Abstract
The Africanization of the honeybee (Apis mellifera) in South America is one of the most spectacular examples of biological invasions. In this study, we analyzed the Africanization process in Central Mexico along an altitudinal transect from 72 to 2800 m, using both mitochondrial and nuclear DNA markers. The mitochondrial analysis revealed that the two high-altitude populations had a significantly greater percentage of African mitotypes (95%) than the three lowland populations (67%), indicating successful spreading of Africanized swarms to these altitudes. All populations (highland and lowland) had a similar overall proportion of African alleles at nuclear loci (58%). Thus, all populations showed an asymmetric introgression of African nuclear and mtDNA. Colonies with African mitotypes had, on average, significantly more African nuclear alleles (60%) than those with European mitotypes (51%). Furthermore, the three lowland populations showed clear signs of linkage disequilibrium, while the two high-altitude populations did not, indicating recent genetic introgression events into the lowland populations.
Similar content being viewed by others
Introduction
Increasing transport of livestock and mobility of humans has led to a sharp increase of invasive species all over the world (Vitousek et al., 1997). Such biological invasions can pose a serious threat to biodiversity as well as to the public health (Mack et al., 2000; McMichael and Beaglehole, 2000). One of the most spectacular, and from the point of view of the invasive organism, most successful biological invasions has been the Africanization of the honeybee (Apis mellifera) in the Americas (Schneider et al., 2004; Moritz et al., 2005).
Originally the natural distribution of A. mellifera was restricted to Africa, Europe and the Middle East, where about 20 subspecies, displaying distinctive morphological and behavioral features, have been described (Ruttner, 1988). The Americas were free of honeybees until the arrival of the Spaniards. Subsequently, domestic honeybees (mostly of European origin) were imported by the European settlers and successfully established domestic and feral populations (Nogueira-Neto, 1972; Spivak et al., 1991). Interestingly these early honeybee populations probably already included, at a low percentage, African mitotypes (Schiff and Sheppard, 1993, 1995; Sheppard et al., 1999; Clarke et al., 2001); however, these did not trigger any invasive Africanization.
In the 1950s, in an attempt to improve the honey harvest and to obtain a hybrid honeybee better adapted to the tropical climate, roughly 50 A. m. scutellata queens were imported from South Africa to Brazil (Kerr, 1967). These 50 queens formed the nucleus for the following biological invasion in which Africanized swarms spread all over South America and finally reached the United States (Hall and Muralidharan, 1989). Their high reproductive (swarming) capacity, their pre-adaptation to tropical and subtropical climate and their unforeseen capacity to supplant honeybee populations of European origin were the key factors in this biological success story. All attempts to stop the progress of the Africanized swarms proved more or less futile and now finally only low temperature and unfavorable climatic conditions seem to have stopped their expansion towards the north in the USA (Visscher et al., 1997) and towards the south of Argentina and Chile (Kerr et al., 1982).
A number of studies have investigated the process of Africanization of the honeybee using various genetic markers (for example, Hall and Muralidharan, 1989; Smith et al., 1989; Del Lama et al., 1990; Rinderer et al., 1991; Sheppard et al., 1991; Lobo 1995; McMichael and Hall, 1996; Suazo et al., 1998; Quezada-Euán, 2000; Clarke et al., 2001, 2002; Pinto et al., 2005; The Honeybee Genome Sequencing Consortium, 2006; Whitfield et al., 2006). Initially, it was believed that the Africanized honeybees were a hybrid swarm of European and African honeybee subspecies, but the use of mitochondrial DNA markers soon undermined that view (Smith et al., 1989). It turned out that the hybridization process is much more complex and frequently results in asymmetrical levels of introgression in the nuclear and mitochondrial compartments (Clarke et al., 2002; Schneider et al., 2004; Moritz et al., 2005). This asymmetric introgression is a consequence of different behavior (migration, reproduction) of the males and females from the different subspecies. While completely Africanized populations have more than 97% African mitotypes, the introgression of African nuclear genes rarely seems to exceed 65–70% (Hall and Muralidharan, 1989; Smith et al., 1989; Clarke et al., 2002; Schneider et al., 2004; Moritz et al., 2005).
In this study, we analyzed five honeybee populations in Veracruz (Central Mexico) along an altitudinal transect from 72 to 2800 m above sea level (Figure 1) using both mtDNA and nuclear microsatellite markers. Africanized bees arrived in this region in 1987 (Moffett et al., 1987), 12 years before the sampling, presumably allowing the Africanization process enough time to be completed.
Map of Mexico and Veracruz; (a) a map of Mexico with the three Mexican sampling locations, Tamaulipas (Mex88, 250 m), Tapachuala (Mex99, 100 m) and the Veracruz populations (V1, 72 m; V2, 700 m; V3, 1300 m, V4, 1900 m, V5, >2500 m). (b) The detailed view of the State of Veracruz and the location of the Veracruz populations there are shown.
The sample design allowed for the assessment of the impact of environmental conditions on the spread of African genes in the populations with different kinds of markers, either inherited maternally (mtDNA) or by both sexes (microsatellites). Since Africanized colonies are supposed to be adapted to tropical conditions, we would expect them to to have become established at lower altitudes. In contrast European colonies, which are more adapted to colder climate, were exptect to have persisted at higher altitudes.
Materials and methods
Sampling and DNA isolation
The first population along the vertical transect was situated at Soledad de Doblado (further abbreviated V1; 30 colonies sampled) at an altitude of 72 m, the second at Córdoba (V2; 30 colonies) at 700 m, the third at Ixcuatlan del Café (V3; 30 colonies) at 1300 m, the fourth at Calcahualco (V4; 14 colonies) at 1900 m and the fifth at Cuiyachapa (V5; 16 colonies) between 2500 and 2800 m (Figure 1). The samples in the lower altitude populations (V1, V2 and V3) were collected directly from colonies in a 3 km radius, exclusively from beekeepers who were keeping colonies established from locally caught swarms which had no imported European queens. In the two highest locations (V4 and V5) the bees had to be collected from feral colonies because there were no beekeepers in these localities.
To be able to perform a comparison, two additional populations were collected in Mexico (Figure 1) (i) a population located in the far south of Mexico in Tapachula, Chiapas State, close to the Guatemalan border at 100 m altitude, was collected in 1999 as reference for earlier Africanized honeybee populations (Mex99; 30 colonies) and (ii) a population located in Tamaulipas State at 250 m altitude, was collected in 1988 as a reference for pre-Africanization honeybee populations (Mex88; 30 colonies). Finally, since our analysis also required a African population as a reference, we used previously published mitochondrial (Franck et al., 2001) and microsatellite data (Franck et al., 2000b) for an A. m. scutellata population from South Africa (Afric) located near Pretoria. In this particular population the allele frequencies at microsatellite loci were calculated based on the genotypes of the parental generation derived by genotyping the worker offspring of six colonies. A summary of the names of all populations analyzed, the sampling locations, their abbreviations, the altitude and the number of colonies sampled are given in Table 1.
All samples were stored in 96% ethanol at −20°C until DNA extraction. Total DNA was isolated from the thorax of one individual bee per colony using a chloroform phenol extraction following the procedure described by Kocher et al. (1989). By sampling one bee per colony the mitotype of a given colony can be described, as well as 50% of the nuclear genome of a given queen and 100% of the genome of one of the drones she mated with. Thus with 14–30 colonies sampled (depending on population and availability of colonies), it is possible to obtain a representative sample of both the nuclear and mitochondrial DNA variation in a population.
mtDNA analysis
The mitochondrial analysis was carried out according to Garnery et al. (1992), amplifying the tRNAleu-COII mitochondrial region by PCR using the primers E2 (5′-GGCAGAATAAGTGCATTG-3′) and H2 (5′-CAATATCATTGATGACC-3′), followed by a restriction of the PCR product with the enzyme DraI. The length of the obtained PCR products was determined on 1% agarose gels. After the digestion of the PCR product with DraI, the restriction fragments were electrophoretically separated on 10% polyacrylamide gels and visualized in UV light after staining with ethidium bromide.
Microsatellite genotyping
We used eight microsatellite loci A014, A043, A(B)024, A088, A113, Ap043, A028 and A008 (Solignac et al., 2003) to genotype the individual bees at nuclear genetic level. Radioactive PCR were performed under standard conditions as reported by Solignac et al. (2003) except for the A014 locus that was amplified with the primers 5′-GTGTCGCAATCGACGTAACC-3′ and 5′-CGATGACGCGTGGATAAGC-3′ with 1.7 mM of MgCl2 in the reaction buffer and 58°C as annealing temperature. The resulting PCR products of each of the eight loci were heated 5 min at 90°C and were electrophoretically separated on 6% denaturizing polyacrylamide sequencing gels for fragment length analysis.
Statistical analysis
Unbiased estimates and s.d. of gene diversity (He) were calculated according to Nei (1973). Using the software HP-Rare (Kalinowski, 2005), we calculated the allelic richness (ag) in the populations taking into account differences in sample size (Leberg, 2002). We further tested for Hardy–Weinberg equilibrium, linkage equilibrium, cytonuclear disequilibrium and population differentiation using Fisher exact tests (Raymond and Rousset, 1995a) implemented in Genepop (online version; Raymond and Rousset, 1995b). Pairwise Fst values were calculated and tested for their significant deviation from zero using FSTAT ver. 2.9.3.2 (Goudet, 2001).
On the basis of the results of the mtDNA analysis, we calculated the degree of Africanization in the maternal mitochondrial genomes as the percentage of African mitotypes present in a given population sample. To infer the degree of introgression of African alleles into the Mexican populations, we used a Bayesian inference method implemented in BAPS v. 3.1 (Bayesian Analysis of Population Structure; Corander et al., 2003, 2004). This software uses allele frequencies at codominant markers (such as microsatellites loci) to infer geographically distinct groups, to estimate posterior mode of population sub-structure (combination of the geographical groups) using Markov Chain Monte Carlo (MCMC) exploration of parameter space with the assumption of Hardy–Weinberg equilibrium and linkage equilibrium. We used the admixture option based on pre-defined clustering, setting the Afric and Mex88 populations as the parental population of each of the other Mexican populations. BAPS v. 3.1 then estimated the proportion of genes from the Afric and Mex88 population in each individual.
Results
mtDNA analysis
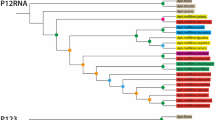
Six different mitotypes (A1, A4, A4′, C1, C2 and M3), previously described (for example, Franck et al., 2001) could be identified among the samples (Table 1). As expected the South African population (Afric) harbored exclusively the mitotypes A1, A4 and A4′, characteristic for the subspecies A. m. scutellata. The other reference population, collected in Mexico in 1988 (Mex88) before the arrival of Africanized swarms, showed a predominantly European pattern (87% C1 and C2) but interestingly also had a small percentage (13%) of the A1 mitotype. The populations sampled in 1999 (V1, V2, V3, V4, V5 and Mex 99) had 78% of mitotypes of the A lineage overall (53% A1 and 25% A4), 20% of C lineage (15% C1 and 5% C2; an eastern European lineages) and 2% of M lineage (M3 a western European lineage). The frequencies of the different mitotypes in the populations are given in Table 1, the joint frequency of African mitotypes is additionally shown in Figure 2.
Degree of introgression of African genes into the populations sampled in Mexico in 1999 (V1, V2, V3, V4, V5 and Mex99). For the mtDNA the frequency of African mtDNA mitotypes is shown, while for the microsatellite markers the degree of Africanization is given as percentage of African nuclear genes estimated with BAPS.
Whereas the three populations at low altitudes (V1, V2 and V3) had only 60–77% of African mitotypes, the two higher altitude populations (V4 and V5), had 94 and 100% of African mitotypes (Table 1). Those differences in mtDNA haplotype frequencies between the low- and high-altitude populations were significant (t-test; t=4.26; P=0.023).
Microsatellite genotyping
A total of 226 individuals were analyzed with the eight microsatellites. The average number of alleles (na) per locus was 9.8±2.1 and the allelic richness (ag) 7.7±1.2 alleles per locus over all analyzed populations (Table 1). The lowest allelic richness (ag) was found in the Mex88 population (4.8±1.5 alleles per locus) and the highest in the Afric population (8.9±1.4 alleles per locus). The Mexican populations from 1999 had an allelic richness between those two extremes, ranging from 7.3±1.6 alleles per locus in the V1 population to 8.5±1.7 alleles per locus in the V2 population. The average number of alleles (na), the allelic richness (ag), the observed heterozygosity (Ho) and the unbiased estimates of gene diversity (He) are given in detail for each of the eight populations in Table 1, the microsatellite allele frequencies of the populations are given in Table 2. One population (population V3 from Mexico 1999) showed a significant deviation from Hardy–Weinberg equilibrium (at the 5% significance level).
The Fisher's exact test for population differentiation revealed that all Mexican populations from 1999 differed significantly from the Mex88 population and from the Afric population, even after Bonferroni adjustment for multiple tests (Table 3; Fst>0.086 and Fst>0.032, respectively). Within the Mexico 1999 populations (V1, V2, V3, V4, V5 and Mex99) 7 out of 15 pairs of populations showed significant differentiation (Table 3). The Mex99 population from the far south of Mexico differed from three out of the five populations from Veracruz (V1, V2 and V4). Interestingly, four pairwise differences were detected between the three low- (V1, V2 and V3) and two high-altitude populations (V4 and V5), while there were no significant pairwise population differentiations within the highland or lowland populations. Overall the Fst values among the Mexico 1999 populations were small and often not significantly different from zero (Table 3).
Even though there was no clear-cut spatial differentiation among the Veracruz populations, we found significant differentiation if we sorted all Veracruz individuals according to their African or European mitotypes. These two resulting groups displayed significant population differentiation (P<0.01) at nuclear loci and apparently form distinct genetic units within the 1999 Veracruz populations.
The tests for linkage- and cyto-nuclear disequilibrium showed a significant deviation from equilibrium in 5 out of 36 pairwise comparisons when applied over the five Veracruz 1999 populations. This is clearly more than the two significant deviations, which would be expected under the null hypothesis (at the 5% level). The three lowland populations showed five significant deviations for linkage and cyto-nuclear disequilibrium, while the two highland populations only displayed two significant deviations. Thus, only the lowland populations displayed more deviations from linkage disequilibrium than expected by chance.
On the basis of the level of admixture calculated by BAPS v. 3.1, the average individual degree of Africanization over all individuals in a given population sample ranged from 54% (±17% s.d.) in the V3 and V5 population to 63% (±15% s.d.) in the V4 population, while V1 (56±20% s.d.) and V2 (61±18% s.d.) displayed intermediate values (Figure 2). The average introgression of African nuclear genes over all populations was 58% (±18% s.d.). Furthermore, no significant differences in the average level of admixture between the three lowland populations and the two highland populations could be detected (t-test; t=−0.28; P=0.41). Also pairwise testing between all five Veracruz populations for differences in the average level of introgression of African nuclear genes yielded no significant results. However, when comparing the degree of nuclear admixture in individuals with African mitotypes vs individuals with European mitotypes in the Veracruz populations, we found a significant difference in the degree of Africanization (t-test; t=2.36; P=0.02). Thus individuals having African mitotypes had an average higher degree of Africanization (n=89; 60±18% s.d.), than individuals with European mitotypes (n=31; 51±18% s.d.).
Discussion
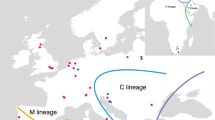
The honeybee samples collected in this study were located along an altitudinal cline in Veracruz (Central Mexico) from 72 to 2800 m with the two extreme points being separated vertically by less than 100 km. Besides the differences in altitude and climate these populations were also influenced in varying degrees by human activity in form of beekeeping, and only the two highest sampling sites are free of beekeeping. We found the highest frequency of African mitotypes (96% African mitotypes) in the two high-altitude populations (V4, V5; 1800–2800 m), while in the three lower altitude populations (V1, V2, V3; 72–1800 m) the frequency of African mitotypes was significantly lower (60–76% African mitotypes). First, this implies that colonies with African mitotypes have no difficulties in coping with the harsher climatic conditions in higher altitudes, since they had nearly completely replaced all European-mitotype colonies there. We would in fact expect an adaptive advantage of the Africanized swarms over the European ones at lower altitudes, and so this result seems unexpected at first glance. The most probable explanation is human influence in form of beekeeping. Even though we took great care to take samples from beekeepers who work with captured feral swarms, there are other beekeepers in the region who regularly import European queens. The more detailed analysis of our mtDNA data is congruent with this hypothesis, since we found almost exclusively eastern European (C1 and C2) mitotypes, and these have been reported in commercially exported A. m. ligustica and A. m. carnica honeybee strains that are favored by beekeepers (Franck et al., 2000a). In contrast, the only European mitotype found in the high-altitude populations was of western European origin (M3). Honeybees of this particular origin (M lineage) are normally not used in commercial beekeeping nowadays and it is thus very unlikely that they were recently introduced by beekeepers.
In a quite extensive study, Clarke et al. (2001) found that during the Africanization of honeybee populations in the Yucatan peninsula, the degree of western European mtDNA (M lineage) increased from 5 to 16%, while the eastern European mitotypes (C lineage) decreased. This effect is commonly explained by the incorporation of western European mitotypes in the advancing Africanized swarms (Schneider et al., 2004) in Brazil already shortly after the first Africanization and hybridization events took place. Also a recent study by Whitfield et al. (2006) using single-nucleotide polymorphisms reports similar results, where Africanized honeybee populations harbur a constant proportion of M alleles, while C-lineage alleles get replaced by A-lineage alleles. Further results of Whitfield et al. (2006) indicate that the M lineage itself is of African origin and thus evolutionarily much closer to the A mitotypes than to the geographically neighboring C mitotypes in Europe. Even though our results show a smaller proportion of western European mitotypes in association with Africanization than reported by Clarke et al. (2001), they are otherwise consistent with the previous studies. In the feral high-altitude populations the only European mitotypes present are M mitotypes while the presence of the C mitotypes is exclusively confined to lower altitudes and thus most likely a result of introductions of C-lineage queens for beekeeping.
No significant differences in the level of African alleles between the low- and high-altitude populations could be detected. However, we found significant linkage disequilibria in the three lowland populations (V1, V2 and V3), while the two highland population (V4 and V5) showed no signs of significant linkage disequilibrium. Again the most likely explanation for this phenomenon is beekeeping and importation of European queens in the lowland populations resulting in recent nuclear introgression.
Interestingly, the colonies displaying African mitotypes within the Veracruz populations had a higher proportion of African alleles (60%) than the colonies displaying European mitotypes (51%). As a result of these different proportions of African alleles there was a significant population differentiation between these two groups, indicating that in fact the colonies with European mitotypes form a genetic subunit within the Veracruz populations. A possible explanation for these differences in proportions of African and European alleles between African and European descendent colonies might be cyto-nuclear incompatibilities and negative heterosis (Smith et al., 1997; Schneider et al., 2003). As the presence of M mitotypes in Africanized populations and the recent findings by Whitfield et al. (2006) suggest, these cyto-nuclear incompatibilities are much more pronounced for the hybrids between the A and C lineage and not so much for hybrids of the A and M lineage.
In this study, we found that Africanized swarms obviously can outcompete European swarms in high altitudes. However, this phenomenon seems not to be universal. Quezada-Euán et al. (2003) found only around 60% African mitotypes in honeybee populations from high-altitude populations (1800–3500 m) in Peru, even though Africanization had started there 18 years before the study. In the Mexican state of Veracruz, the climate is subtropical at low altitude. At higher altitude, that is, above 2500 m, there is an arid desert-like climate, with hot days (commonly above 30°C) and cold nights, especially during the winter, where the temperature can be as low as −5°C. Thus even though the climatic conditions in high altitudes in Veracruz are surely less suitable than at low altitudes, the Africanized bees still have a selective advantage over the European descendent populations, and are able to cope with periods of night frost.
Our study also included one ‘pre-Africanization’ population from Tamaulipas collected in 1988 (Mex88), which contained a small percentage (13%) of African (A1) mitotypes. This result supports the findings of previous studies which also detected African mitotypes in pre-Africanization populations in South- and Meso-America (Schiff and Sheppard, 1993, 1995; Sheppard et al., 1999; Clarke et al., 2001). This pre-Africanization occurrence of A1 mitotypes may reflect the composition of the Iberian honeybees introduced into the Americas by settlers more than 100 years ago. In accordance with this hypothesis, the A1 mitotype has been observed in samples from Spain and Portugal at frequencies reaching up to 40% (Franck et al., 2001).
In sum, these Veracruz populations showed two distinct patterns of Africanization depending on the degree of beekeeping activity present. While the three lowland populations (V1, V2 and V3), which are heavily influenced by beekeeping, displayed only a moderate asymmetry in introgression of African genes (67% African mitotypes, 57% African nuclear genes), the two highland populations (V4 and V5) did not. These two populations at high altitudes, where no beekeeping is present, were nearly exclusively constituted of African mitotype colonies (97% African mitotypes), but had a much lower degree of African nuclear alleles (59% African alleles), thus showing a strong asymmetrical introgression of African genes. The difference in the lowland populations can be explained by the constant introduction of European queens, reflected in the linkage disequlibrium in these populations. We conjecture that, without constant importation of queens, the African mitotype colonies would rapidly replace the European mitotypes leading to a strong asymmetric genetic pattern as found in the feral highland populations.
References
Clarke KE, Oldroyd BP, Javier J, Quezada Euán G, Rinderer TE (2001). Origin of honeybees (Apis mellifera L.) from the Yucatan peninsula inferred from mitochondrial DNA analysis. Mol Ecol 10: 1347–1355.
Clarke KE, Rinderer TE, Franck P, Quezada-Eván JG, Oldroyd BP (2002). The Africanization of honey bees (Apis mellifera L.) of theYucatan: a study of a massive hybridization event across time. Evolution 56: 1462–1474.
Corander J, Waldmann P, Marttinen P, Sillanpaa MJ (2004). BAPS 2: enhanced possibilities for the analysis of genetic population structure. Bioinformatics 20: 2363–2369.
Corander J, Waldmann P, Sillanpaa MJ (2003). Bayesian analysis of genetic differentiation between populations. Genetics 163: 367–374.
Del Lama MA, Lobo JA, Soares AEE, Del Lama SN (1990). Genetic differentiation estimated by isozymic analysis of Africanized honey bee populations from Brazil and from Central America. Apidologie 21: 271–280.
Franck P, Garnery L, Celebrano G, Solignac M, Cornuet JM (2000a). Hybrid origins of honeybees from Italy (Apis mellifera ligustica) and Sicily (A. m. sicula). Mol Ecol 9: 907–921.
Franck P, Garnery L, Loiseau A, Oldroyd BP, Hepburn HR, Solignac M et al. (2001). Genetic diversity of the honeybee in Africa: microsatellite and mitochondrial data. Heredity 86: 420–430.
Franck P, Koeniger N, Lahner G, Crewe RM, Solignac M (2000b). Evolution of extreme polyandry: an estimate of mating frequency in two African honeybee subspecies, Apis mellifea monticola and A.m. scutellata. Insect Soc 47: 364–370.
Garnery L, Cornuet JM, Solignac M (1992). Evolutionary history of the honeybee Apis mellifera inferred from mitochondrial DNA analysis. Mol Ecol 1: 145–154.
Goudet J (2001). FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). J Hered 86: 485–486.
Hall HG, Muralidharan K (1989). Evidence from mitochondrial DNA that African honey bees spread as continuous maternal lineages. Nature 339: 211–213.
Kalinowski ST (2005). HP-Rare: a computer program for performing rarefaction on measures of allelic diversity. Mol Ecol Notes 5: 187–189.
Kerr WE (1967). The history of the introduction of the African bee in Brazil. S African Bee J 39: 3–5.
Kerr WE, de Leon del Rio S, Barrionuevo MD (1982). The southern limits of the distribution of the Africanized honey bee in South America. Am Bee J 122: 196–198.
Kocher TD, Thomas WK, Meyer A, Edwards SV, Paabo S, Villablanca FX et al. (1989). Dynamics of mitochondrial DNA sequence evolution in animals. Proc Natl Acad Sci USA 86: 6196–6200.
Leberg PL (2002). Estimating allelic richness: effects of sample size and bottlenecks. Mol Ecol 11: 2445–2449.
Lobo JA (1995). Morphometric, isozymic and mitochondrial variability of africanized honeybees in Costa Rica. Heredity 75: 133–141.
Mack RN, Simberloff CD, Lonsdale WM, Evans H, Clout M, Bazzaz FA (2000). Biotic invasion: causes, epidemiology, global consequences and control. Ecol Appl 10: 689–710.
McMichael AJ, Beaglehole R (2000). The changing global context of public health. Lancet 356: 495–499.
McMichael M, Hall HG (1996). DNA RFLPs at a highly polymorphic locus distinguish European and African subspecies of the honey bee, Apis mellifera L., and suggest geographical origins of NewWorld honey bees. Mol Ecol 5: 403–416.
Moffett JO, Dale LM, Thomas A, Fierro MM (1987). The Africanized bee in Chiapas, Mexico. Am Bee J 127: 517–519.
Moritz RFA, Härtel S, Neumann P (2005). Global invasions of the western honey bee (Apis mellifera) and the consequences for biodiversity. Ecoscience 12: 289–301.
Nei M (1973). Analysis of gene diversity in subdivided populations. Proc Nat Acad Sci USA 70: 3321–3323.
Nogueira-Neto P (1972). Notas sobre a historia da Apicultura Brasileira. In: De Camargo JMF (ed). Manual de Apicultura. Editora Agronomica Ceres: Sao Paulo. pp 17–32.
Pinto MA, Rubink WL, Patton JC, Coulson RN, Johnston JS (2005). Africanization in the United States: replacement of feral European honeybees (Apis mellifera L.) by an African hybrid swarm. Genetics 170: 1653–1665.
Quezada-Euán JJG (2000). Hybridization between European and Africanized honey bees in tropical Yucatan, Mexico. II. Morphometric, allozymic and mitochondrial DNA variability in feral colonies. Apidologie 31: 1–10.
Quezada-Euán JJG, Perez-Castro E, May-Itza WDJ (2003). Hybridization between European and African-derived honeybee populations (Apis mellifera) at different altitudes in Peru. Apidologie 34: 217–225.
Raymond M, Rousset F (1995a). An exact test for population differentiation. Evolution 49: 1280–1283.
Raymond M, Rousset F (1995b). GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86: 248–249.
Rinderer TE, Stelzer JA, Oldroyd BP, Buco SM, Rubink WL (1991). Hybridization between European and Africanized honey bees in the Neotropical Yucatan Peninsula. Science 253: 309–311.
Ruttner F (1988). Biogeography and Taxonomy of Honeybees. Springer Verlag: Berlin.
Schiff NM, Sheppard WS (1993). Mitochondrial DNA evidence for the 19th century introduction of African honey bees into the United States. Experientia 49: 350–352.
Schiff NM, Sheppard WS (1995). Genetic analysis of commercial honey bees (Hymenoptera: Apidae) from the southern United States. J Econ Entomol 88: 1216–1220.
Schneider SS, DeGrandi-Hoffman G, Smith DR (2004). The African honey bee: factors contributing to a successful biological invasion. Annu Rev Entomol 49: 351–376.
Schneider SS, Leamy LJ, Lewis LA, DeGrandi-Hoffman G (2003). The influence of hybridization between African and European honey bees, Apis mellifera, on asymmetries in wing size and shape. Evolution 57: 2350–2364.
Sheppard WS, Rinderer TE, Garnery L, Shimanuki H (1999). Analysis of Africanized honey bee mitochondrial DNA reveals further diversity of origin. Genet Mol Biol 22: 73–75.
Sheppard WS, Rinderer TE, Mazzoli JA, Stelzer JA, Shimanuki H (1991). Gene flow between African- and European-derived honey bee populations in Argentina. Nature 349: 782–784.
Smith DR, Crespi BJ, Bookstein FL (1997). Fluctuating asymmetry in the honeybee, Apis mellifera: effects of ploidy and hybridization. J Evol Biol 10: 551–574.
Smith DR, Taylor OR, Brown WW (1989). Neotropical Africanized honey bees have African mitochondrial DNA. Nature 339: 213–215.
Solignac M, Vautrin D, Loiseau A, Mougel F, Baudry E, Estoup A et al. (2003). Five hundred and fifty microsatellite markers for the study of the honeybee (Apis mellifera L. genome). Mol Ecol Notes 3: 307–311.
Spivak M, Fletcher DJC, Breeds MD (eds) (1991). The ‘African’ Honeybee. Westview Press: San Francisco, CA.
Suazo A, McTiernan R, Hall HG (1998). Differences between African and European honey bees (Apis mellifera L.) in random amplified polymorphic DNA (RAPD). J Hered 89: 32–36.
The Honeybee Genome Sequencing Consortium (2006). Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443: 931–949.
Visscher PK, Vetter RS, Baptista FC (1997). Africanized bees, 1990–1995: initial rapid expansion has slowed in the US. Calif Agric 51: 22–25.
Vitousek PM, D’Antonio CM, Loope LL, Rejmánek M, Westbrooks R (1997). Introduced species: a significant component of human-caused global change. New Zeal J Ecol 21: 1–16.
Whitfield CW, Behura SK, Berlocher SH, Clark AG, Johnston SG, Sheppard WS et al. (2006). Thrice out of Africa: ancient and recent expansions of the honey bee Apis mellifera. Science 314: 642–645.
Acknowledgements
We thank Daniel Grimanelli for his basic and motivating training, as well as Jean-Marie Cornuet and Marc-Edouard Colin for their initial support to this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kraus, F., Franck, P. & Vandame, R. Asymmetric introgression of African genes in honeybee populations (Apis mellifera L.) in Central Mexico. Heredity 99, 233–240 (2007). https://doi.org/10.1038/sj.hdy.6800988
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800988
Keywords
This article is cited by
-
Prevalence and distribution of Varroa destructor and Nosema spp. in symptomatic honey bee colonies across the USA from 2015 to 2022
Scientific Reports (2024)
-
Coexistence of honeybees with distinct mitochondrial haplotypes and hybridised nuclear genomes on the Comoros Islands
The Science of Nature (2021)
-
The genetic consequences of the anthropogenic movement of social bees
Insectes Sociaux (2016)
-
Stock composition of northern neotropical honey bees: mitotype and morphotype diversity in Mexico (Hymenoptera: Apidae)
Apidologie (2016)
-
Impact of Varroa destructor on honeybee (Apis mellifera scutellata) colony development in South Africa
Experimental and Applied Acarology (2015)