, clinical, environmental, and food microbiologists. Microbiol Mol Biol Rev 75: 583–609.
Gardes M, Bruns TD . (1993). ITS primers with enhanced specificity for Basidiomycetes-application to the identification of mycorrhizae and rusts. Mol Ecol 2: 113–118.
Grace JB . (2006) Structural Equation Modeling and Natural Systems. Cambridge University Press: Cambridge, UK.
Grace JB, Schoolmaster DR, Guntenspergen GR, Little AM, Mitchell BR, Miller KM et al. (2012). Guidelines for a graph-theoretic implementation of structural equation modeling. Ecosphere 3: 1–44.
Herrera J, Khidir HH, Eudy DM, Porras-Alfaro A, Natvig DO, Sinsabaugh RL . (2010). Shifting fungal endophyte communities colonize Bouteloua gracilis: effect of host tissue and geographical distribution. Mycologia 102: 1012–1026.
Hiiesalu I, ÖPik M, Metsis M, Lilje L, Davison J, Vasar M et al. (2012). Plant species richness belowground: higher richness and new patterns revealed by next-generation sequencing. Mol Ecol 21: 2004–2016.
Ingham RE, Trofymow JA, Ingham ER, Coleman DC . (1985). Interactions of bacteria, fungi, and their nematode grazers: effects on nutrient cycling and plant growth. Ecol Monogr 55: 119–140.
Johnson D, Booth RE, Whiteley AS, Bailey MJ, Read DJ, Grime JP et al. (2003). Plant community composition affects the biomass, activity and diversity of microorganisms in limestone grassland soil. Eur J Soil Sci 54: 671–677.
Johnson D, Vandenkoornhuyse PJ, Leake JR, Gilbert L, Booth RE, Grime JP et al. (2004). Plant communities affect arbuscular mycorrhizal fungal diversity and community composition in grassland microcosms. N Phytol 161: 503–515.
Klironomos JN . (2002). Feedback with soil biota contributes to plant rarity and invasiveness in communities. Nature 417: 67–70.
Kobayashi DY, Crouch JA . (2009). Bacterial/fungal interactions: from pathogens to mutualistic endosymbionts. Annu Rev Phytopathol 47: 63–82.
Kõljalg U, Larsson K-H, Abarenkov K, Nilsson RH, Alexander IJ, Eberhardt U et al. (2005). UNITE: a database providing web-based methods for the molecular identification of ectomycorrhizal fungi. N Phytol 166: 1063–1068.
Kourtev PS, Ehrenfeld JG, Häggblom M . (2002). Exotic plant species alter the microbial community structure and function in the soil. Ecology 83: 3152–3166.
Krüger M, Stockinger H, Krüger C, Schüßler A . (2009). DNA-based species level detection of Glomeromycota: one PCR primer set for all arbuscular mycorrhizal fungi. N Phytol 183: 212–223.
Lamb E, Shirtliffe S, May W . (2011). Structural equation modeling in the plant sciences: an example using yield components in oat. Can J Plant Sci 91: 603–619.
Lamb E, Winsley T, Piper CL, Freidrich SA, Siciliano SD . (2016). A high-throughput belowground plant diversity assay using next-generation sequencing of the trnL intron. Plant Soil 404: 361–372.
Legendre P, Legendre L . (2012) Numerical Ecology. Developments in Environmental Modeling 24. 3rd edn. Elsevier Science B.V.: Amsterdam, The Netherlands.
Lozupone C, Knight R . (2005). UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71: 8228–8235.
Lozupone C, Lladser ME, Knights D, Stombaugh J, Knight R . (2011). UniFrac: an effective distance metric for microbial community comparison. ISME J 5: 169–172.
Lynch MDJ, Neufeld JD . (2015). Ecology and exploration of the rare biosphere. Nat Rev Microbiol 13: 217–229.
Ma C, Xiong Y, Li L, Guo D, Briones MJ . (2016). Root and leaf decomposition become decoupled over time: implications for below- and above-ground relationships. Funct Ecol 30: 1239–1246.
Manter DK, Vivanco JM . (2007). Use of the ITS primers, ITS1F and ITS4, to characterize fungal abundance and diversity in mixed-template samples by qPCR and length heterogeneity analysis. J Microbiol Methods 71: 7–14.
Marchant R . (1999). How important are rare species in aquatic community ecology and bioassessment? A comment on the conclusions of Cao et al. Limnol Oceanogr 44: 1840–1841.
McLellan CA, Turbyville TJ, Wijeratne EMK, Kerschen A, Vierling E, Queitsch C et al. (2007). A rhizosphere fungus enhances Arabidopsis thermotolerance through production of an HSP90 inhibitor. Plant Physiol 145: 174–182.
Millard P, Singh BK . (2009). Does grassland vegetation drive soil microbial diversity? Nutr Cycling Agroecosyst 88: 147–158.
Mooney HA, Hobbs RJ . (2000) Invasive Species in a Changing World. Island Press: Washington, DC, USA.
Nielsen UN, Osler GHR, Campbell CD, Burslem DFRP, van der Wal R . (2010). The influence of vegetation type, soil properties and precipitation on the composition of soil mite and microbial communities at the landscape scale. J Biogeogr 37: 1317–1328.
Pearl J . (2012) The causal foundations of structural equation modeling. In: Hoyle RH (ed). Handbook of Structural Equation Modeling. Guilford Press: New York, NY USA, pp 68–91.
Pellissier L, Niculita-Hirzel H, Dubuis A, Pagni M, Guex N, Ndiribe C et al. (2014). Soil fungal communities of grasslands are environmentally structured at a regional scale in the Alps. Mol Ecol 23: 4274–4290.
Piper CL, Siciliano SD, Winsley T, Lamb EG . (2015a). Smooth brome invasion increases rare soil bacterial species prevalence, bacterial species richness and evenness. J Ecol 103: 386–396.
Piper CL, Lamb EG, Siciliano SD . (2015b). Smooth brome changes gross soil nitrogen cycling processes during invasion of a rough fescue grassland. Plant Ecol 216: 235–246.
Poos MS, Jackson DA . (2012). Addressing the removal of rare species in multivariate bioassessments: the impact of methodological choices. Ecol Indicators 18: 82–90.
Prober SM, Leff JW, Bates ST, Borer ET, Firn J, Harpole WS et al. (2015). Plant diversity predicts beta but not alpha diversity of soil microbes across grasslands worldwide. Ecol Lett 18: 85–95.
Prosser JI, Nicol GW . (2008). Relative contributions of archaea and bacteria to aerobic ammonia oxidation in the environment. Environ Microbiol 10: 2931–2941.
Reinhart KO, Royo AA, Van Der Putten WH, Clay K . (2005). Soil feedback and pathogen activity in Prunus serotina throughout its native range. J Ecol 93: 890–898.
Rodrigues RR, Pineda RP, Barney JN, Nilsen ET, Barrett JE, Williams MA . (2015). Plant invasions associated with change in root-zone microbial community structure and diversity. PLoS One 10: e0141424.
Rosseel Y . (2012). lavaan: an R Package for Structural Equation Modeling. Journal of Statistical Software 48: 36.
Rudrappa T, Czymmek KJ, Paré PW, Bais HP . (2008). Root-secreted malic acid recruits beneficial soil bacteria. Plant Physiol 148: 1547–1556.
Scherber C, Eisenhauer N, Weisser WW, Schmid B, Voigt W, Fischer M et al. (2010). Bottom-up effects of plant diversity on multitrophic interactions in a biodiversity experiment. Nature 468: 553–556.
Scherer-Lorenzen M, Palmborg C, Prinz A, Schulze E-D . (2003). The role of plant diversity and composition for nitrate leaching in grasslands. Ecology 84: 1539–1552.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al. (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75: 7537–7541.
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA et al. (2012). Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci USA 109: 6241–6246.
Schröter D, Brussaard L, De Deyn G, Poveda K, Brown VK, Berg MP et al. (2004). Trophic interactions in a changing world: modelling aboveground–belowground interactions. Basic Appl Ecol 5: 515–528.
Shaffer JP . (1995). Multiple hypothesis testing. Annu Rev Psychol 46: 561–584.
Shannon-Firestone S, Reynolds HL, Phillips RP, Flory SL, Yannarell A . (2015). The role of ammonium oxidizing communities in mediating effects of an invasive plant on soil nitrification. Soil Biol Biochem 90: 266–274.
Shipley B . (2016) Cause and Correlation in Biology: a User's Guide to Path Analysis, Structural Equations and Causal Inference with R. 2nd edn. Cambridge University Press: Cambridge, UK.
Siciliano SD, Palmer AS, Winsley T, Lamb E, Bissett A, Brown MV et al. (2014). Soil fertility is associated with fungal and bacterial richness, whereas pH is associated with community composition in polar soil microbial communities. Soil Biol Biochem 78: 10–20.
Simon HM, Dodsworth JA, Goodman RM . (2000). Crenarchaeota colonize terrestrial plant roots. Environ Microbiol 2: 495–505.
Smith B, Wilson JB . (1996). A consumer's guide to evenness indices. Oikos 76: 70–82.
Smith MS, Fridley JD, Goebel M, Bauerle TL . (2014). Links between belowground and aboveground resource-related traits reveal species growth strategies that promote invasive advantages. PLoS One 9: e104189.
Soil Classification Working Group. (1998) The Canadian system of soil classification. Agriculture and Agri-Food Canada Publication 1646. 3rd edn. NRC Research Press: Ottawa, ON, Canada.
Stackebrandt E, Goebel BM . (1994). Taxonomic note: a place for DNA-DNA reassociation and 16 S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Evol Microbiol 44: 846–849.
Stockinger H, Walker C, Schüßler A . (2009). ‘Glomus intraradices DAOM197198’, a model fungus in arbuscular mycorrhiza research, is not Glomus intraradices. N Phytol 183: 1176–1187.
Stockinger H, Krüger M, Schüßler A . (2010). DNA barcoding of arbuscular mycorrhizal fungi. N Phytol 187: 461–474.
Taberlet P, Gielly L, Pautou G, Bouvet J . (1991). Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17: 1105–1109.
Taberlet P, Coissac E, Pompanon F, Gielly L, Miquel C, Valentini A et al. (2007). Power and limitations of the chloroplast trnL (UAA) intron for plant DNA barcoding. Nucleic Acids Res 35: e14.
Teske A, Sørensen KB . (2007). Uncultured archaea in deep marine subsurface sediments: have we caught them all? ISME J 2: 3–18.
van der Heijden MGA . (2008). The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11: 296–310.
van der Putten WH, Vet LEM, Harvey JA, Wäckers FL . (2001). Linking above- and belowground multitrophic interactions of plants, herbivores, pathogens, and their antagonists. Trends Ecol Evol 16: 547–554.
van der Putten WH, Klironomos JN, Wardle DA . (2007). Microbial ecology of biological invasions. ISME J 1: 28–37.
van der Putten WH, Bardgett RD, Bever JD, Bezemer TM, Casper BB, Fukami T et al. (2013). Plant–soil feedbacks: the past, the present and future challenges. J Ecol 101: 265–276.
Vogelsang KM, Bever JD . (2009). Mycorrhizal densities decline in association with nonnative plants and contribute to plant invasion. Ecology 90: 399–407.
Wang Q, Garrity GM, Tiedje JM, Cole JR . (2007). Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73: 5261–5267.
Wang ZY, Kelly JM, Kovar JL . (2004). In situ dynamics of phosphorus in the rhizosphere solution of five species. J Environ Qual 33: 1387–1392.
Weiss S, Van Treuren W, Lozupone C, Faust K, Friedman J, Deng Y et al. (2016). Correlation detection strategies in microbial data sets vary widely in sensitivity and precision. ISME J 10: 1669–1681.
White JF, Morgan-Jones G . (1987). Endophyte-host associations in forage grasses. IX. Concerning Acremonium typhinum, the anamorph of Epichloë typhina. Mycotaxon 29: 489–500.
Wilson SD, Pinno BD . (2013). Environmentally-contingent behaviour of invasive plants as drivers or passengers. Oikos 122: 129–135.
Zuur AF . (2009) Mixed Effects Models and Extensions in Ecology with R. Springer: New York, NY, USA.
Acknowledgements
Funding came from a Natural Sciences and Research Council (NSERC) PGSM scholarship to CLP, NSERC Discovery grants to EGL and SDS, the Department of Plant Sciences, and a Canadian Foundation for Innovation grant to EGL. Alix Schebel, Galen Seilis, Sarah Hardy, Mark Sigouin, Amanda Guy, Sherri Friedrich, and Martin Brummell assisted in the field and/or laboratory, and Joanne Sanche, Jonathan Bennett, Andrew Bissett, and Bobbi Helgason provided insightful comments on an earlier draft of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Mamet, S., Lamb, E., Piper, C. et al. Archaea and bacteria mediate the effects of native species root loss on fungi during plant invasion. ISME J 11, 1261–1275 (2017). https://doi.org/10.1038/ismej.2016.205
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.205
This article is cited by
-
Altered diversity and functioning of soil and root-associated microbiomes by an invasive native plant
Plant and Soil (2022)
-
Archaea: An Agro-Ecological Perspective
Current Microbiology (2021)
-
A survey of invasive plants on grassland soil microbial communities and ecosystem services
Scientific Data (2020)
-
Structural equation modeling of a winnowed soil microbiome identifies how invasive plants re-structure microbial networks
The ISME Journal (2019)
-
Rhizocompartments and environmental factors affect microbial composition and variation in native plants
Journal of Microbiology (2019)